Single Nucleotide Polymorphisms Jennifer Lyon Eskind Biomedical Library

Single Nucleotide Polymorphisms Jennifer Lyon Eskind Biomedical Library May 1, 2009 CRC Workshop Series
Types of Genetic Variations • • • Single Nucleotide Polymorphisms (SNP) – Single base pair changes GTCATTCGATT GTCAGTCGATT Indels – Small insertion/deletions CTT------GATC CTTACGGATC Small variable repeats – microsatellites – ACGACGACG (6 copies) – ACGACGACGACG (7 copies) Variable Long tandem repeats (can be dozens to hundreds to thousands) Chromosomal Aberrations: Translocations, Inversions, etc.

Focusing on SNPs • • Types of SNPs SNP nomenclature Resources for SNPs Examples and Challenges in Finding SNPs http: //learn. genetics. utah. edu/content/health/pharma/snips/

SNPs Types • SNPs can be categorized in a number of ways, the most common are by location and function (relative to a gene) • Intragenic SNPs are often categorized by function – are they in a coding region, an intron, part of the m. RNA, outside the m. RNA but still in the gene locus (i. e. , in the promoter) • Extragenic SNPs may be considered simply ‘genomic’ or might be labeled relative to the nearest gene, ie. 5’ or 3’ to a gene An ‘extragenic’ SNP may affect regulatory regions important in gene expression or other DNA functions such as DNA replication.

SNP Functional Categories • coding nonsynonymous – Missense, nonsense, frame shift • coding synonymous • Intronic – splice site • m. RNA utr – 5' utr or 3' utr • (gene) locus region (5’ or 3’ to the gene) – ‘near gene’ usually means within ~2000 bp of gene • genomic/extragenic (distant from any gene)
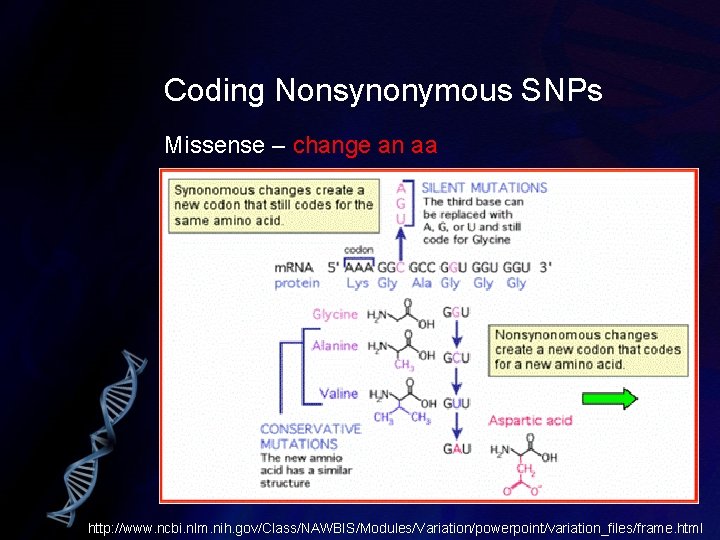
Coding Nonsynonymous SNPs Missense – change an aa http: //www. ncbi. nlm. nih. gov/Class/NAWBIS/Modules/Variation/powerpoint/variation_files/frame. html

Coding Non-Synonymous SNPs • Nonsense – Change an aa to a stop codon – Results in a shortened protein • Frame Shift – Are really single-base indels – Drop or add one base and the triplet reading frame is thrown out of shift, altering all downstream aa’s and usually resulting in an earlier stop codon

SNP Nomenclature • The Human Genome Variation Society (http: //www. hgvs. org/mutnomen/recs. html) has proposed some guidelines for SNP nomenclature, but at the moment, there is minimal consistency. • Different sources will refer to the same SNP in different ways • While db. SNP identifiers (rs#12345678) are becoming common, they are not required of publishing authors and not used in all cases.

SNPs at Base-Pair Level • The base-pair change is given in various forms: A/C T→G C>T 432 G>C T 73 C The HGVS nomenclature recommendations: "c. " for a coding DNA sequence (like c. 76 A>T) "g. " for a genomic sequence (like g. 476 A>T) "m. " for a mitochondrial sequence (like m. 8993 T>C "r. " for an RNA sequence (like r. 76 a>u)

Position, position! • The big issue with SNPs is identifying their location (numerically). • Position can be specified: – Number location within a specific sequence – Relative to another genetic landmark • Start site for a coding region of a gene • Start or end of an exon or intron • Relative to a marker • Published articles are not always clear on this!!! • Different resources may use different landmarks/numbering • Numbering is always relative to the chosen sequence

Coding SNPs • These are easier because they can be identified by the amino acid position rather than the basepair position • Most common nomenclature uses either 3 -letter or single amino acid codes: Asn 332 Asp OR A 95 V • The HGVS recommendation is similar: "p. " for a protein sequence (like p. Lys 76 Asn) • Amino Acid (protein) coding sequence positions becoming more consistent, but are not always consistent

Database of SNPs (db. SNP) db. SNP • is the international central repository for both single base nucleotide substitutions and short deletion and insertion polymorphisms • accepts data submissions from scientists • is integrated with the NCBI’s Entrez system

db. SNP Content The SNP database has two major classes of content: • Submitted data, i. e. , original observations of sequence variation: Submitted SNPs (SS) with ss# (ss 5586300) • Computed/curated data: Reference SNP Clusters (Ref SNP) with rs# (rs 4986582)

Reference SNP Clusters • Ref SNP clusters are computer-generated and curated by NCBI staff • Ref SNP Clusters define a non-redundant set of SNPs • All individual SNPs submitted by a researcher are given a submitter SNP number (ss#) and then redundant (repetitive) submitter SNPs are combined into a Ref. SNP cluster record, with a unique rs# • Ref SNP clusters may contain multiple submitted SNPs

Searching db. SNP • db. SNP is searched like any other Entrez db • Specialized fields include: Field Tag Notes Allele [Allele] Uses IUPAC codes for bases Chromosomal Location [CHRPOS] Uses chromosomal base-pair locations Contig Position [ctpos] Uses contig base-pair locations Function Class [Func] Includes coding synonymous, missense, nonsense, intron, utr, etc. SNP Class [SNP_Class] Includes snp, indel, mixed
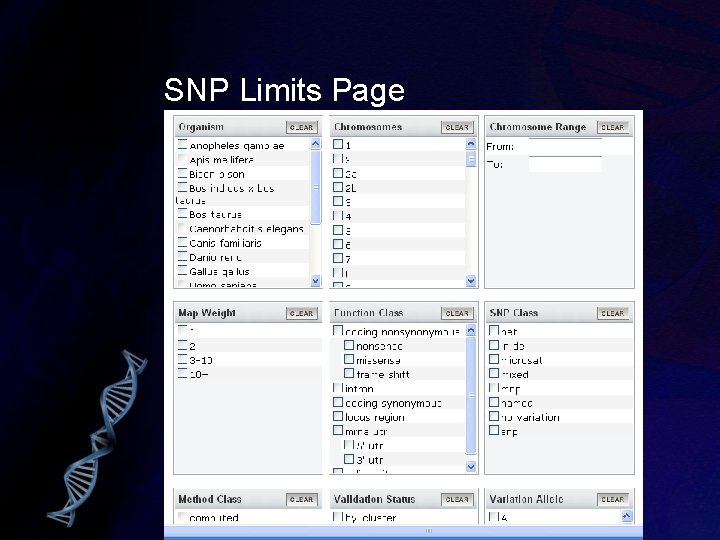
SNP Limits Page

Creating a Complex Search Retrieve all synonymous coding reference SNPs for the human norepinephrine transporter gene (Slc 6 a 2) from db. SNP Search Strategy: human[orgn] AND Slc 6 a 2[gene] AND “coding synonymous” [FUNC] Note: To use the [gene] (gene name) field, it is necessary to have the official gene name or gene symbol as per the Human Gene Nomenclature Committee. Entrez Gene can be used to find these.
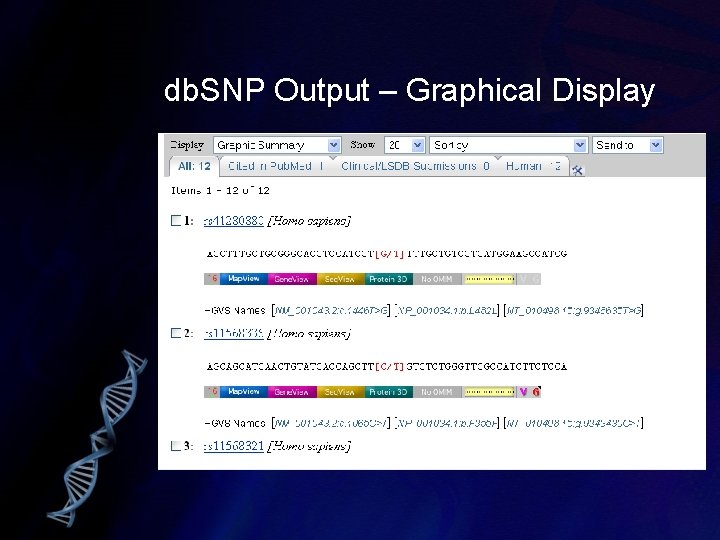
db. SNP Output – Graphical Display

db. SNP - Live • Let’s look at a db. SNP reference SNP page: • http: //www. ncbi. nlm. nih. gov/SNP/snp_ref. cgi? rs =3743788

Finding SNPs - Challenges • If rs# is available – start with it • Not all rs#s have information in all databases • Another database of interest is the Online Mendelian Inheritance in Man (OMIM) • OMIM doesn’t always provide rs#s even when there is one • db. SNP records may link to OMIM or may not, even if the SNP is in an OMIM record

Example 1 • rs 1800888 • (C>T) → Ile 164 Thr in ADRB 2 gene • HGVS nomenclature – NP_000015. 1: p. T 164 I To Find in OMIM • Search with rs 1800888 – yield nothing • Search with ADRB 2[gene] – find record • Look at allelic variants: . 0003 BETA-2 ADRENORECEPTOR AGONIST, REDUCED RESPONSE TO [ADRB 2, THR 164 ILE ] • It is a match

Example 2 • rs 2740574 • A/G SNP located 5’ to CYP 3 A 4 • HGVS nomenclature: – NT_007933. 14: g. 24616372 C>T To find in OMIM • Search with rs 2740574 yields nothing • Search with gene name CYP 3 A 4 – find record • Find list of allelic variants -. 0001 CYP 3 A 4 PROMOTER POLYMORPHISM [CYP 3 A 4, a-g PROMOTER] • Compare info in db. SNP to info in OMIM (look at sequence)

Other Databases • • OMIM – NCBI Hap. Map - International Hap. Map Project ALFRED – Allele Frequence Databases HGVbase. G 2 P - Human Genome Variation database of Genotype-to-Phenotype information • Pharm. GKB – Pharmacogenomics Knowledgebase • F-SNP – Functional SNPs
- Slides: 23