Simon Fraser University Department of Molecular Biology and

Simon Fraser University Department of Molecular Biology and Biochemistry Vancouver Bioinformatics Users Group (Van. BUG) Polymorphic segmental duplications in the nematode Caenorhabditis elegans Ismael Vergara Chen Lab Thursday February 12 th, 2009


Introduction – Different types of genomic structural variations have been found in most eukaryotic genomes. – Duplications at the gene, segment and genome level have been detected. – Duplications are a driving force for the generation of new genes.

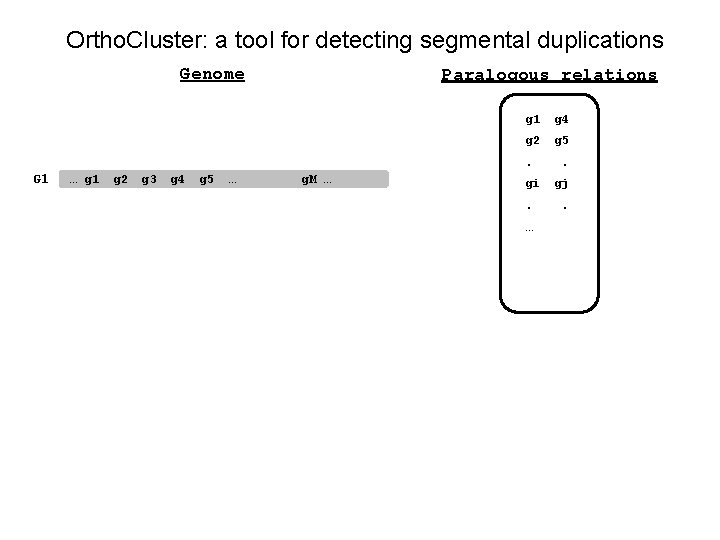
Ortho. Cluster: a tool for detecting segmental duplications Genome Paralogous relations g 1 g 4 g 2 g 5 . G 1 … g 1 g 2 g 3 g 4 g 5 … g. M … gi. … . gj.
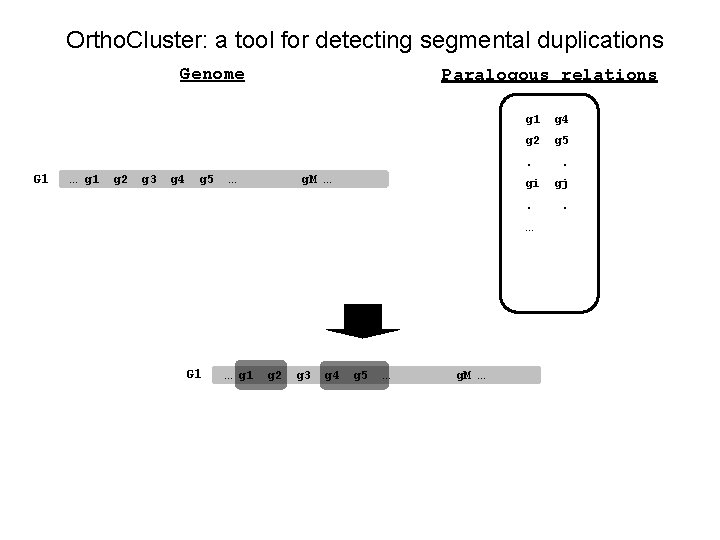
Ortho. Cluster: a tool for detecting segmental duplications Genome Paralogous relations g 1 g 4 g 2 g 5 . G 1 … g 1 g 2 g 3 g 4 g 5 … g. M … gi. … G 1 … g 1 g 2 g 3 g 4 g 5 … g. M … . gj.

C. elegans: a model organism

C. elegans: a model organism Sydney Brenner Established C. elegans as a model organism. Nobel Prize in Physiology or Medicine (2002)

C. elegans: a model organism Sydney Brenner Established C. elegans as a model organism. Nobel Prize in Physiology or Medicine (2002) The C. elegans Sequencing Consortium, 1998

(1) Distribution of perfect segmental duplications in C. elegans
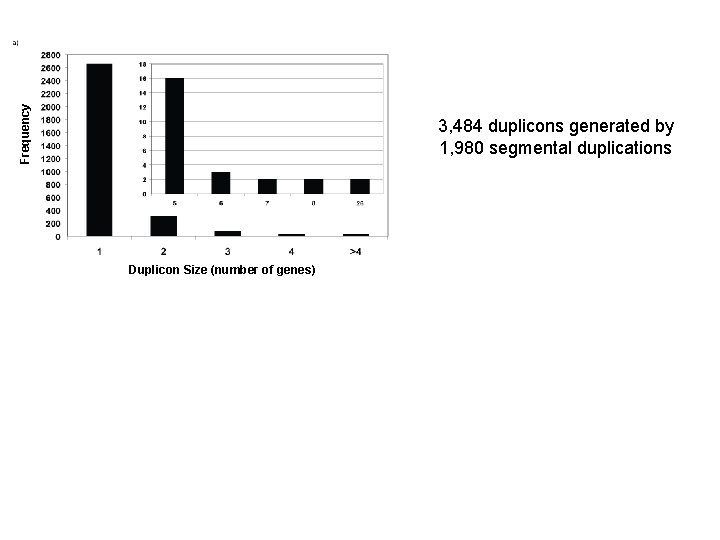
Frequency 3, 484 duplicons generated by 1, 980 segmental duplications Duplicon Size (number of genes)
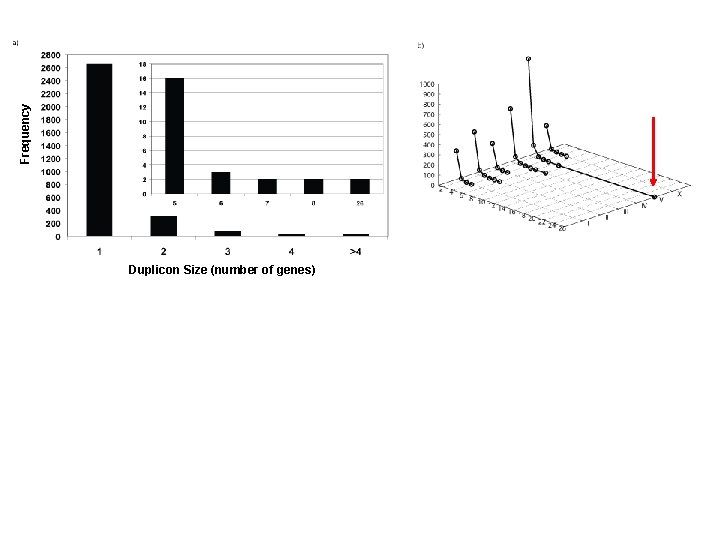
Frequency Duplicon Size (number of genes)
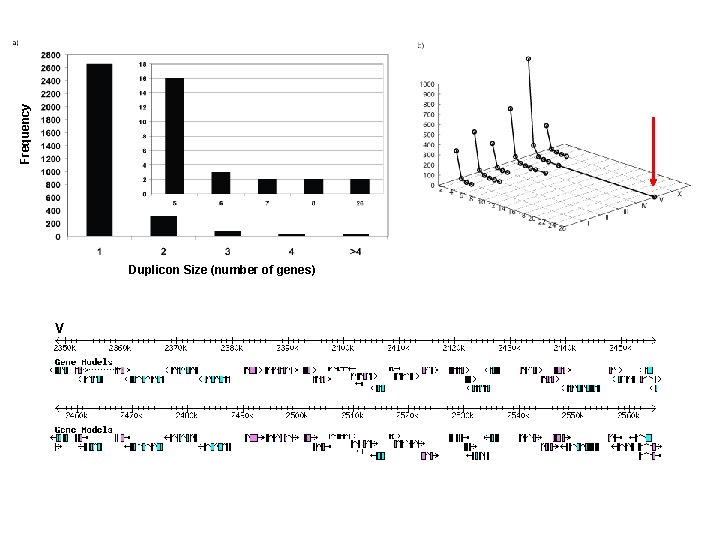
Frequency Duplicon Size (number of genes) V

(2) Molecular characterization of the largest segmental duplication


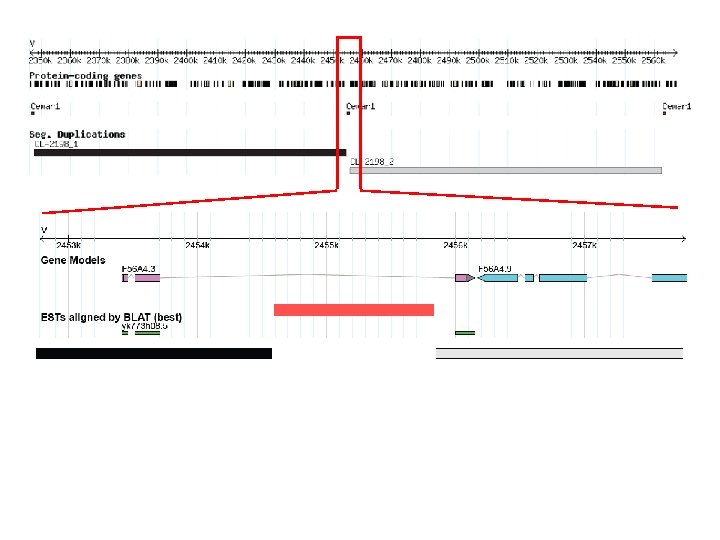

99. 7% similarity

99. 7% similarity

(3) Experimental characterization of the largest segmental duplication
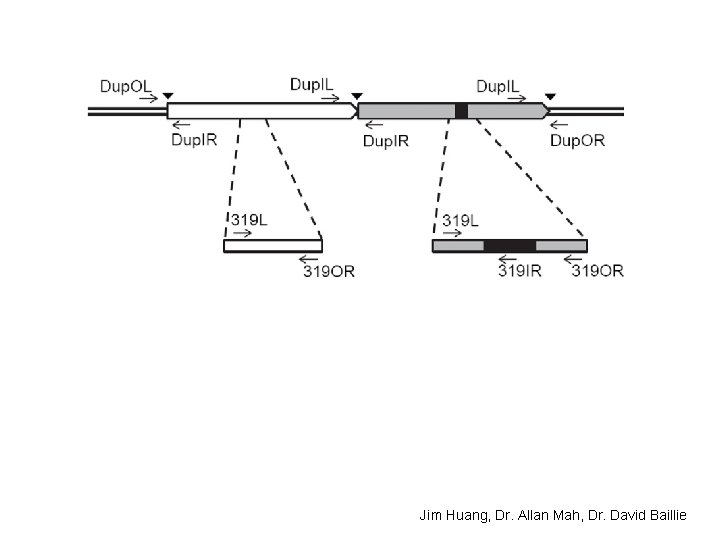
Jim Huang, Dr. Allan Mah, Dr. David Baillie
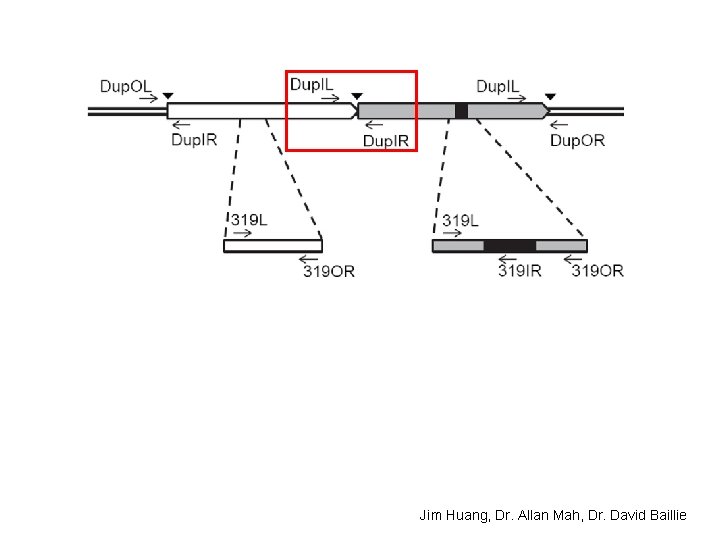
Jim Huang, Dr. Allan Mah, Dr. David Baillie
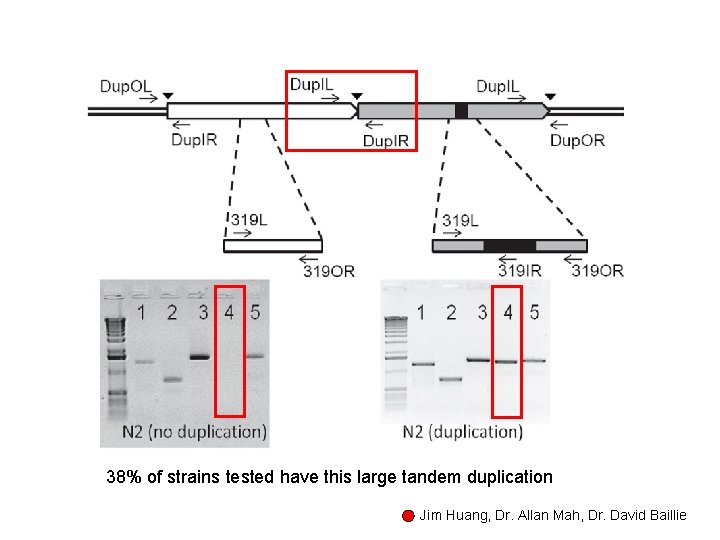
38% of strains tested have this large tandem duplication Jim Huang, Dr. Allan Mah, Dr. David Baillie
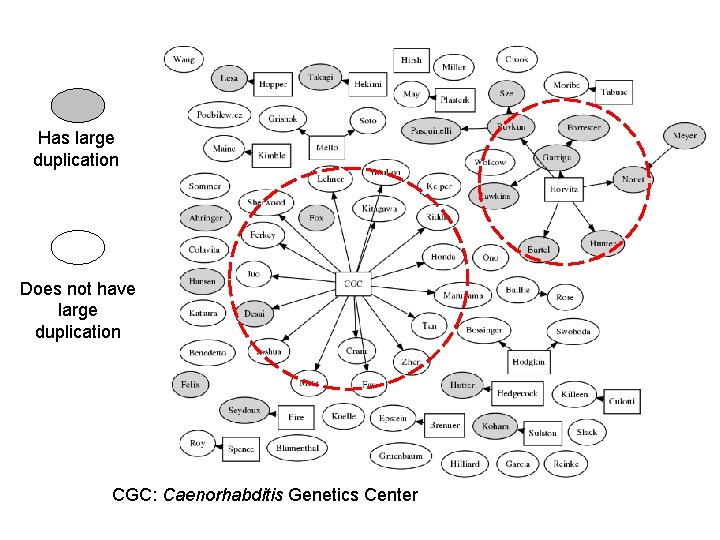
Has large duplication Does not have large duplication CGC: Caenorhabditis Genetics Center
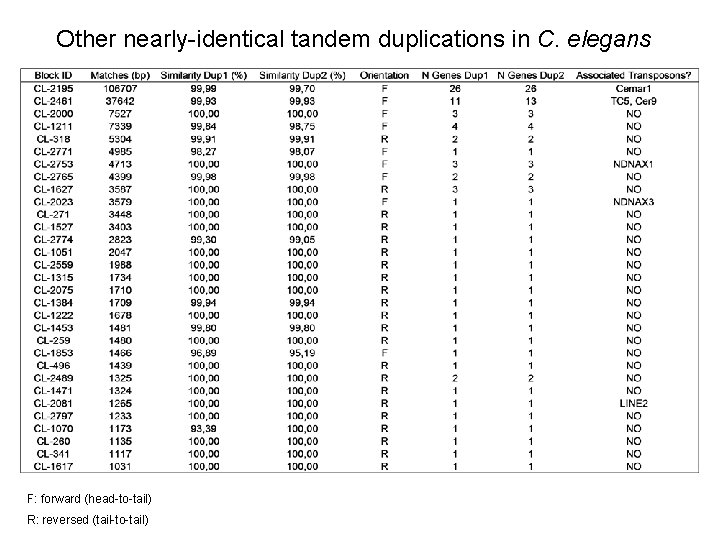
Other nearly-identical tandem duplications in C. elegans F: forward (head-to-tail) R: reversed (tail-to-tail)

Next Steps • Extend this analysis to all nearly-identical tandem duplications. • Inspect further the correlation between different types of segmental duplications and Transposable Elements. • Analyze the function of F 56 A 4. 3 and its potential role in stabilizing the segmental duplication.

Conclusions • We have developed a computer program, Ortho. Cluster, for the genome-wide detection of segmental duplications. • We have identified 1, 980 perfect segmental duplications in C. elegans. • The largest segmental duplication is polymorphic in Wild Type C. elegans strains. • This large duplication may represent only one of many cases of polymorphic regions within C. elegans strains.

Ortho. Cluster. DB http: //genome. sfu. ca/orthoclusterdb

Acknowledgements • Dr. Jack Chen • Dr. Maja Tarailo • Dr. Jian Pei (CS) • Dr. Ke Wang • Xinghuo Zeng • • Dr. David Baillie (MBB) Dr. Allan Mah Dr. Bob Johnsen Jim Huang • The C. elegans community • Supported by Weyerhaeuser Molecular Biology and Biochemistry Fellowship, NSERC, and MSFHR.


Ortho. Cluster: a tool for the detection of synteny blocks Genomes G 1 … g 11 g 12 g 13 g 14 g 15 … g 1 M … G 2 … g 21 g 22 g 23 g 24 g 25 … g 2 M … G 3 … g 31 g 32 g 33 g 34 g 35 … g 3 M … . . . GN … g. N 1 g. N 2 g. N 3 g. N 4 g. N 5 … g. NM …

Ortho. Cluster: a tool for the detection of synteny blocks Genomes Correspondence File … G 1 … g 11 g 12 g 13 g 14 g 15 … g 1 M … G 2 … g 21 g 22 g 23 g 24 g 25 … g 2 M … G 3 … g 31 g 32 g 33 g 34 g 35 … g 3 M … … g. N 1 g. N 2 g. N 3 g 21 g 31 … g. N 1 g 12 g 22 g 32 … g. N 2 g 13 g 23 g 33 … g. N 3 g 14 g 24 g 34 … g. N 4 . . . GN g 11 g. N 4 g. N 5 … g. NM … g 1 M … g 2 M g 3 M … g. NM

Ortho. Cluster: a tool for the detection of synteny blocks Genomes Correspondence File … G 1 … g 11 g 12 g 13 g 14 g 15 … g 1 M … G 2 … g 21 g 22 g 23 g 24 g 25 … g 2 M … G 3 … g 31 g 32 g 33 g 34 g 35 … g 3 M … … g. N 1 g. N 2 g. N 3 g 21 g 31 … g. N 1 g 12 g 22 g 32 … g. N 2 g 13 g 23 g 33 … g. N 3 g 14 g 24 g 34 … g. N 4 . . . GN g 11 g. N 4 g. N 5 … g. NM … g 1 M g 2 M g 3 M … G 1 … g 11 g 12 g 13 g 14 g 15 … g 1 M … G 2 … g 21 g 22 g 23 g 24 g 25 … g 2 M … G 3 … g 31 g 32 g 33 g 34 g 35 … g 3 M … . . . GN … g. N 1 g. N 2 g. N 3 g. N 4 g. N 5 … g. NM … … g. NM
- Slides: 34