Shrunken Centroid Ordering by Orthogonal Projections SCOOP method

Shrunken Centroid Ordering by Orthogonal Projections (SCOOP) method of variable selection Joe Verducci Ohio State University

Outline ¡ Motivation—gene expression l l l Variable selection for LDA Large p Moderate n Advantages in gene selection Method ¡ Model Justification ¡ Measures of Performance ¡ Modifications ¡

LDA Motivation ¡ ¡ ¡ Non-greedy selection: l preserve (augmented) discriminant information Variables with between group differences Variables highly correlated with these

Fisher’s Linear Discriminant Function and A Stupid Generalization where

Why It’s Stupid S= m 1 = 0 0 1 1 0 0 0 0 m 2 = 0 0 -1 Results from Bickel and Levina (2004) imply that the eigenvectors of within and between group covariance matrices approach orthogonality under n fixed p infinity asymptotics.

Genetic Motivation ¡ Wound Healing l l 80 National Wound Healing Clinics 1000 patients Initial + 1 -week samples ¡ Clinical records of patients ¡ l l l ~10 K genes of potential interest in myocytes Subsets of genes act in concert A single gene may be active in several subsystems

P 53 ¡ ¡ ¡ When the DNA in a cell becomes damaged by agents such as toxic chemicals or ultraviolet (UV) rays from sunlight, this protein plays a critical role in determining whether the DNA will be repaired or the cell will undergo programmed cell death (apoptosis). If the DNA can be repaired, tumor protein p 53 activates other genes to fix the damage. If the DNA cannot be repaired, tumor protein p 53 prevents the cell from dividing and signals it to undergo apoptosis. This process prevents cells with mutated or damaged DNA from dividing, which helps prevent the development of tumors.
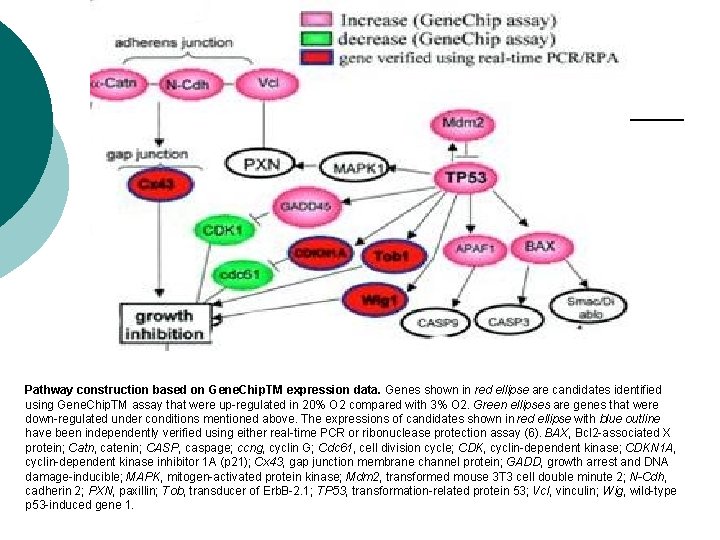
Pathway construction based on Gene. Chip. TM expression data. Genes shown in red ellipse are candidates identified using Gene. Chip. TM assay that were up-regulated in 20% O 2 compared with 3% O 2. Green ellipses are genes that were down-regulated under conditions mentioned above. The expressions of candidates shown in red ellipse with blue outline have been independently verified using either real-time PCR or ribonuclease protection assay (6). BAX, Bcl 2 -associated X protein; Catn, catenin; CASP, caspage; ccng, cyclin G; Cdc 61, cell division cycle; CDK, cyclin-dependent kinase; CDKN 1 A, cyclin-dependent kinase inhibitor 1 A (p 21); Cx 43, gap junction membrane channel protein; GADD, growth arrest and DNA damage-inducible; MAPK, mitogen-activated protein kinase; Mdm 2, transformed mouse 3 T 3 cell double minute 2; N-Cdh, cadherin 2; PXN, paxillin; Tob, transducer of Erb. B-2. 1; TP 53, transformation-related protein 53; Vcl, vinculin; Wig, wild-type p 53 -induced gene 1.

Motivating Simple Example ¡ Two groups l ¡ 50 samples in each P= 4000 normal variables l l All have variance 1 First 10 variables ¡ correlation =. 75 between all pairs ¡ Difference of 2 between group means Second 10 variables ¡ correlation =. 75 between all pairs ¡ Difference of 1 between group means Last 3980 variables ¡ independent ¡ same mean in both groups

Results from 100 Simulations ¡ Individual t-test ranking by p-values l l ¡ 73% of top 20 selected are correct On average need to select 400 variables to ensure inclusion of all 20 SCOOP l l 91% of top 20 selected are correct On average need to select 200 variables to ensure inclusion of all 20

Shrunken Centroid Method for K groups Tibshirani, Hastie, Narasimhan & Chu ¡ For each gene i, l l ¡ xik = sample mean in group k, xi = overall sample mean sik = estimated std. error of xik ¡ Based on pooled std deviation dik = (xik - xi)/sik is a t-statistic Shrinking by an amount D > 0 gives l Shrunken difference • Shrunken centroid

Properties of Shrunken Centroid ¡ ¡ ¡ When K = 2, ordering of variables/genes is same as t-test Keeps “redundant” predictors Can be modified to regularize the estimated std errors Shrunken centroids used directly for classification Shrinkage by amount D is simultaneous in all coordinates on standardized scale Shrinkage parameter D chosen by crossvalidation

Reformulating the Goals ¡ Genetic studies l Find biomarkers classification/prediction ¡ Use small number of classifiers/predictors ¡ l Understand genetic pathways Discover which genes work together to make a difference ¡ possible intervention ¡ ¡ Other studies l Improve efficiency in difficult discrimination problems

SCOOP Method (version 1) ¡ ¡ ¡ Define the Augmented Discriminant Space: ADS = span of eigenvectors of Within and Between Covariance Matrices Modify shrinkage so as not to distort configuration of data in the ADS shrink variables differentially along directions orthogonal to the ADS Note: Unlike the reference, we do not standardize, but scale only at the shrinkage stage. Keep track of the amount of shrinkage li needed to eliminate the ith variable
![SCOOP Algorithm for K groups 1. Between Group eigenvectors DB = [(xik - xi)] SCOOP Algorithm for K groups 1. Between Group eigenvectors DB = [(xik - xi)]](http://slidetodoc.com/presentation_image/1cff8c3df7f92ea41adbf7779ec1cb2e/image-15.jpg)
SCOOP Algorithm for K groups 1. Between Group eigenvectors DB = [(xik - xi)] p x K matrix Use Singular Value Decompostion (SVD) on DB. The singular vectors of DB are the eigenvectors of DB (DB)T 2. Within Group eigenvectors

Algorithm (part 2) ¡ Orthogonalize the Between group (BG) eigenvectors to the Within group (WG) eigenvectors l ¡ ¡ Note: residuals from orthogonalization will no longer be orthogonal to each other Renormalize compute projection operator onto complement of the ADS l Note: do not need to use p x p storage

Algorithm (part 3) ¡ Order variables by scaled shrinkage distances {li} l l For each variable i, compute a scale value = (squared) length of its projection onto the orthogonal complement of the ADS Then calculate how many [li] such units are needed to shrink each of the K mean differences to 0

Notes ¡ Shrinking is non-linear l l ¡ it truncates at 0 shrinks each group only as much as it needs to What to use as a stopping rule? l l l Some measure of preserved information Elbow in the distribution of {li} Reference to extreme value distribution

Theoretical Concern Inconsistency of sample eigenvectors l if p(n)/n c > 0 ¡ l Johnstone and Lu (2004) Unless sparse representation (offset) factor model ¡ Latent factors account for both ¡ l l Correlation among variables Group mean differences

Modeling considerations ¡ Common offset factor model for gene expression l l ¡ Normally distributed data l l l ¡ latent factors represent biological variation random measurement error are “uniqueness” components of individual genes. two populations share the same factor structure differ only by the means of the underlying factors the restricted maximum likelihood procedure is the (stupid) generalization of Fisher’s Linear Discriminant Analysis (SLDA) that incorporates a generalized inverse of the pooled sample covariance matrix. SLDA seldom works well for real data l amend overly restrictive assumptions on both means and covariances.

More model considerations ¡ Factors underlying biological variation l Common factors in 2 groups ¡ ¡ l Group specific factors ¡ ¡ ¡ Some with different means in 2 groups Some with same mean Some may have non-zero means Some have 0 means Unique variation among genes l l . Most is noise A few of the genes that do not load on any factor may have different means in the two groups

Model

Simulation ¡ ¡ ¡ ¡ n=100 ¡ Loadings on common factors l l 1 indicates 1 st 10 variables [1] p=4000 l l 2 indicates 2 nd 10 variables [. 55] G=2 l l 3 indicates 3 rd 10 variables [0] K=3 ¡ Loadings on Group-specific factors (g) J =1 l L 1(1) indicates 4 th 10 variables [. 55] s=1 l L 1(2) indicates 5 th 10 variables [0] sk. F=1 Sj(g)=1 Here [] is the difference in means
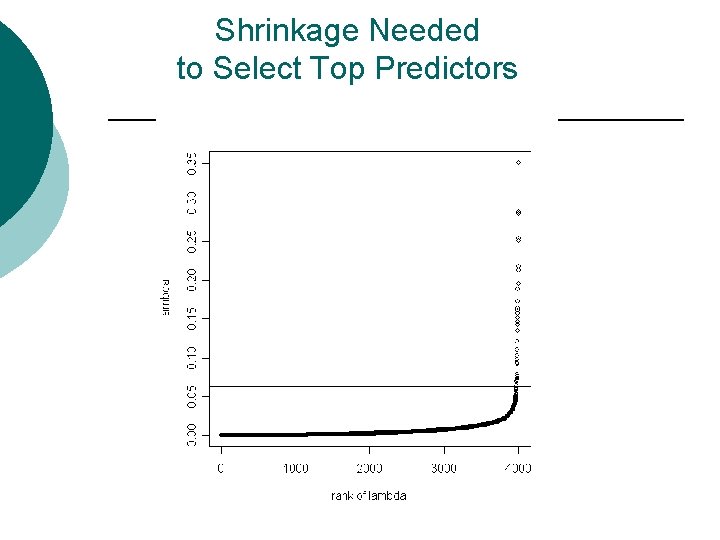
Shrinkage Needed to Select Top Predictors

Measures of Performance ¡ Individual t-test ranking by p-values l l ¡ 49% of top 30 selected are correct On average need to select 400 variables to ensure inclusion of all 30 SCOOP l l 61% of top 30 selected are correct On average need to select 200 variables to ensure inclusion of all 30

Modifications ¡ Preserve common and group-distinct within group sample eigenvectors ¡ Regularize sample eigenvectors using Linear Perturbation Theory This is piecewise linear until adjacent eigenvalues become equal

Conclusions To the extent that something like an offset factor model holds, incorporating correlations may substantially improve selection of discriminating variables (DVs) ¡ Clustering of non-DVs does not seem to have any serious ill effect ¡ SCOOP is one way to use covariance structure efficiently ¡

References ¡ Bickel PJ and Levina E (2004). Some theory for Fisher's linear discriminant function, `naive Bayes', and some alternatives when there are many more variables than observations. Bernoulli 10, no. 6 989– 1010. ¡ Tibshirani R, Hastie T, Narasimhan & Chu (2002) Diagnosis of multiple cancer types by shrunken centroids of gene expression. PNAS 99, no. 10 6567 -6572. ¡ Sen, CK, Verducci, JS, Melfi, VF, Khanna, S, Barbacioru, C and Roy, S (2005). Post-reperfusion healing of the heart: Focus on oxygen-sensitive genes and DNA microarray as a tool. Mathematical Biosciences Institute Technical Report No. 31 (available at http: //mbi. osu. edu/publications/pub 2005. html)
- Slides: 28