Sharing Models How Can I Exchange Models SBML

Sharing Models

How Can I Exchange Models? SBML (Systems Biology Markup Language): de facto standard for representing cellular networks. A large number (>200) of tools support SBML. Cell. ML: Stores models in mathematical form, therefore is quite general, but biological information is lost. Not possible to reconstruct network. Less than a hand-full of tools support Cell. ML SBGN: A proposed standard for visually representing cellular networks. No persistent format has yet been devised which limits use in software. Matlab: Proprietary math based scripting language
SBML 3

Systems Biology Markup Language Originally developed in 2000 to allow users to exchange models between the small number of simulators that existed at that time. Since then it has become the de facto standard for model exchange in systems biology SBML represents models using XML by describing: 1. 2. 3. 4. 5. 6. Compartment Molecular Species Chemical and Enzymatic Reactions (including gene regulatory) Parameters Kinetic Rate Laws Additional Mathematical Equations when necessary 4

Systems Biology Markup Language <? xml version="1. 0" encoding="UTF-8"? > <!-- Created by XMLPretty. Printer on 7/30/2012 --> <sbml xmlns = "http: //www. sbml. org/sbml/level 2" level = "2" version = "1"> <model id = "cell"> <list. Of. Compartments> <compartment id = "compartment" size = "1"/> </list. Of. Compartments> <list. Of. Species> <species id = "S 1" boundary. Condition = "true" initial. Concentration = "1" compartment = "compartment"/> <species id = "S 3" boundary. Condition = "true" initial. Concentration = "0" compartment = "compartment"/> <species id = "S 2" boundary. Condition = "false" initial. Concentration = "1. 33" compartment = "compartment"/> </list. Of. Species> <list. Of. Parameters> <parameter id = "k 1" value = "3. 4"/> <parameter id = "k 2" value = "2. 3"/> </list. Of. Parameters> <list. Of. Reactions> <reaction id = "J 1" reversible = "false"> <list. Of. Reactants> <species. Reference species = "S 1" stoichiometry = "1"/> </list. Of. Reactants> <list. Of. Products> <species. Reference species = "S 2" stoichiometry = "1"/> </list. Of. Products> 5

Systems Biology Markup Language <kinetic. Law> <math xmlns = "http: //www. w 3. org/1998/Math. ML"> <apply> <times/> <ci> k 1 </ci> <ci> S 1 </ci> </apply> </math> </kinetic. Law> </reaction> <reaction id = "J 2" reversible = "false"> <list. Of. Reactants> <species. Reference species = "S 2" stoichiometry = "1"/> </list. Of. Reactants> <list. Of. Products> <species. Reference species = "S 3" stoichiometry = "1"/> </list. Of. Products> <kinetic. Law> <math xmlns = "http: //www. w 3. org/1998/Math. ML"> <apply> <times/> <ci> k 2 6

Model Repositories 421 Curated models as of July 2012 433 Non-curated Models. Biomodels. net At the EBI near Cambridge, UK 7

Parts Repository: Max Neals This tools decomposes all biomodels into their constituent parts. For example, search for pfk to locate all pfk parts in the biomodels database. See the following web site for details: http: //sites. google. com/site/semanticsofbiologicalprocesses/projects/sbmlrxnfinder
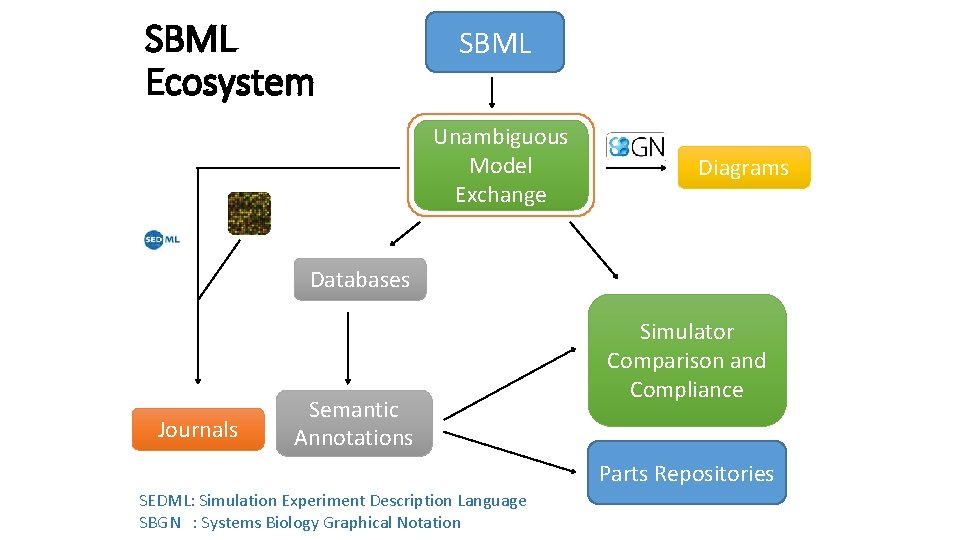
SBML Ecosystem SBML Unambiguous Model Exchange Diagrams Databases Journals Semantic Annotations Simulator Comparison and Compliance Parts Repositories SEDML: Simulation Experiment Description Language SBGN : Systems Biology Graphical Notation

Exporting/Importing Models Importing: 1. Antimony (using loada) 2. SBML (using roadrunner. Road. Runner (sbml model) Exporting: 1. r. get. Antimony() 2. r. get. SBML() 3. r. get. Matlab()

Exercise Build a simple model and export the SBML and Matlab
- Slides: 11