Sequencing Methods CAGE vs m RNASEQ Tru Seq







- Slides: 22

Sequencing Methods

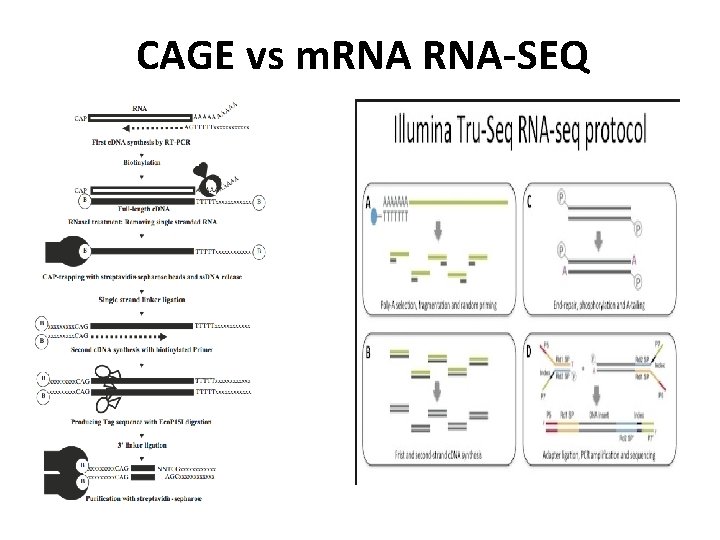
CAGE vs m. RNA-SEQ

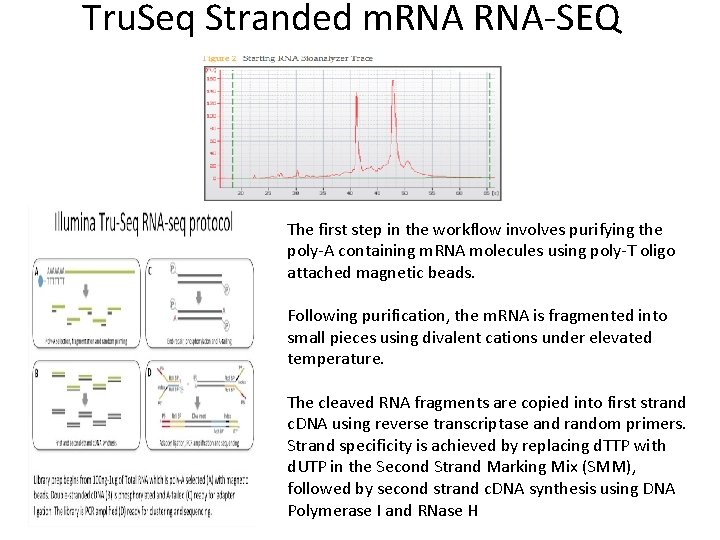
Tru. Seq Stranded m. RNA-SEQ The first step in the workflow involves purifying the poly-A containing m. RNA molecules using poly-T oligo attached magnetic beads. Following purification, the m. RNA is fragmented into small pieces using divalent cations under elevated temperature. The cleaved RNA fragments are copied into first strand c. DNA using reverse transcriptase and random primers. Strand specificity is achieved by replacing d. TTP with d. UTP in the Second Strand Marking Mix (SMM), followed by second strand c. DNA synthesis using DNA Polymerase I and RNase H

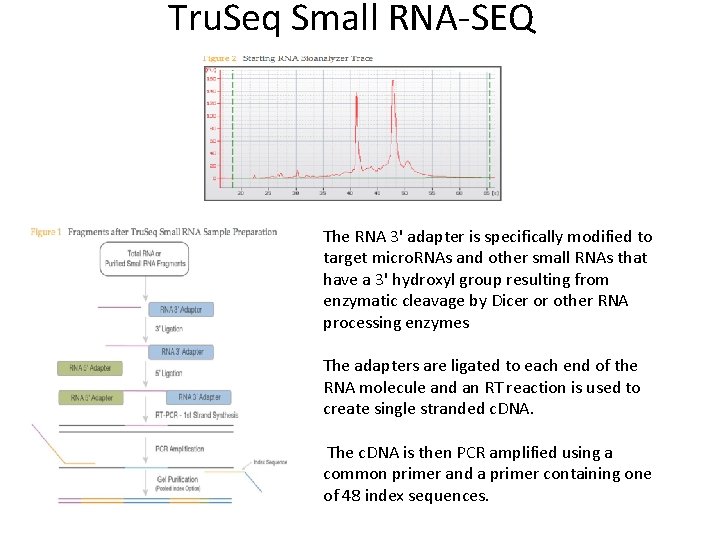
Tru. Seq Small RNA-SEQ The RNA 3' adapter is specifically modified to target micro. RNAs and other small RNAs that have a 3' hydroxyl group resulting from enzymatic cleavage by Dicer or other RNA processing enzymes The adapters are ligated to each end of the RNA molecule and an RT reaction is used to create single stranded c. DNA. The c. DNA is then PCR amplified using a common primer and a primer containing one of 48 index sequences.

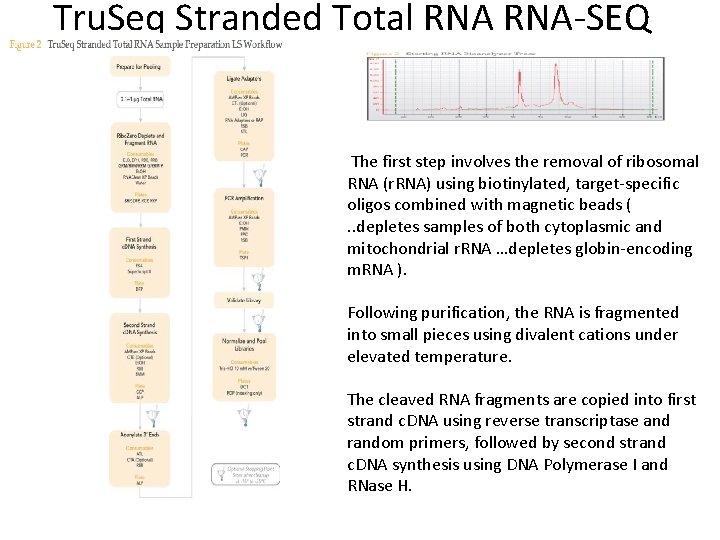
Tru. Seq Stranded Total RNA-SEQ The first step involves the removal of ribosomal RNA (r. RNA) using biotinylated, target-specific oligos combined with magnetic beads (. . depletes samples of both cytoplasmic and mitochondrial r. RNA …depletes globin-encoding m. RNA ). Following purification, the RNA is fragmented into small pieces using divalent cations under elevated temperature. The cleaved RNA fragments are copied into first strand c. DNA using reverse transcriptase and random primers, followed by second strand c. DNA synthesis using DNA Polymerase I and RNase H.

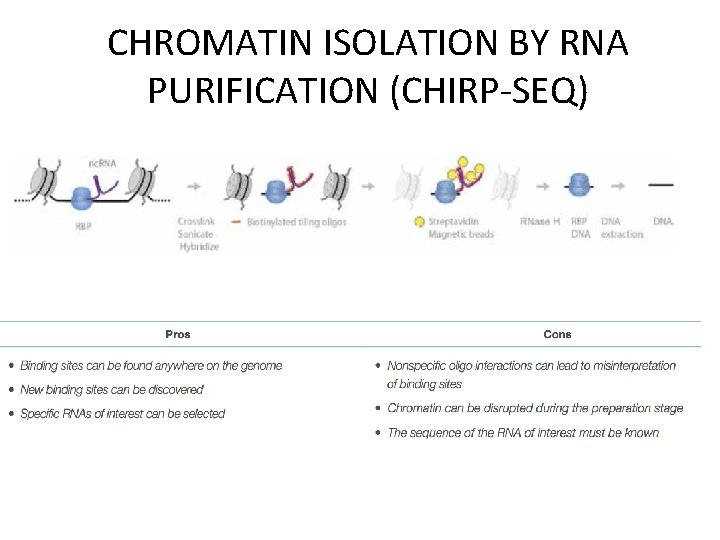
CHROMATIN ISOLATION BY RNA PURIFICATION (CHIRP-SEQ) A. Samples are first crosslinked and sonicated. B. Biotinylated tiling oligos are hybridized to the RNAs of interest. C. The complexes are captured with streptavidin magnetic beads. D. After treatment with RNase H the DNA is extracted and sequenced.

CHROMATIN ISOLATION BY RNA PURIFICATION (CHIRP-SEQ)

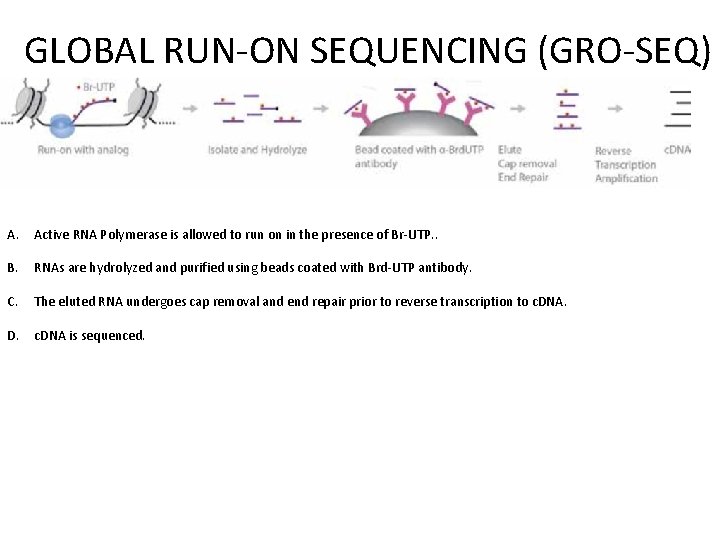
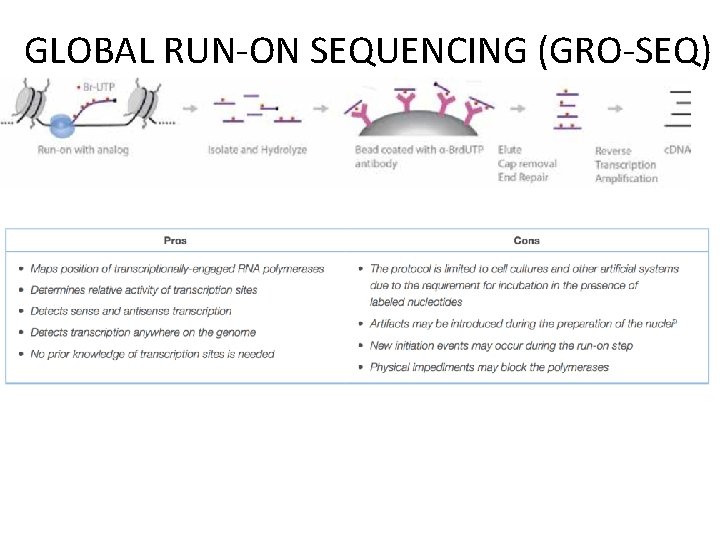
GLOBAL RUN-ON SEQUENCING (GRO-SEQ) A. Active RNA Polymerase is allowed to run on in the presence of Br-UTP. . B. RNAs are hydrolyzed and purified using beads coated with Brd-UTP antibody. C. The eluted RNA undergoes cap removal and end repair prior to reverse transcription to c. DNA. D. c. DNA is sequenced.

GLOBAL RUN-ON SEQUENCING (GRO-SEQ)

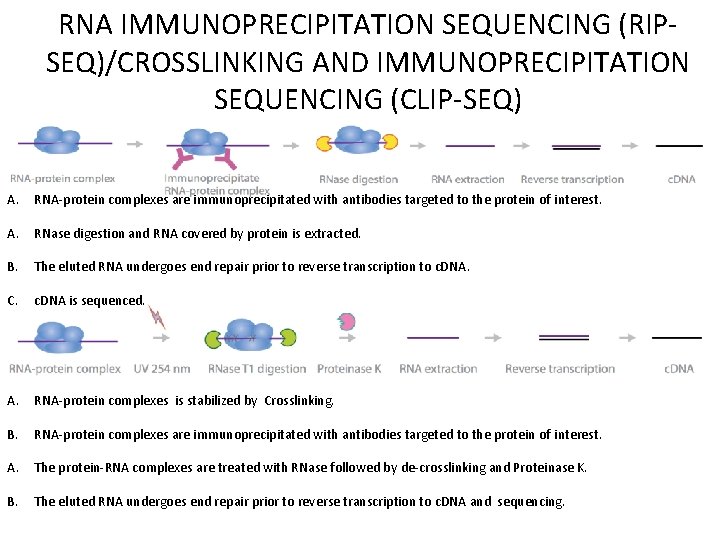
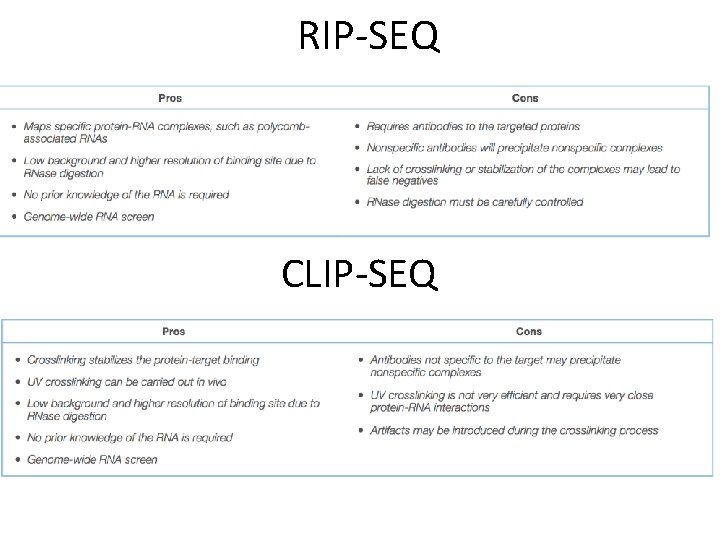
RNA IMMUNOPRECIPITATION SEQUENCING (RIPSEQ)/CROSSLINKING AND IMMUNOPRECIPITATION SEQUENCING (CLIP-SEQ) A. RNA-protein complexes are immunoprecipitated with antibodies targeted to the protein of interest. A. RNase digestion and RNA covered by protein is extracted. B. The eluted RNA undergoes end repair prior to reverse transcription to c. DNA. C. c. DNA is sequenced. A. RNA-protein complexes is stabilized by Crosslinking. B. RNA-protein complexes are immunoprecipitated with antibodies targeted to the protein of interest. A. The protein-RNA complexes are treated with RNase followed by de-crosslinking and Proteinase K. B. The eluted RNA undergoes end repair prior to reverse transcription to c. DNA and sequencing.

RIP-SEQ CLIP-SEQ


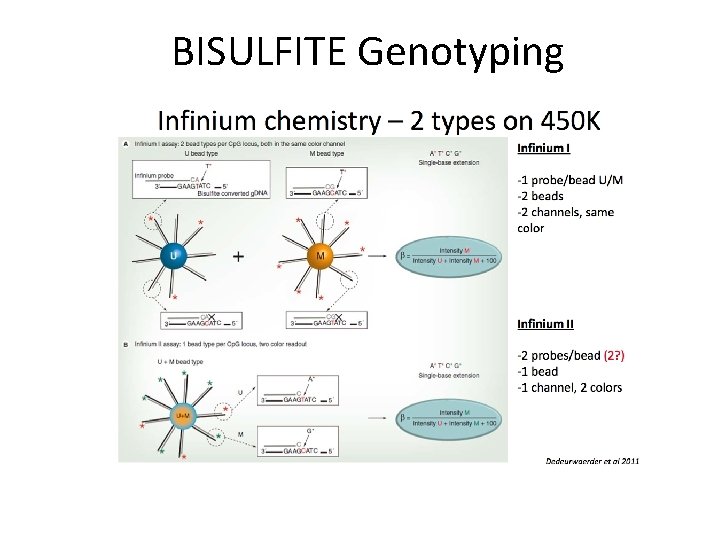
BISULFITE Genotyping

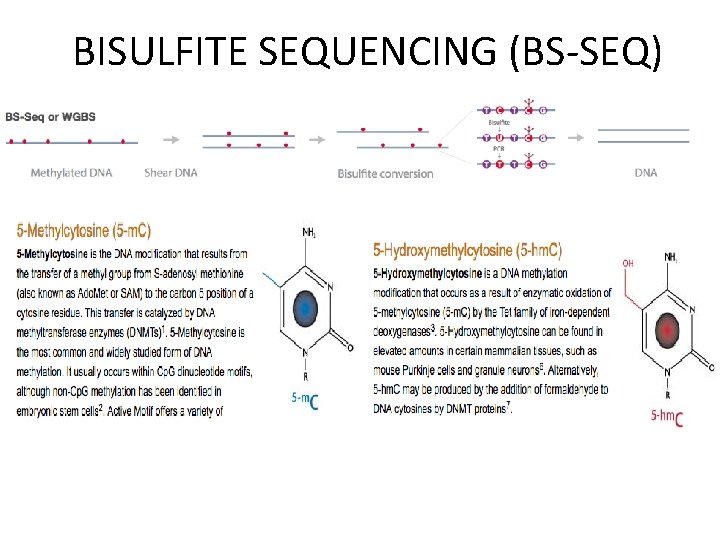
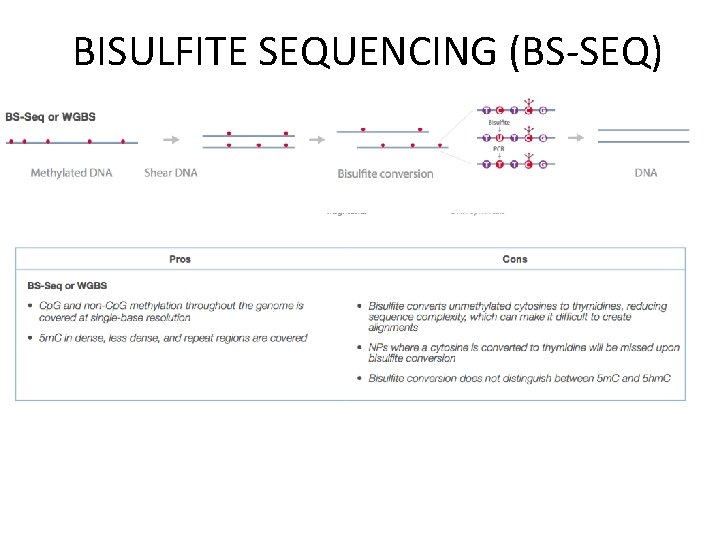
BISULFITE SEQUENCING (BS-SEQ)

BISULFITE SEQUENCING (BS-SEQ)

METHYLATED DNA IMMUNOPRECIPITATION SEQUENCING (MEDIP-SEQ)

METHYLATED DNA IMMUNOPRECIPITATION SEQUENCING (MEDIP-SEQ)

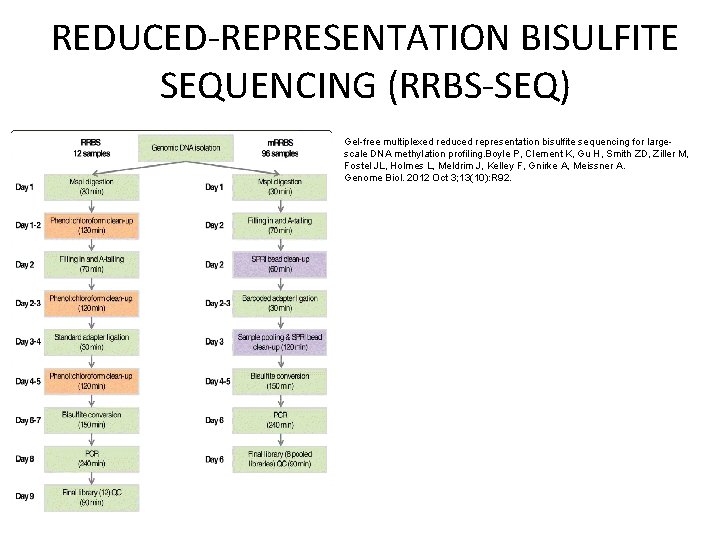
REDUCED-REPRESENTATION BISULFITE SEQUENCING (RRBS-SEQ) Gel-free multiplexed reduced representation bisulfite sequencing for largescale DNA methylation profiling. Boyle P, Clement K, Gu H, Smith ZD, Ziller M, Fostel JL, Holmes L, Meldrim J, Kelley F, Gnirke A, Meissner A. Genome Biol. 2012 Oct 3; 13(10): R 92.

REDUCED-REPRESENTATION BISULFITE SEQUENCING (RRBS-SEQ)

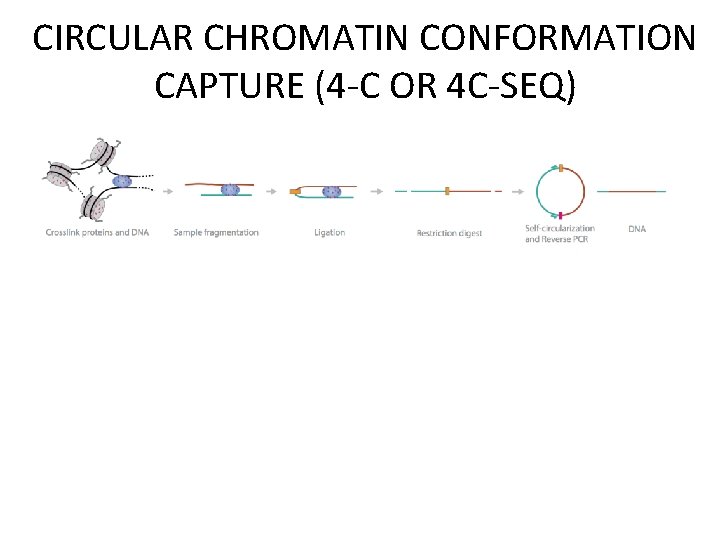
CIRCULAR CHROMATIN CONFORMATION CAPTURE (4 -C OR 4 C-SEQ)

Overview of 3 C-derived methods. de Wit E , and de Laat W Genes Dev. 2012; 26: 11 -24

CIRCULAR CHROMATIN CONFORMATION CAPTURE (4 -C OR 4 C-SEQ)