Sequence Based Analysis Tutorial March 26 2004 NIH

Sequence Based Analysis Tutorial March 26, 2004 NIH Proteomics Workshop Lai-Su L. Yeh, Ph. D. Protein Science Team Lead Protein Information Resource at Georgetown University Medical Center

Retrieval, Sequence Search & Classification Methods Ø Retrieve protein info by text / UID Ø Sequence Similarity Search l BLAST, FASTA, Dynamic Programming Ø Family Classification l Patterns, Profiles, Hidden Markov Models, Sequence Alignments, Neural Networks Ø Integrated Search and Classification System 2

Sequence Similarity Search Based on Pair-Wise Comparisons Ø Dynamic Programming Algorithms Ø l l Ø Global Similarity: Needleman-Wunch Local Similarity: Smith-Waterman Heuristic Algorithms l l l FASTA: Based on K-Tuples (2 -Amino Acid) BLAST: Triples of Conserved Amino Acids Gapped-BLAST: Allow Gaps in Segment Pairs PHI-BLAST: Pattern-Hit Initiated Search PSI-BLAST: Position-Specific Iterated Search 3

Sequence Similarity Search Ø Similarity Search Parameters l l Ø Scoring Matrices – Based on Conserved Amino Acid Substitution • Dayhoff Mutation Matrix, e. g. , PAM 250 (~20% Identity) • Henikoff Matrix from Ungapped Alignments, e. g. , BLOSUM 62 Gap Penalty Search Time Comparisons l l l Smith-Waterman: 10 Min FASTA: 2 Min BLAST: 20 Sec 4

Feature Representation Features: Residue Physicochemical Properties, Context (Local & Global) Features, Evolutionary Features Ø Alternative Alphabets: Classification of Amino Acids To Capture Different Features of Amino Acid Residues Ø 5

Substitution Matrix Ø Ø Likelihood of One Amino Acid Mutated into Another Over Evolutionary Time Negative Score: Unlikely to Happen (e. g. , Gly/Trp, -7) Positive Score: Conservative Substitution (e. g. , Lys/Arg, +3) High Score for Identical Matches: Rare Amino Acids (e. g. , Trp, Cys) 6

BLAST (Basic Local Alignment Search Tool) Ø To search a sequence against the database Ø Extremely fast Ø Robust Ø Most widely used It finds very short segment pairs between the query and sequence in the database These segments are then extended in both directions until the maximum possible score of this particular segment is reached 7
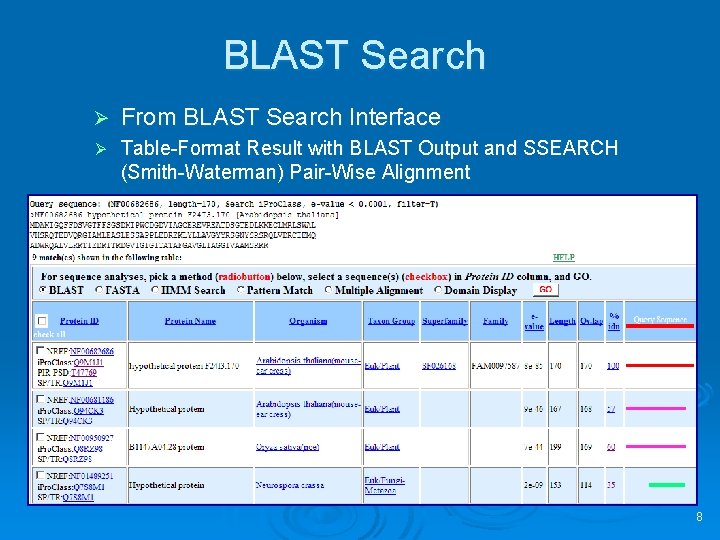
BLAST Search Ø From BLAST Search Interface Ø Table-Format Result with BLAST Output and SSEARCH (Smith-Waterman) Pair-Wise Alignment 8
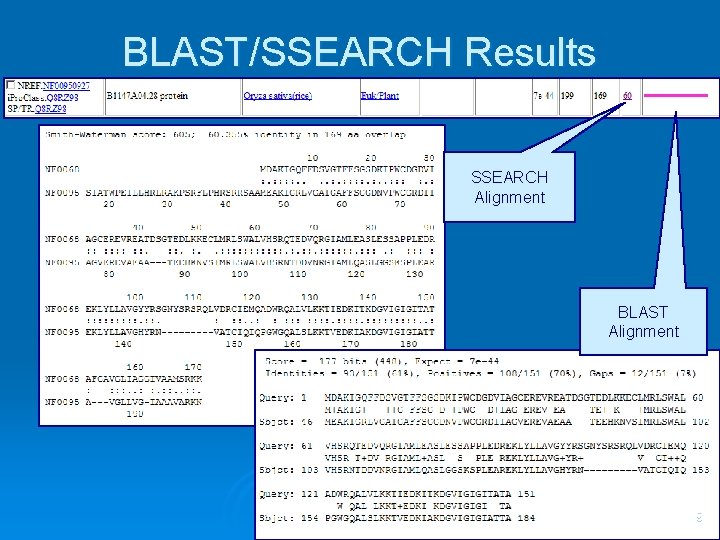
BLAST/SSEARCH Results SSEARCH Alignment BLAST Alignment 9

Family Classification Methods Based on Family Information Ø Clustal. W Multiple Sequence Alignment Ø Pro. Site Pattern Search Ø Profile Search Ø Hidden Markov Models (HMMs) Ø Neural Networks Ø Integrated Analysis Ø 10

Multiple Sequence Alignment Ø Clustal. W Progressive Pairwise Approach l Base on Exhaustive Pairwise Alignments Ø Neighbor Joining l Joining Order Corresponding to a Tree Ø Alignment Varies l Dependent on Joining Order Ø 11

How do you build a tree? Ø Pick sequences to align Ø Align them Ø Verify the alignment Ø Keep the parts that are aligned correctly Ø Build and evaluate a phylogenetic tree 12

Multiple Alignment and Tree Ø From Text/Sequence Search Result or Clustal. W Alignment Interface 13
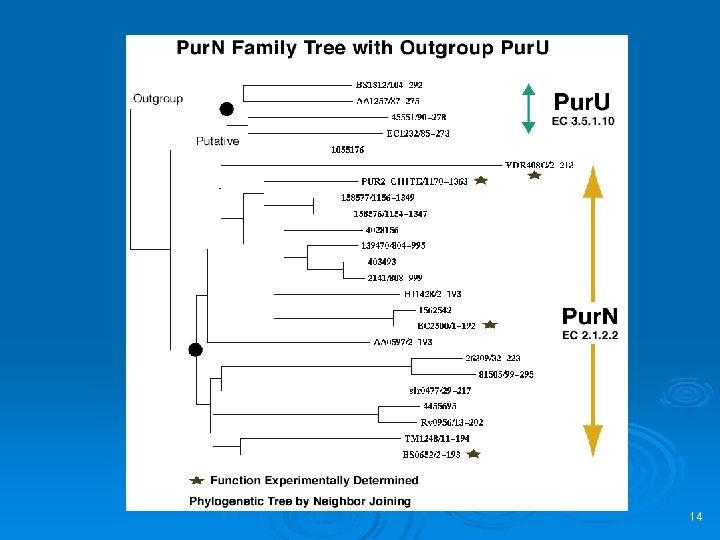
14

Motif Patterns (Regular Expressions) Ø Signature Patterns for Functional Motifs Pro. Class Motif Alignments 15

PIR Pattern Search From Text/Sequence Search Result or Pattern Search Interface Ø One Query Sequence Against PROSITE Pattern Database Ø One Query Pattern (PROSITE or User-Defined) Against Sequence DB Ø 16

Pattern Search Result (I) Ø One Query Sequence Against PROSITE Pattern Database 17
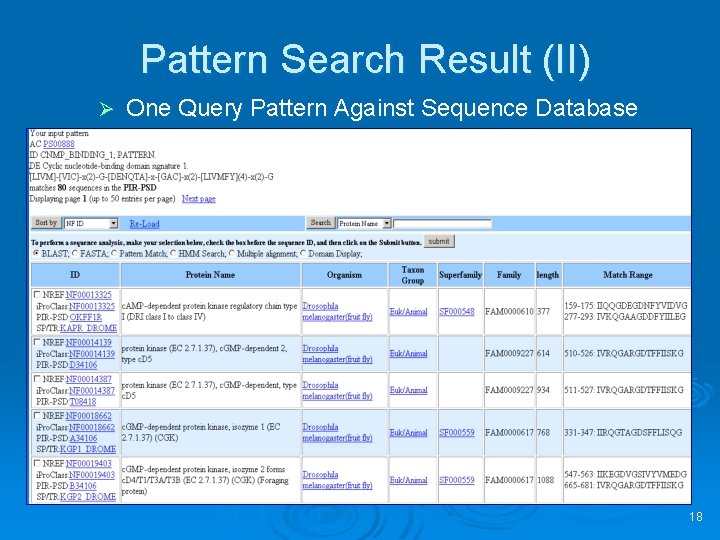
Pattern Search Result (II) Ø One Query Pattern Against Sequence Database 18

Profile Method Profile: A Table of Scores to Express Family Consensus Derived from Multiple Sequence Alignments l Num of Rows = Num of Aligned Positions l Each row contains a score for the alignment with each possible residue. Ø Profile Searching l Summation of Scores for Each Amino Acid Residue along Query Sequence l Higher Match Values at Conserved Positions Ø 19

PIR HMM Domain/Motif Search Ø From Text/Sequence Search Result or HMM Search Interface Ø HMMER Model Building & Sequence Search Ø Search One Query Protein Against All HMMs Ø Search One HMM Against Sequence DB 20

HMM Search Result (I) Ø One Query Protein Against All Pfam HMMs 21
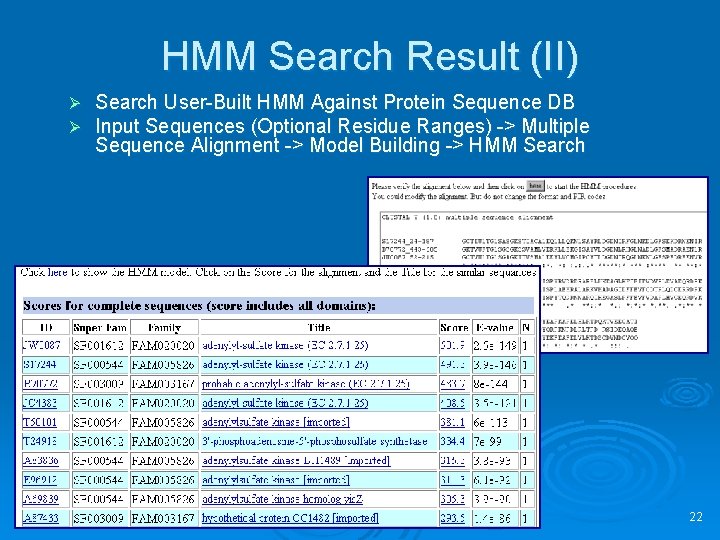
HMM Search Result (II) Ø Ø Search User-Built HMM Against Protein Sequence DB Input Sequences (Optional Residue Ranges) -> Multiple Sequence Alignment -> Model Building -> HMM Search 22

Secondary Structure Features a Helix Patterns of Hydrophobic Residue Conservation Showing I, I+3, I+4, I+7 Pattern Are Highly Indicative of an a Helix (Amphipathic) Ø b Strands That Are Half Buried in the Protein Core Will Tend to Have Hydrophobic Residues at Positions I, I+2, I+4, I+6 Ø 23

Integrated Bioinformatics System for Function and Pathway Discovery Data Integration Ø Associative Analysis Ø 24

Query Sequence PIR-NREF i. Pro. Class Family Classification & Functional Analysis BLAST Search HMM Domain Search Analytical Pipeline Top-Matched Superfamilies/Domains HMM Motif Search Pattern Search Signal. P/TMHMM Predicated Superfamilies/Domains/Motifs/Sites/Signal. Peptides/TMHs SSEARCH CLUSTALW Superfamily/Domain/Motif Alignments Family Relationships & Functional Features 25

Integrated Bioinformatics System Ø Global Bioinformatics Analysis of 1000’s of Genes and Proteins Ø Pathway Discovery, Target Identification 26

27

Lab Section 28
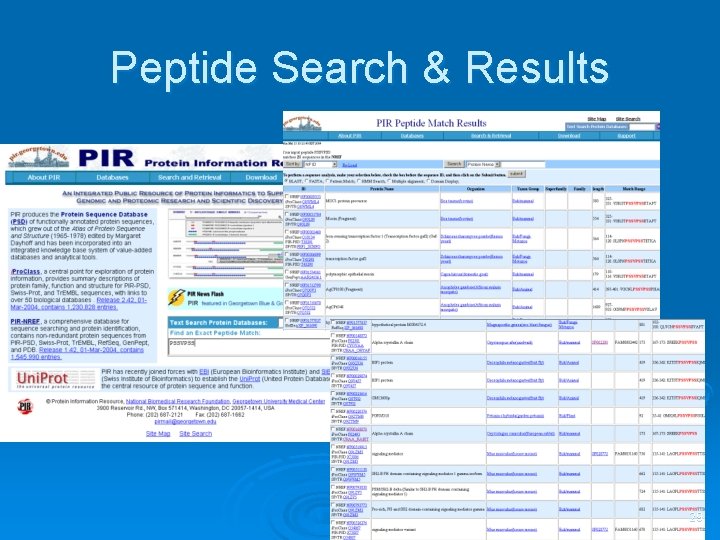
Peptide Search & Results 29
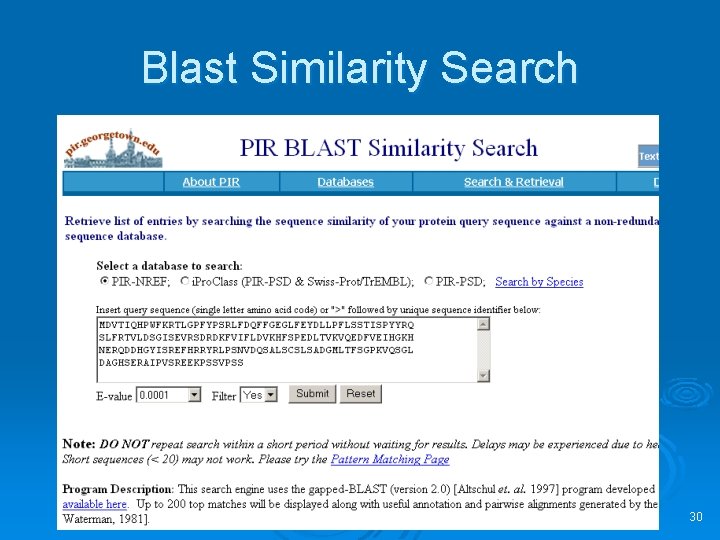
Blast Similarity Search 30
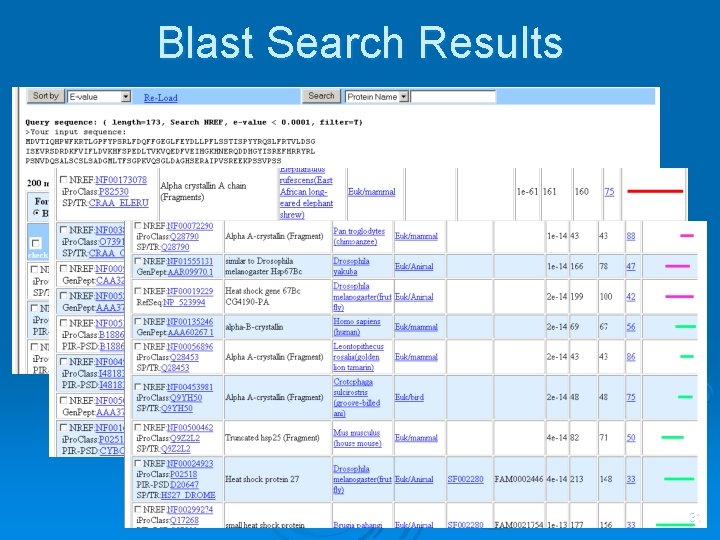
Blast Search Results 31

Pair-Wise Alignment 32
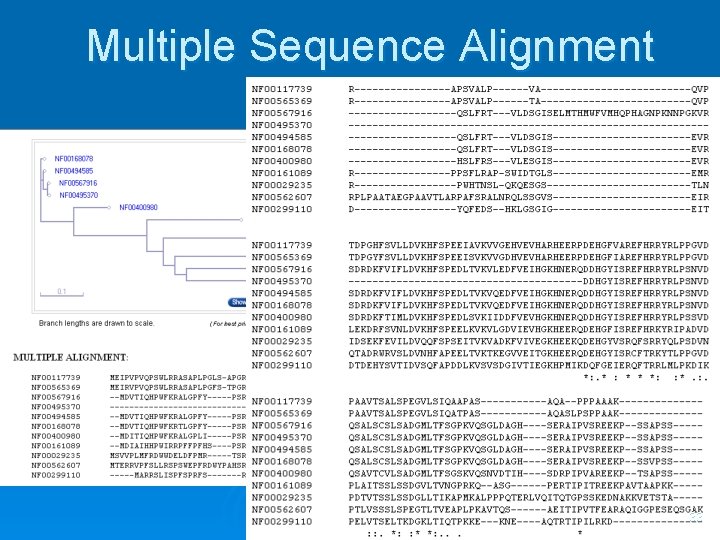
Multiple Sequence Alignment 33

Pattern Search Results 34

HMM Domain Search Result 35
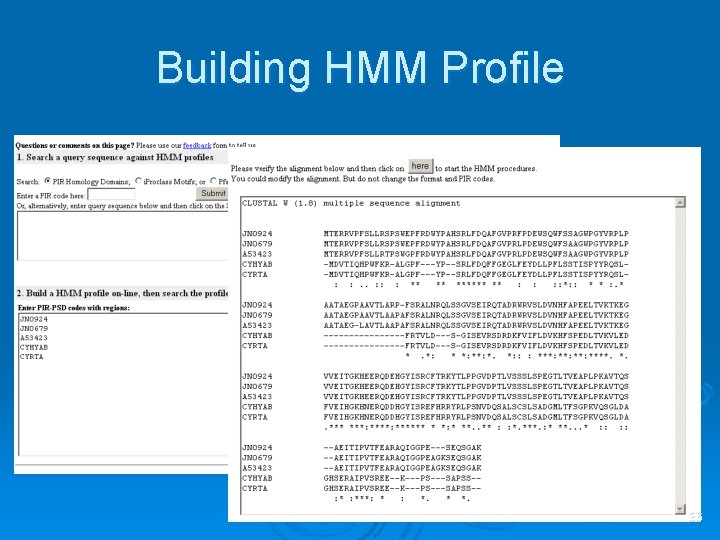
Building HMM Profile 36
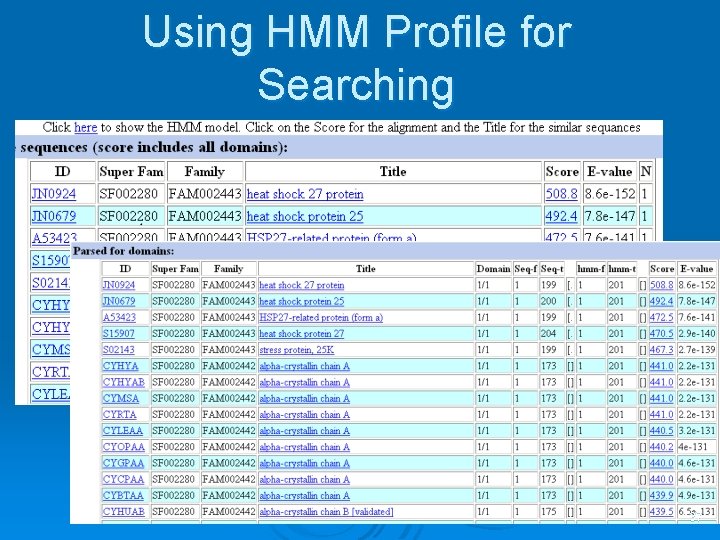
Using HMM Profile for Searching 37
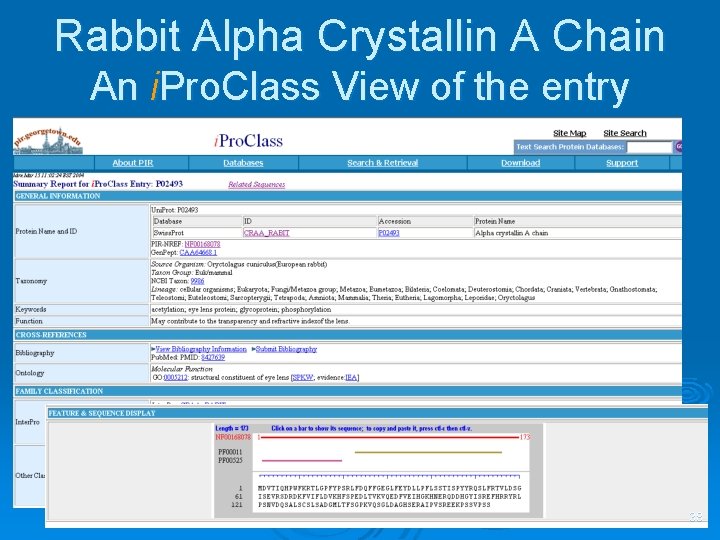
Rabbit Alpha Crystallin A Chain An i. Pro. Class View of the entry 38
- Slides: 38