Se deben administrar las terapias siempre personalizadas en

¿Se deben administrar las terapias siempre personalizadas en cáncer de pulmón? : A FAVOR Enriqueta Felip Hospital Vall d’Hebron; Barcelona

DONDE ESTAMOS CON EL TRATAMIENTO CON “TARGETED THERAPIES”? Estos agentes tienen actividad en subgrupos de pacientes; pero ninguno tiene beneficio universal

Fase III positivos en 1 línea con “targeted therapies” en población no seleccionada • ECOG carbo/pac/beva vs carbo/pac ü PFS 6, 2 vs 4, 5 meses; HR 0. 66, p<0. 001 ü MS 12, 3 vs 10, 3; HR 0. 79, p<0. 003 • AVAIL cis/gem/beva 15 vs cis/gem/beva 7. 5 vs cis/gem/placebo ü PFS 6, 6 vs 6, 8 vs 6, 2 meses; HR 0. 85/p=0. 0456; HR 0. 75/p=0. 0003 ü MS 13, 4 vs 13, 6 vs 13, 1 meses; HR 1. 03/p=0. 76; HR 0. 93/p=0. 42 • FLEX cis/vin/cetuximab vs cis/vin (EGFR IH +) ü PFS 4, 8 vs 4, 8 meses; HR 0. 943, p=NS ü MS 11, 3 vs 10, 1 meses; HR 0. 871, p=0. 044

Fase III positivos en 2 -3 línea con “targeted therapies” en población no seleccionada • BR. 21 erlotinib vs placebo ü Mediana de supervivencia 6, 7 vs 4, 7 meses; HR 0. 72; p=0. 001 • INTEREST gefitinib vs docetaxel ü Mediana de supervivencia 7, 8 vs 8. 0 meses; HR=1. 02

Cronología interpretación mutaciones de EGFR • 2004 mutaciones de EGFR; predictivas • 2005 número copias del gen predictivo; mutaciones no • 2007 mutaciones de EGFR pronosticas no predictivas • 2008 mutaciones de EGFR predictivas de supervivencia libre de progresión
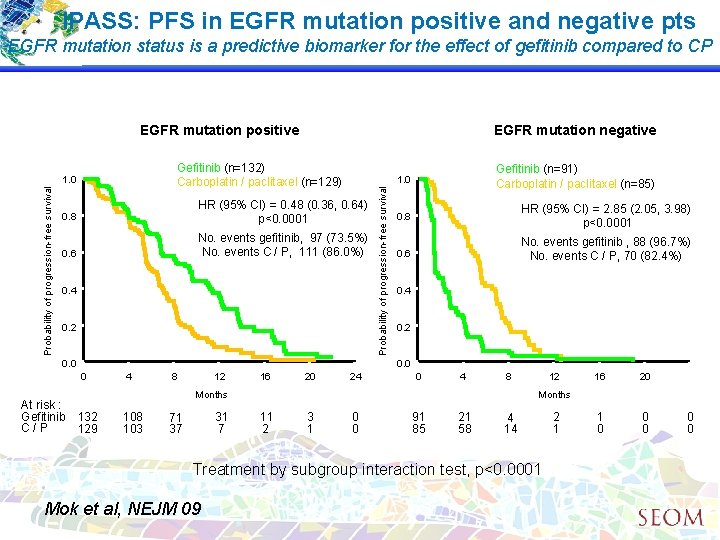
IPASS: PFS in EGFR mutation positive and negative pts EGFR mutation status is a predictive biomarker for the effect of gefitinib compared to CP EGFR mutation positive Gefitinib (n=132) Carboplatin / paclitaxel (n=129) 0. 8 HR (95% CI) = 0. 48 (0. 36, 0. 64) p<0. 0001 0. 6 No. events gefitinib, 97 (73. 5%) No. events C / P, 111 (86. 0%) 0. 4 0. 2 0. 0 0. 8 HR (95% CI) = 2. 85 (2. 05, 3. 98) p<0. 0001 0. 6 No. events gefitinib , 88 (96. 7%) No. events C / P, 70 (82. 4%) 0. 4 0. 2 0. 0 0 At risk : Gefitinib C/P Gefitinib (n=91) Carboplatin / paclitaxel (n=85) 1. 0 Probability of progression-free survival EGFR mutation negative 4 8 12 16 20 24 0 4 8 Months 132 129 108 103 31 7 71 37 12 20 1 0 0 0 Months 11 2 3 1 0 0 91 85 21 58 4 14 Treatment by subgroup interaction test, p<0. 0001 Mok et al, NEJM 09 16 2 1 0 0
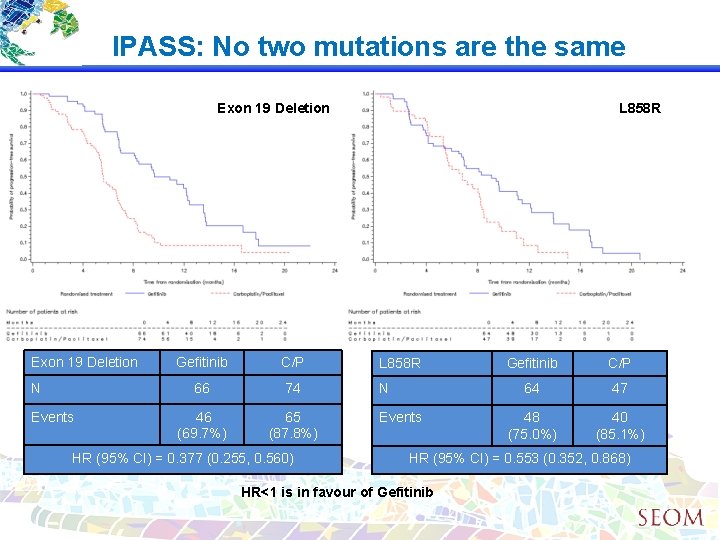
IPASS: No two mutations are the same Exon 19 Deletion N Events L 858 R Gefitinib C/P L 858 R 66 74 N 46 (69. 7%) 65 (87. 8%) HR (95% CI) = 0. 377 (0. 255, 0. 560) Events Gefitinib C/P 64 47 48 (75. 0%) 40 (85. 1%) HR (95% CI) = 0. 553 (0. 352, 0. 868) HR<1 is in favour of Gefitinib
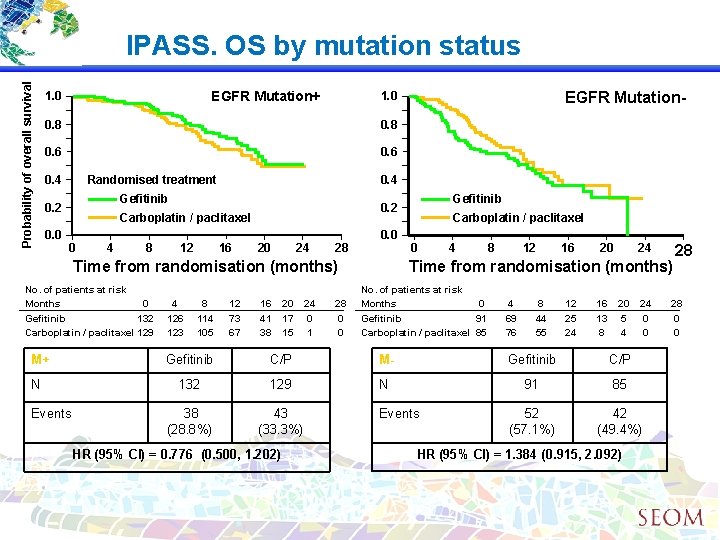
Probability of overall survival IPASS. OS by mutation status EGFR Mutation+ 1. 0 0. 8 0. 6 0. 4 0. 0 0. 4 Randomised treatment Gefitinib 0. 2 4 8 12 16 20 24 28 0. 0 Time from randomisation (months) No. of patients at risk Months 0 Gefitinib 132 Carboplatin / paclitaxel 129 M+ N Events 4 126 123 8 114 105 12 73 67 Gefitinib 0. 2 Carboplatin / paclitaxel 0 EGFR Mutation- 1. 0 16 41 38 20 17 15 24 0 1 28 0 0 Carboplatin / paclitaxel 0 4 8 12 16 20 24 Time from randomisation (months) No. of patients at risk Months 0 Gefitinib 91 Carboplatin / paclitaxel 85 4 69 76 8 44 55 12 25 24 16 13 8 20 5 4 24 0 0 Gefitinib C/P M- Gefitinib C/P 132 129 N 91 85 38 (28. 8%) 43 (33. 3%) 52 (57. 1%) 42 (49. 4%) HR (95% CI) = 0. 776 (0. 500, 1. 202) Events HR (95% CI) = 1. 384 (0. 915, 2. 092) 28 28 0 0
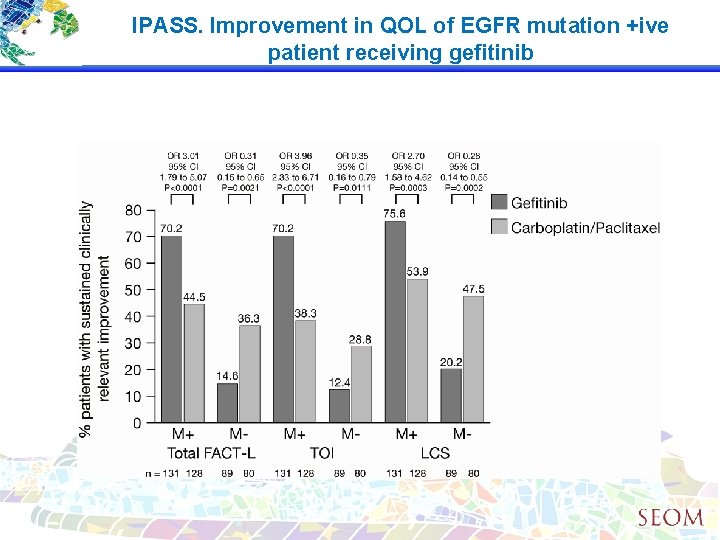
IPASS. Improvement in QOL of EGFR mutation +ive patient receiving gefitinib
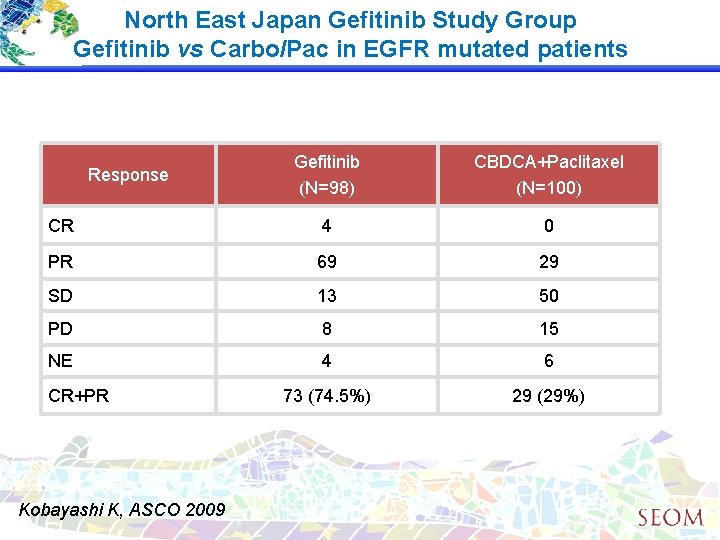
North East Japan Gefitinib Study Group Gefitinib vs Carbo/Pac in EGFR mutated patients Gefitinib (N=98) CBDCA+Paclitaxel (N=100) CR 4 0 PR 69 29 SD 13 50 PD 8 15 NE 4 6 73 (74. 5%) 29 (29%) Response CR+PR Kobayashi K, ASCO 2009
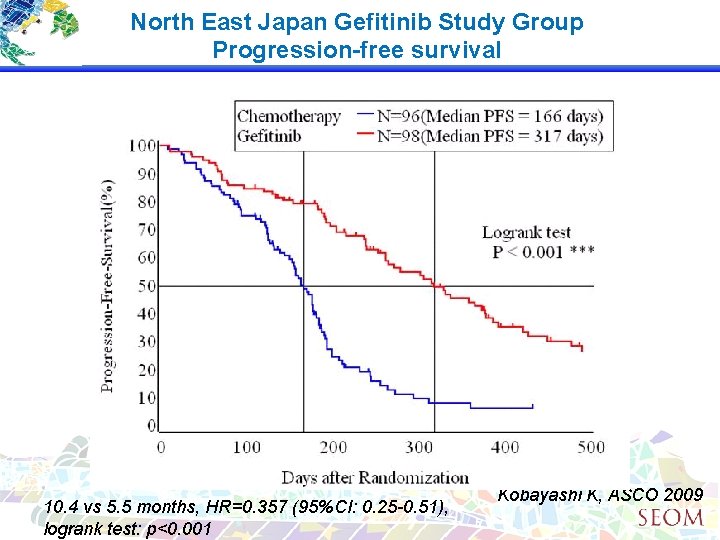
North East Japan Gefitinib Study Group Progression-free survival 10. 4 vs 5. 5 months, HR=0. 357 (95%CI: 0. 25 -0. 51), logrank test: p<0. 001 Kobayashi K, ASCO 2009

Screening for EGFR mutations in lung cancer (Rosell et al, NEJM 09) • Analizan mutaciones EGFR en 2105 pacientes de 129 centros (2005 -08) ü 350 mutaciones (16, 6%) ü > frecuente en mujeres (69, 7%) y no fumadores (66, 6%) ü Deleciones exon 19, 62% / mutaciones L 858 R, 38% • 217 pacientes recibieron erlotinib ü PFS 14 meses; MS 27 meses
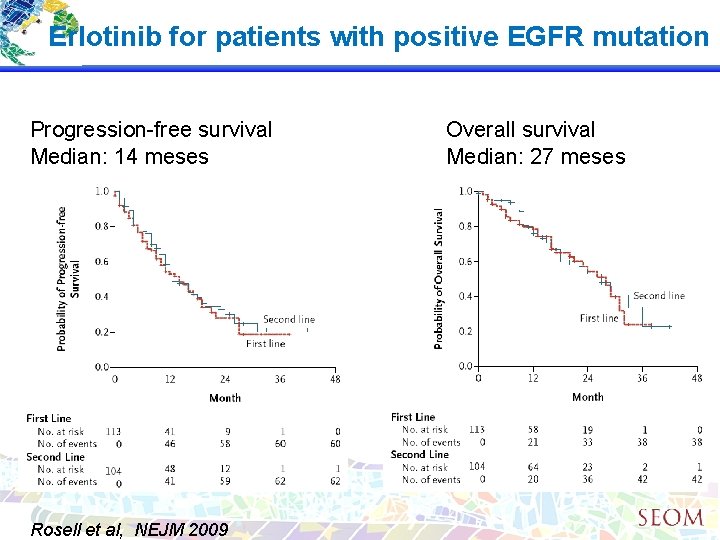
Erlotinib for patients with positive EGFR mutation Progression-free survival Median: 14 meses Rosell et al, NEJM 2009 Overall survival Median: 27 meses
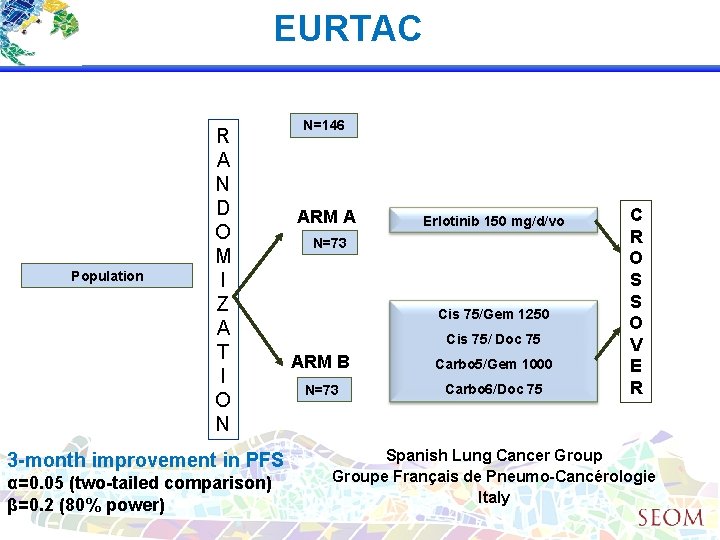
EURTAC Population R A N D O M I Z A T I O N 3 -month improvement in PFS α=0. 05 (two-tailed comparison) β=0. 2 (80% power) N=146 ARM A Erlotinib 150 mg/d/vo N=73 Cis 75/Gem 1250 Cis 75/ Doc 75 ARM B Carbo 5/Gem 1000 N=73 Carbo 6/Doc 75 C R O S S O V E R Spanish Lung Cancer Groupe Français de Pneumo-Cancérologie Italy
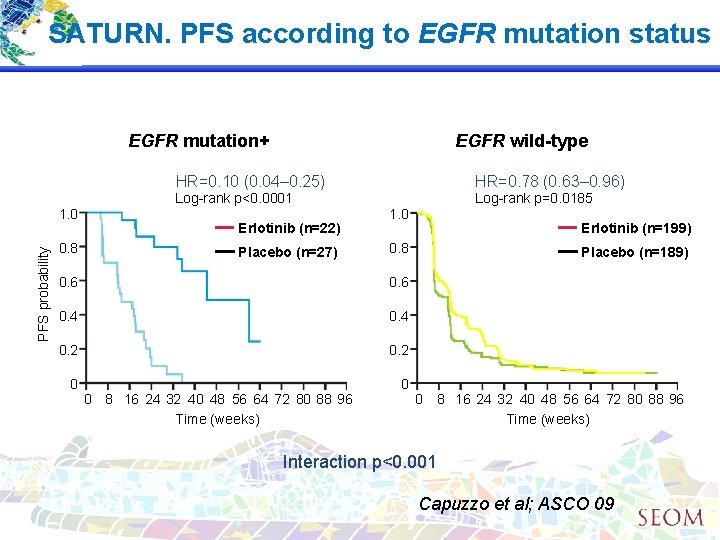
SATURN. PFS according to EGFR mutation status EGFR mutation+ PFS probability 1. 0 0. 8 EGFR wild-type HR=0. 10 (0. 04– 0. 25) HR=0. 78 (0. 63– 0. 96) Log-rank p<0. 0001 Log-rank p=0. 0185 Erlotinib (n=22) Placebo (n=27) 1. 0 0. 8 0. 6 0. 4 0. 2 0 0 8 16 24 32 40 48 56 64 72 80 88 96 Time (weeks) Erlotinib (n=199) 0 Placebo (n=189) 0 8 16 24 32 40 48 56 64 72 80 88 96 Time (weeks) Interaction p<0. 001 Capuzzo et al; ASCO 09
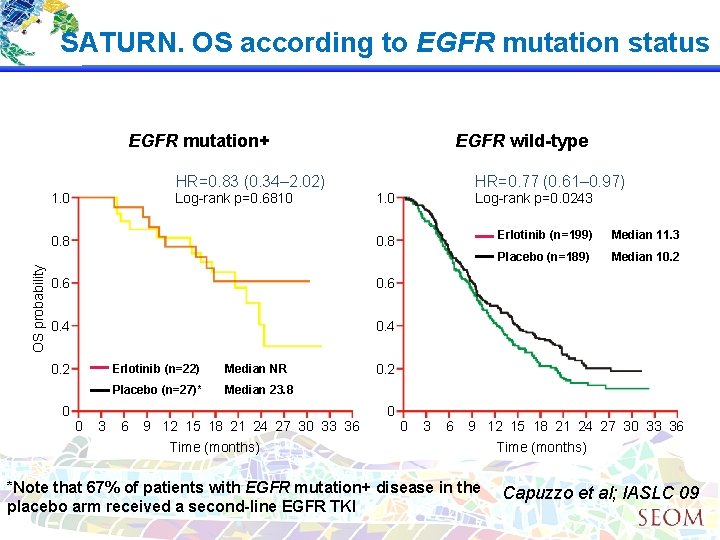
SATURN. OS according to EGFR mutation status EGFR mutation+ HR=0. 83 (0. 34– 2. 02) 1. 0 Log-rank p=0. 6810 OS probability 0. 8 EGFR wild-type 1. 0 Log-rank p=0. 0243 0. 8 0. 6 0. 4 0. 2 0 HR=0. 77 (0. 61– 0. 97) 0 3 Erlotinib (n=22) Median NR Placebo (n=27)* Median 23. 8 6 9 12 15 18 21 24 27 30 33 36 Time (months) Erlotinib (n=199) Median 11. 3 Placebo (n=189) Median 10. 2 0 0 3 6 9 12 15 18 21 24 27 30 33 36 Time (months) *Note that 67% of patients with EGFR mutation+ disease in the placebo arm received a second-line EGFR TKI Capuzzo et al; IASLC 09

EGFR TKIs works well on EGFR mutation positive patients: Be it first or second line
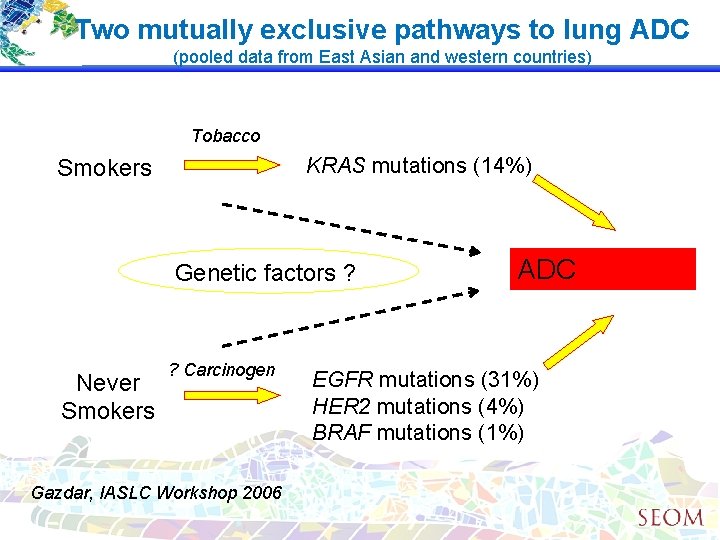
Two mutually exclusive pathways to lung ADC (pooled data from East Asian and western countries) Tobacco KRAS mutations (14%) Smokers Genetic factors ? Never Smokers ? Carcinogen Gazdar, IASLC Workshop 2006 ADC EGFR mutations (31%) HER 2 mutations (4%) BRAF mutations (1%)

Mutaciones de KRAS en ADC de pulmón y beneficio a EGFR TKIs • Mutaciones de KRAS; 20 -30% de ADC, sobretodo en fumadores • Resultados NO concluyentes, pero: ü En KRAS mutado; respuesta a EGFR TKIs < 1% ü Meta-análisis de estudios publicados (Linardou et al; Lancet Oncol 08); mutación de KRAS se asocia a ausencia de respuesta a TKIs ü Evidencia insuficiente para confirmar que mutaciones de KRAS no son una contraindicación a cetuximab en pacientes con ADC (FLEX: KRAS analizado en 395 muestras / 75 mutaciones)

Características clínicas de pacientes con NSCLC con EML 4 -ALK + (Shaw AT, JCO 09) • EMLK 4 -ALK; proteína tyrosine kinasa de fusión • “Screening” EMLK 4 -ALK / EGFR si > 2 características: mujer, etnia asiática, no fumador, ADC ü 141 analizados, 13% EMLK 4 -ALK + / 22% mutaciones EGFR ü EMLK 4 -ALK +: − Excluyente con mutaciones de EGFR / KRAS − Pacientes más jóvenes y frecuentemente hombres − Frecuentemente no fumadores − Asociado a resistencia a EGFR TKIs • EMLK 4 -ALK +, subgrupo molecular de NSCLC diferente

Estudio fase I PF-02341066 (Kwak et al ASCO 09) • PF-02341066 inhibidor oral de c-MET y receptor tyrosine kinasa ALK • Cohorte enriquecida en fase I; 29 pacientes con NSCLC con translocación EML 4 -ALK en tumor (76% no fumadores / 90% ADC): – Respuesta parcial 59% pacientes – Control de la enfermedad 83% pacientes
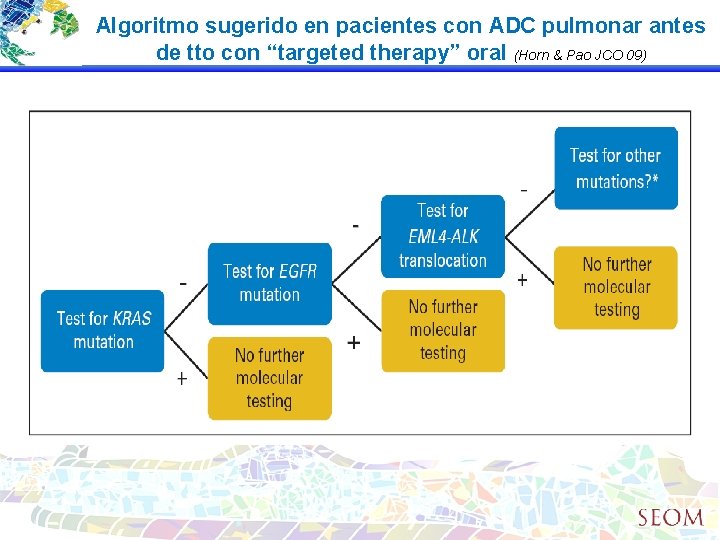
Algoritmo sugerido en pacientes con ADC pulmonar antes de tto con “targeted therapy” oral (Horn & Pao JCO 09)
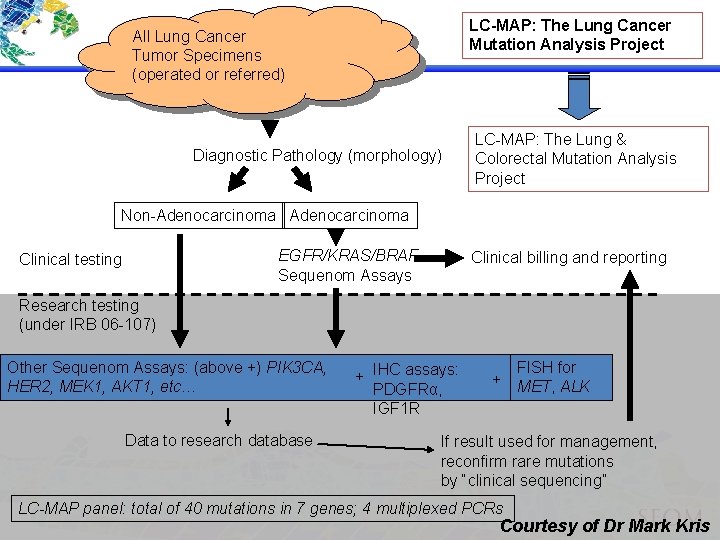
LC-MAP: The Lung Cancer Mutation Analysis Project All Lung Cancer Tumor Specimens (operated or referred) Diagnostic Pathology (morphology) LC-MAP: The Lung & Colorectal Mutation Analysis Project Non-Adenocarcinoma EGFR/KRAS/BRAF Sequenom Assays Clinical testing Clinical billing and reporting Research testing (under IRB 06 -107) Other Sequenom Assays: (above +) PIK 3 CA, HER 2, MEK 1, AKT 1, etc… Data to research database + IHC assays: PDGFRα, IGF 1 R FISH for + MET, ALK If result used for management, reconfirm rare mutations by “clinical sequencing” LC-MAP panel: total of 40 mutations in 7 genes; 4 multiplexed PCRs Courtesy of Dr Mark Kris

Genetic profiling in ADC at MSKCC Courtesy of Dr Mark Kris Routine use of molecular characteristics to select therapies for patients with ADC EGFR Mutation Use erlotinib KRAS Mutation Do not use of erlotinib Do not use cisplatin-based adjuvant therapy HER 2 Mutation or Amplification Use trastuzumab Molecular characteristics used to select patients for protocols at MSK EGFR BIBW 2992 Erlotinib (Adjuvant) KRAS Salirasib - Deforolimus EML 4 -ALK MET Amplification PF-02341066 BRAF AZ 6244
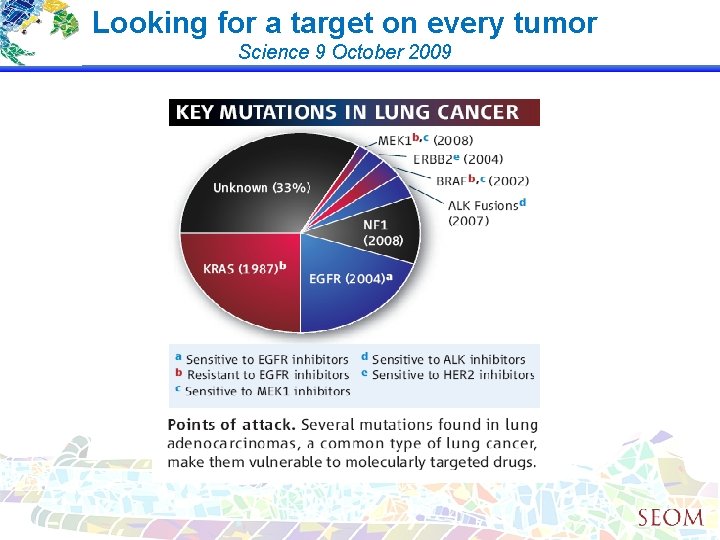
Looking for a target on every tumor Science 9 October 2009
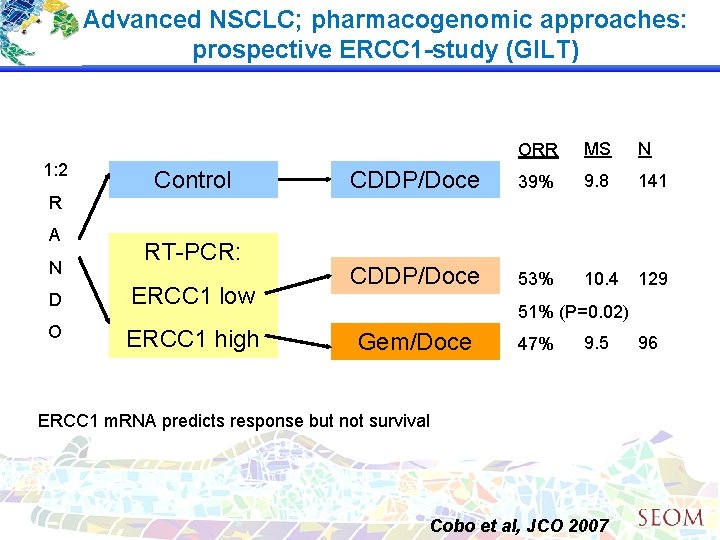
Advanced NSCLC; pharmacogenomic approaches: prospective ERCC 1 -study (GILT) 1: 2 R A N Control RT-PCR: D ERCC 1 low O ERCC 1 high ORR MS N CDDP/Doce 39% 9. 8 141 CDDP/Doce 53% 10. 4 129 51% (P=0. 02) Gem/Doce 47% 9. 5 ERCC 1 m. RNA predicts response but not survival Cobo et al, JCO 2007 96
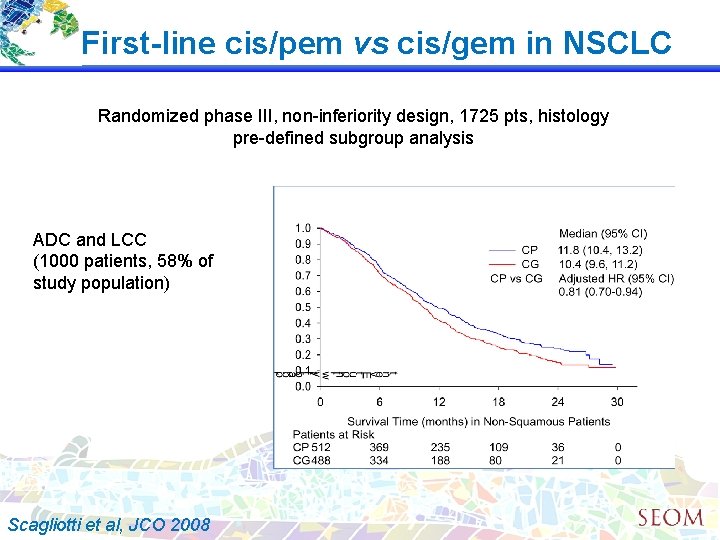
First-line cis/pem vs cis/gem in NSCLC Randomized phase III, non-inferiority design, 1725 pts, histology pre-defined subgroup analysis ADC and LCC (1000 patients, 58% of study population) Scagliotti et al, JCO 2008
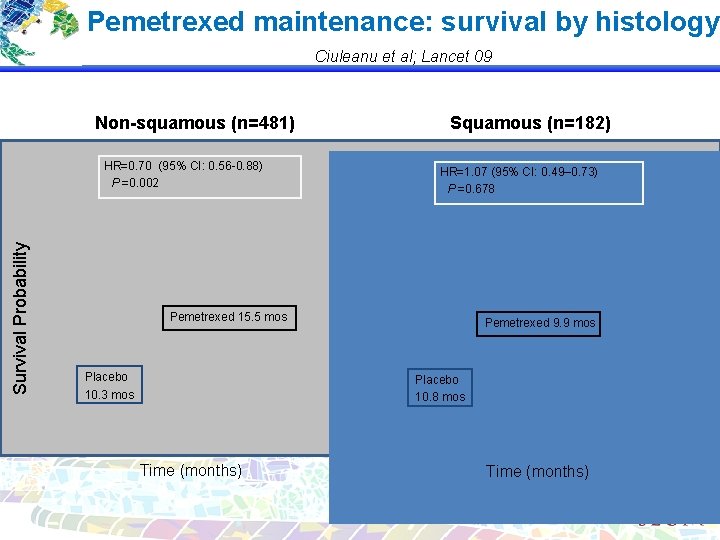
Pemetrexed maintenance: survival by histology Ciuleanu et al; Lancet 09 Non-squamous (n=481) Survival Probability HR=0. 70 (95% CI: 0. 56 -0. 88) P =0. 002 Squamous (n=182) HR=1. 07 (95% CI: 0. 49– 0. 73) P =0. 678 Pemetrexed 15. 5 mos Placebo 10. 3 mos Pemetrexed 9. 9 mos Placebo 10. 8 mos Time (months)

Histology: surrogate of molecular markers
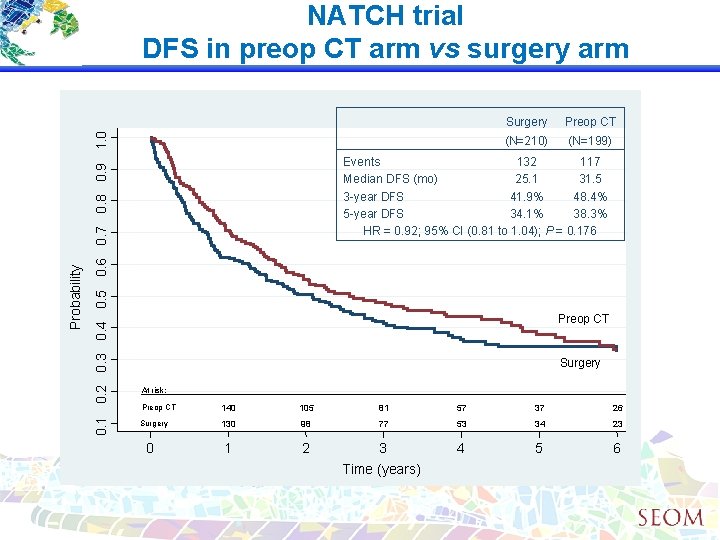
0. 1 0. 2 0. 3 0. 4 0. 5 0. 6 0. 7 0. 8 0. 9 1. 0 Probability NATCH trial DFS in preop CT arm vs surgery arm Surgery Preop CT (N=210) (N=199) Events 132 117 Median DFS (mo) 25. 1 31. 5 3 -year DFS 41. 9% 48. 4% 5 -year DFS 34. 1% 38. 3% HR = 0. 92; 95% CI (0. 81 to 1. 04); P = 0. 176 Preop CT Surgery At risk: Preop CT 140 105 81 57 37 26 Surgery 130 98 77 53 34 23 1 2 3 Time (years) 4 5 6 0
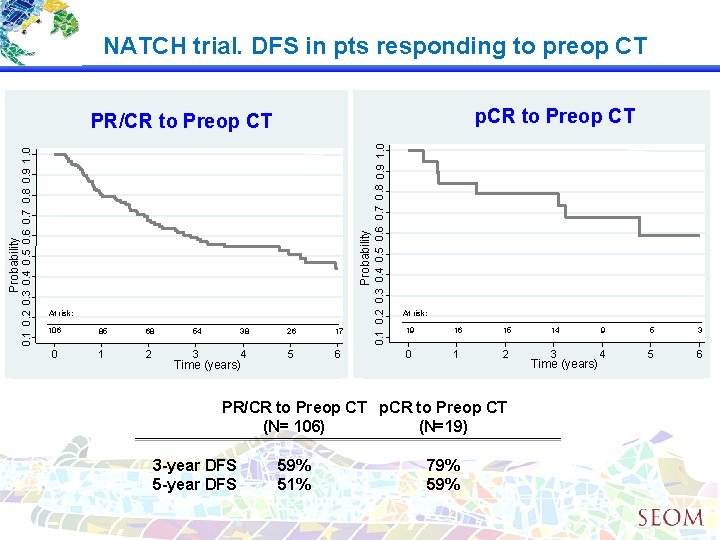
NATCH trial. DFS in pts responding to preop CT p. CR to Preop CT Probability At risk: 106 85 68 0 1 2 54 38 26 17 3 4 5 6 Time (years) 0. 1 0. 2 0. 3 0. 4 0. 5 0. 6 0. 7 0. 8 0. 9 1. 0 Probability 0. 1 0. 2 0. 3 0. 4 0. 5 0. 6 0. 7 0. 8 0. 9 1. 0 PR/CR to Preop CT At risk: 19 16 15 0 1 2 PR/CR to Preop CT p. CR to Preop CT (N= 106) (N=19) 3 -year DFS 59% 51% 79% 59% 14 9 5 3 3 4 5 6 Time (years)
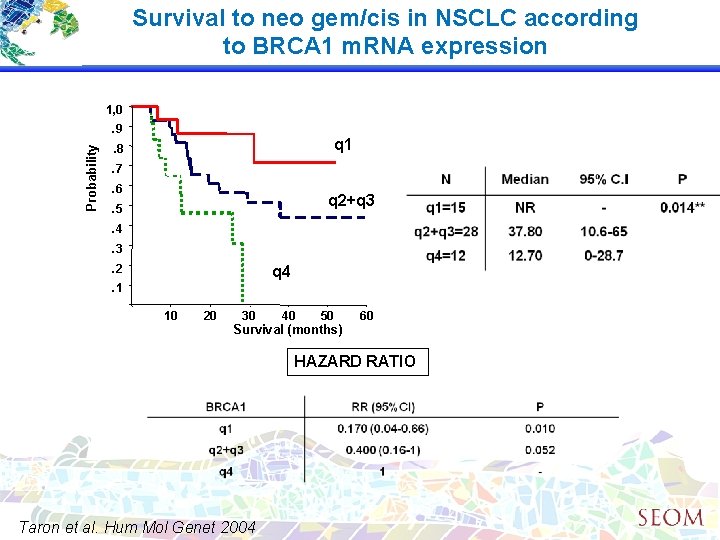
Survival to neo gem/cis in NSCLC according to BRCA 1 m. RNA expression 1, 0 Probability . 9 q 1 . 8. 7. 6 q 2+q 3 . 5. 4. 3. 2 q 4 . 1 10 20 30 40 50 Survival (months) 60 HAZARD RATIO Taron et al. Hum Mol Genet 2004

¿Se deben administrar las terapias siempre personalizadas en cáncer de pulmón? : SI • Mutaciones de EGFR predictivas de beneficio a EGFR TKIs • Mutaciones de KRAS y EMLK 4 -ALK + definen subgrupos de pacientes con diferentes evoluciones a inhibidores TKIs • Histología: surrogado de marcadores moleculares • Aplicación clínica marcadores genéticos en estadios iniciales; cuando se plantee QT neoadyuvante El tratamiento personalizado es una realidad en cáncer de pulmón
- Slides: 33