Science Technology Multiscale Modeling of Lipid Bilayer Interactions

Science & Technology Multiscale Modeling of Lipid Bilayer Interactions with Solid Substrates David R. Heine, Aravind R. Rammohan, and Jitendra Balakrishnan October 23 rd, 2008 RPI High Performance Computing Conference

Outline • Background – structure of lipid bilayers – applications of supported lipid bilayers • • • Modeling challenges Atomistic modeling Mesoscale modeling Experimental work Conclusions Science & Technology 2
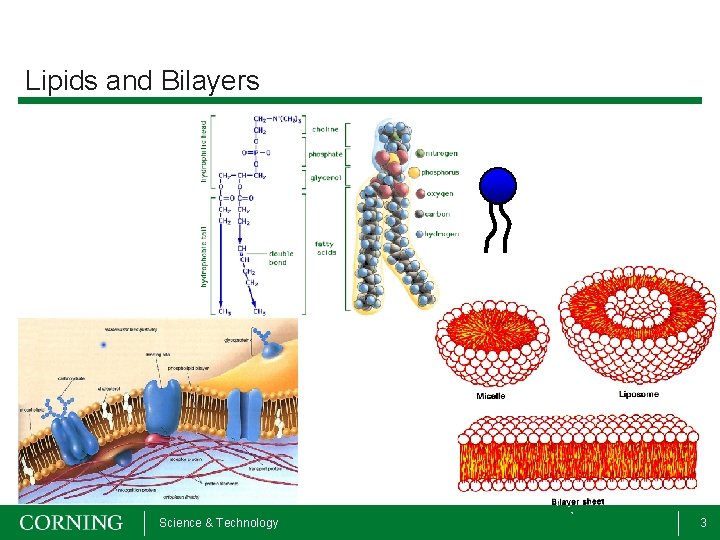
Lipids and Bilayers Science & Technology 3

Technological Relevance of Supported Lipid Bilayers • SLBs are important for various biotech applications – Biological research • • Model systems to study the properties of cell membranes Stable, immobilized base for research on membrane moieties Biosensors for the activity of various biological species Cell attachment surfaces – Pharmaceutical research • Investigation of membrane receptor drug targets • Membrane microarrays: High throughput screening for drug discovery – How does bilayer-substrate interaction affect bilayer behavior? Science & Technology 4

Supported Lipid Bilayers at Corning • Applications: Membrane-protein microarrays for pharmaceutical drug discovery • Substrate texture is important in the adhesion and conformation of bilayers on the surface – Crucial for the biological functionality of bilayers • Objective: Quantify the effect of substrate topography and chemical composition on bilayer conformation and dynamics Science & Technology 5

Bilayer Length & Time Scales • Bilayer dynamics vary over large length and time scales, suggesting a multiscale approach. Time Scales Length Scales Bond Vibrations: fs Stokes Radius: 2. 4 nm Lateral Diffusion Time: 4 ps Bilayer Thickness: 4 nm Area per lipid: 60 +/- 2 Undulations: 4 Å – 0. 25 mm Å2 Peristaltic Modes: 1 -10 ns Undulatory Modes 0. 1 ns – 0. 1 ms Membrane Fusion: 1 -10 s Science & Technology 6

Multiscale Approach • Atomistic model – capture local structure and short term dynamics • Mesoscale model – capture longer length and time scales – sufficient to look at interaction with rough surfaces Science & Technology 7
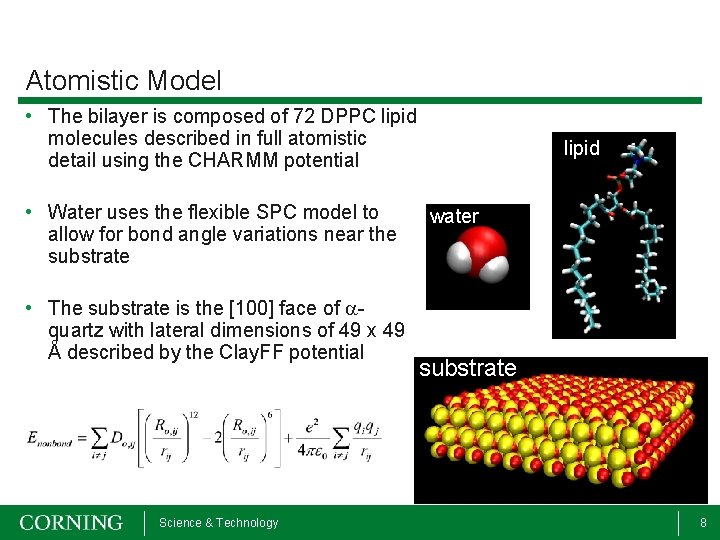
Atomistic Model • The bilayer is composed of 72 DPPC lipid molecules described in full atomistic detail using the CHARMM potential • Water uses the flexible SPC model to allow for bond angle variations near the substrate • The substrate is the [100] face of aquartz with lateral dimensions of 49 x 49 Å described by the Clay. FF potential Science & Technology lipid water substrate 8

Simulation Technique • System is periodic in x and y directions with a repulsive wall above the water surface in the z direction Water • NVT ensemble must be used since pressure control is prohibited by the solid substrate Upper leaflet • Temperature is maintained at 323 K with a Nose-Hoover thermostat • Total energy and force on the bilayer are extracted during the simulation. Heine et al. Molecular Simulations, 2007, 33(4 -5), pp. 391 -397. Science & Technology Lower leaflet Bilayer Lipids Water Substrate 9

Simulation Technique • System is periodic in x and y directions with a repulsive wall above the water surface in the z direction • NVT ensemble must be used since pressure control is prohibited by the solid substrate • Temperature is maintained at 323 K with a Nose-Hoover thermostat • Total energy and force on the bilayer are extracted during the simulation. Heine et al. Molecular Simulations, 2007, 33(4 -5), pp. 391 -397. Science & Technology 10
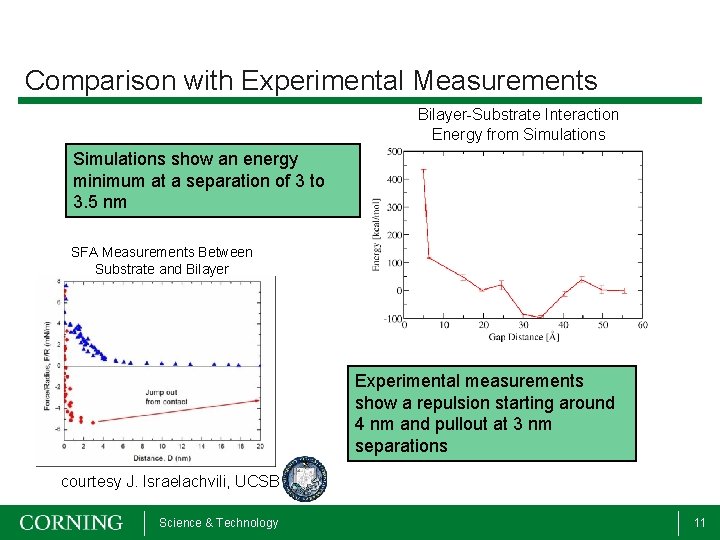
Comparison with Experimental Measurements Bilayer-Substrate Interaction Energy from Simulations show an energy minimum at a separation of 3 to 3. 5 nm SFA Measurements Between Substrate and Bilayer Experimental measurements show a repulsion starting around 4 nm and pullout at 3 nm separations courtesy J. Israelachvili, UCSB Science & Technology 11
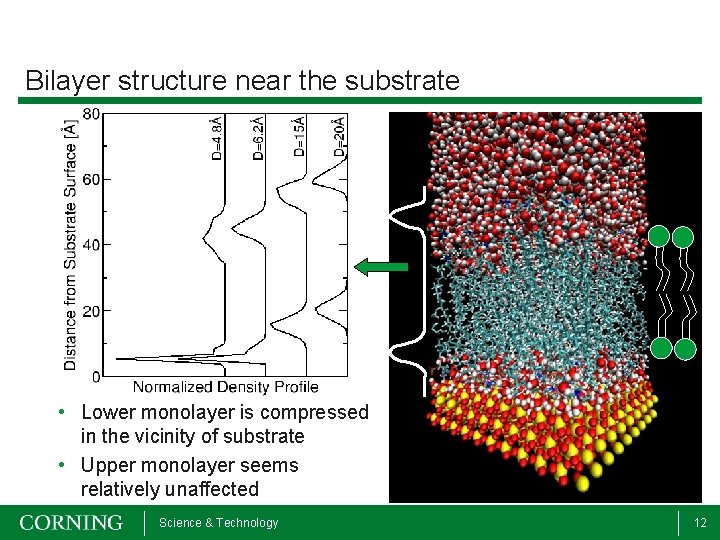
Bilayer structure near the substrate • Lower monolayer is compressed in the vicinity of substrate • Upper monolayer seems relatively unaffected Science & Technology 12
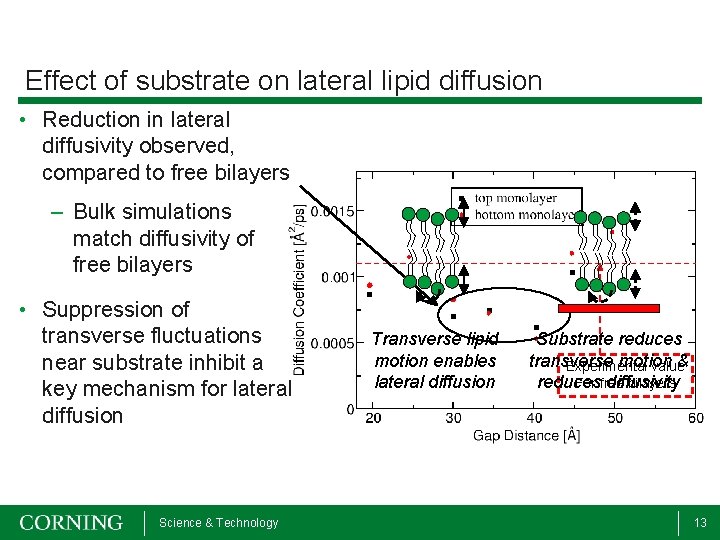
Effect of substrate on lateral lipid diffusion • Reduction in lateral diffusivity observed, compared to free bilayers – Bulk simulations match diffusivity of free bilayers • Suppression of transverse fluctuations near substrate inhibit a key mechanism for lateral diffusion Science & Technology Transverse lipid motion enables lateral diffusion Substrate reduces transverse motion & Experimental value For free bilayers reduces diffusivity 13

Atomistic Simulation Results • MD simulations show bilayer-substrate equilibrium separation of 3 – 3. 5 nm, in agreement with SFA experiments • Lateral diffusion of the lipid head groups decreases as the bilayer approaches the substrate • Suppression of transverse fluctuations may be responsible for reduced lateral diffusion Science & Technology 14
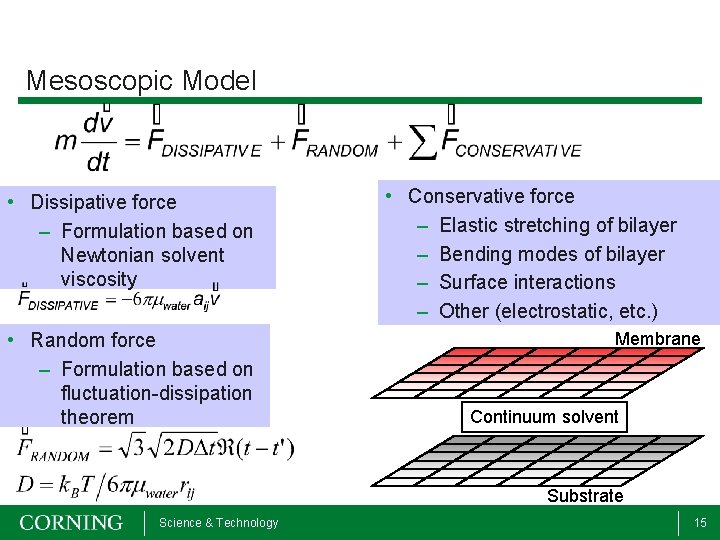
Mesoscopic Model • Dissipative force – Formulation based on Newtonian solvent viscosity • Random force – Formulation based on fluctuation-dissipation theorem • Conservative force – Elastic stretching of bilayer – Bending modes of bilayer – Surface interactions – Other (electrostatic, etc. ) Membrane Continuum solvent Substrate Science & Technology 15
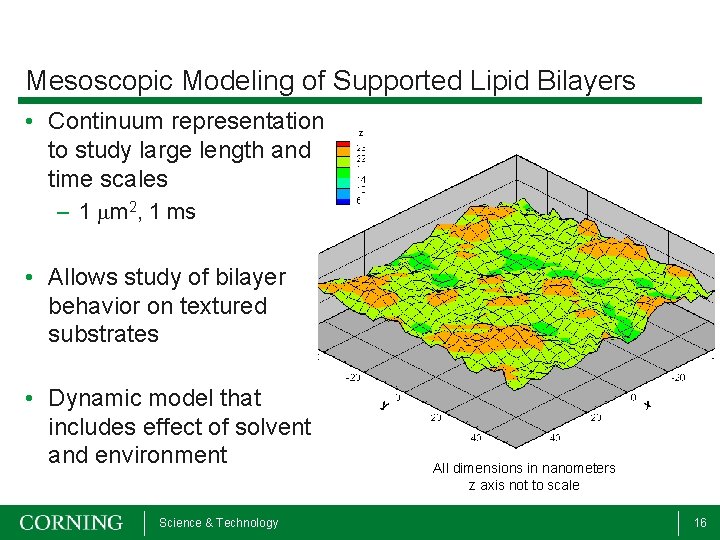
Mesoscopic Modeling of Supported Lipid Bilayers • Continuum representation to study large length and time scales – 1 mm 2, 1 ms • Allows study of bilayer behavior on textured substrates • Dynamic model that includes effect of solvent and environment Science & Technology All dimensions in nanometers z axis not to scale 16
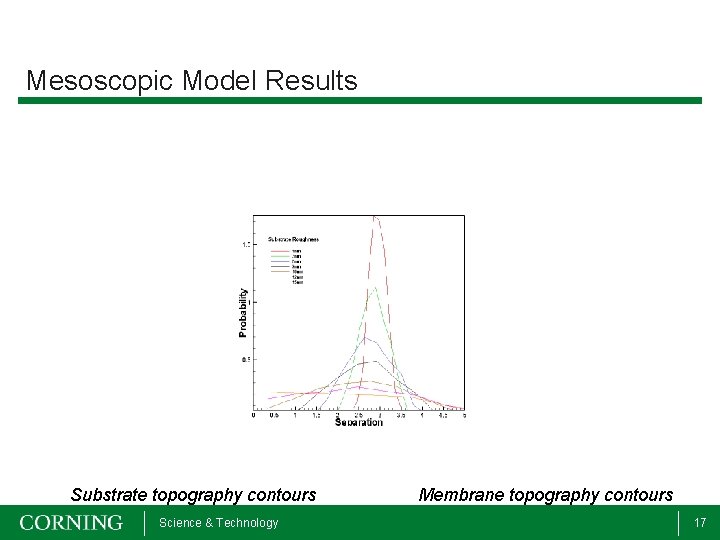
Mesoscopic Model Results Substrate topography contours Science & Technology Membrane topography contours 17
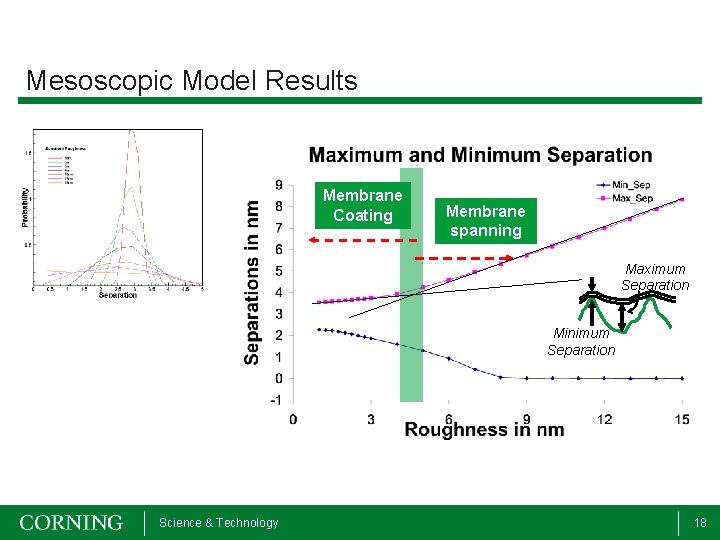
Mesoscopic Model Results Membrane Coating Membrane spanning Maximum Separation Minimum Separation Science & Technology 18

Mesoscopic Model Results • Allows study of bilayer on micron and microsecond scales • Minimum surface roughness of 4 -5 nm required for membrane spanning conformation • Spanning configuration important for maintaining bilayer mobility Science & Technology 19

AFM measurements Spreading of Bilayer on Synthetic Substrates AFM image & measurements courtesy Sergiy Minko, Clarkson University Ref: Nanoletters, 2008, 8(3), 941 -944 Science & Technology 20

AFM measurements Smoothening of membrane on rough substrates AFM image & measurements courtesy Sergiy Minko, Clarkson University Science & Technology 21
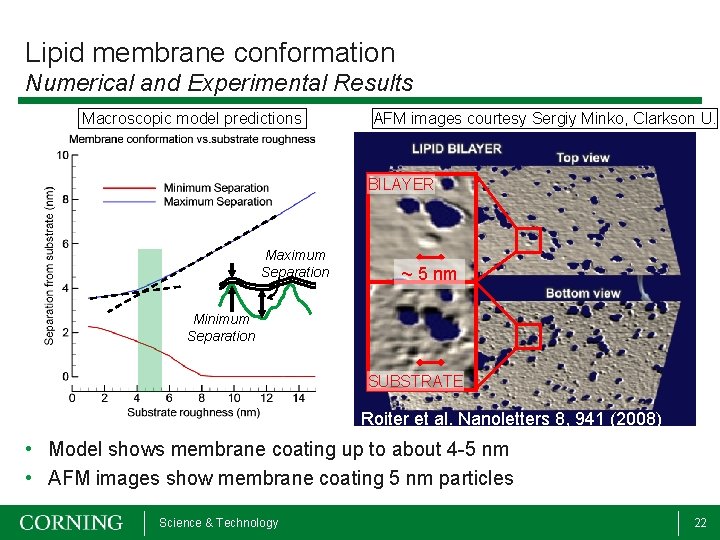
Lipid membrane conformation Numerical and Experimental Results Macroscopic model predictions AFM images courtesy Sergiy Minko, Clarkson U. BILAYER Maximum Separation ~ 5 nm Minimum Separation SUBSTRATE Roiter et al. Nanoletters 8, 941 (2008) • Model shows membrane coating up to about 4 -5 nm • AFM images show membrane coating 5 nm particles Science & Technology 22

Conclusions • MD simulations show bilayer-substrate separation of 3 – 3. 5 nm, in agreement with SFA experiments • MD simulations show reduced lateral diffusion in lipids as the bilayer approaches the substrate • Mesoscopic model shows membranes coat particles up to 4 – 5 nm in diameter, in agreement with AFM observations • Larger surface features are needed to achieve separation between bilayer and substrate • High-performance computing has opened up new approaches for understanding biomolecule-substrate interactions, which aids design • There is still plenty of room to grow as these models are still restricted in terms of size, timescale, and complexity Science & Technology 23

Acknowledgements • Professor Sergiy Minko & his group at Clarkson U. • Professor Jacob Israelachvili & his group at U. C. Santa Barbara Science & Technology 24

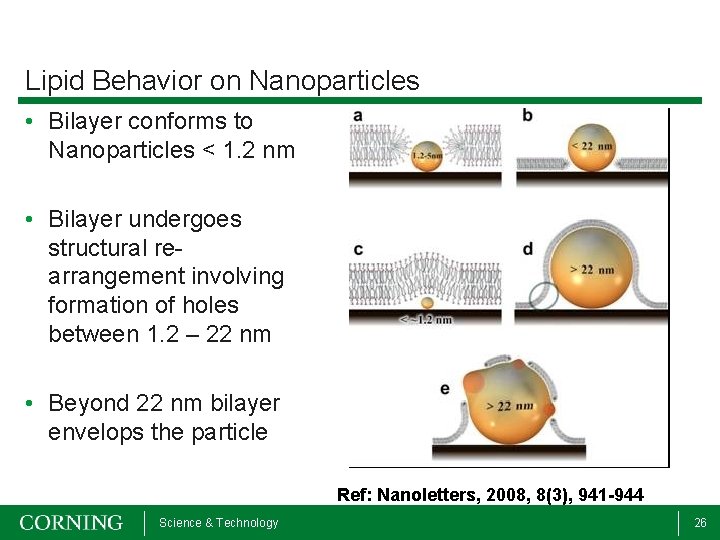
Lipid Behavior on Nanoparticles • Bilayer conforms to Nanoparticles < 1. 2 nm • Bilayer undergoes structural rearrangement involving formation of holes between 1. 2 – 22 nm • Beyond 22 nm bilayer envelops the particle Ref: Nanoletters, 2008, 8(3), 941 -944 Science & Technology 26
- Slides: 26