Schedule Last week pAcceptor Ligands and Biology CO


Schedule • Last week: p-Acceptor Ligands and Biology CO, O 2, N 2 and NO complexes, introduction to M-M bonds • Lecture 7: M-M bonds d-bonds and bonding in metal clusters • Lecture 8: Rates of reaction Ligand-exchange reactions, labile and inert metal ions • Lecture 9: Redox reactions Inner and outer-sphere reactions Slide 2/20

Summary of Course – week 6 Metal-metal bonding • be able to predict bond order for M 2 Lx dimers using d-electron count and s, p and d molecular orbital diagram • be able to predict bond order in larger metal-halide clusters using delectron count shared over edges of cluster • be able to predict bond order in metal carbonyl clusters using 18 e- rule Reaction mechanisms • be able to describe ligand exchange mechanisms • be able to explain role of metal charge and LFSE in rate of ligand exchange • be able to describe electron transfer reaction mechanisms • be able to predict relative rate of outer sphere reaction for different metals Resources • Slides for lectures 7 -9 • Shriver and Atkins “Inorganic Chemistry” Chapter 18. 11, 21. 20, 20. 1 -20. 13 Slide 3/20

Summary of Last Lecture Metal-N 2 complexes • N 2 is isoelectronic with CO but M-N 2 bonding is much weaker • N 2 is non-polar and bond is strong NO complexes • Can bond as 1 electron donor (NO-: bent M-NO) • Can bond as 2 electron donor (NO+ linear M-NO) Today’s lecture • Metal-Metal bonding in complexes Slide 4/20
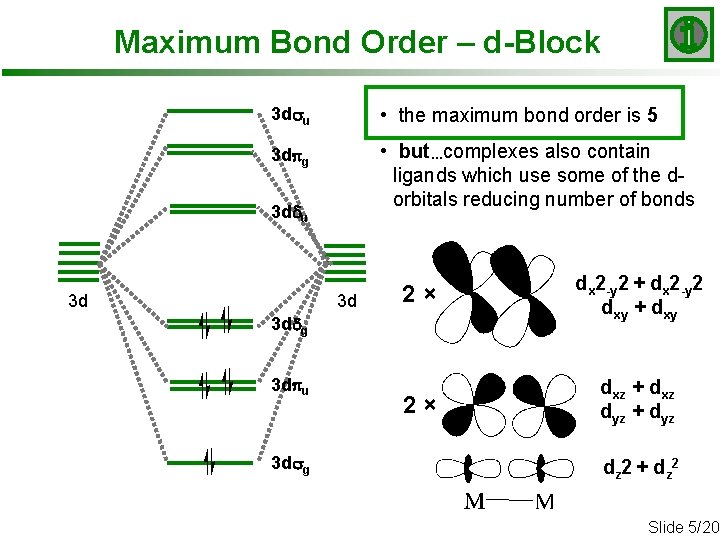
Maximum Bond Order – d-Block 3 dsu • the maximum bond order is 5 3 dpg • but…complexes also contain ligands which use some of the dorbitals reducing number of bonds 3 ddu 3 d 3 d 2× dx 2 -y 2 + dx 2 -y 2 dxy + dxy 2× dxz + dxz dyz + dyz 3 ddg 3 dpu 3 dsg dz 2 + d z 2 Slide 5/20
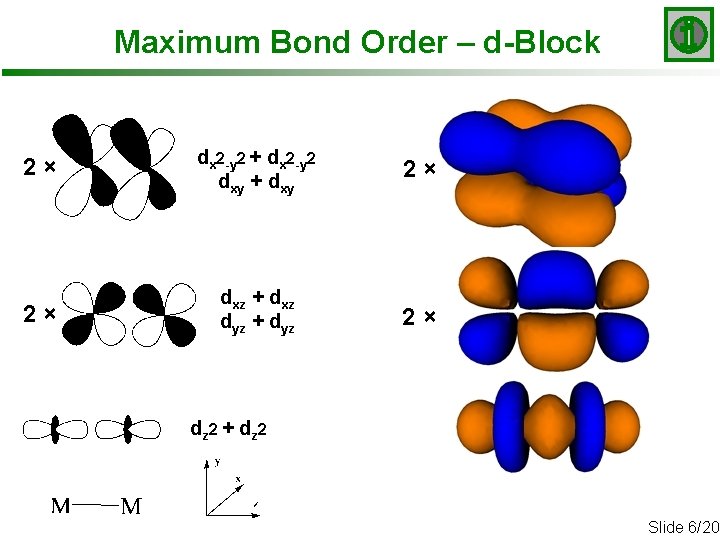
Maximum Bond Order – d-Block 2× dx 2 -y 2 + dx 2 -y 2 dxy + dxy 2× 2× dxz + dxz dyz + dyz 2× dz 2 + d z 2 Slide 6/20
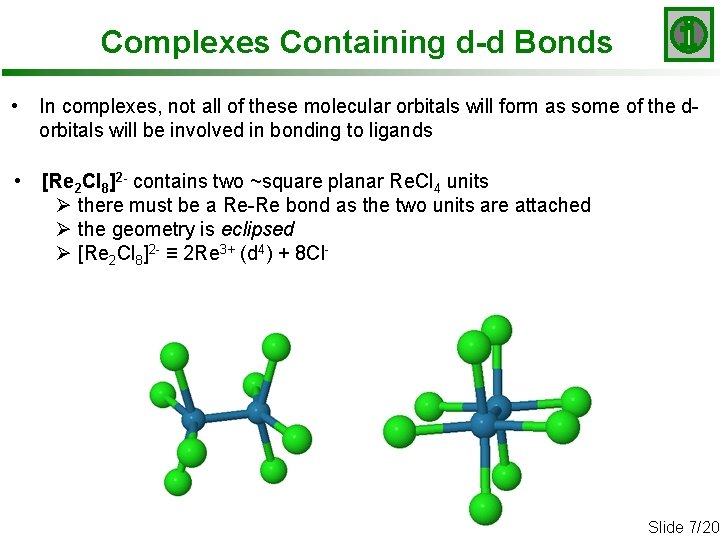
Complexes Containing d-d Bonds • In complexes, not all of these molecular orbitals will form as some of the dorbitals will be involved in bonding to ligands • [Re 2 Cl 8]2 - contains two ~square planar Re. Cl 4 units Ø there must be a Re-Re bond as the two units are attached Ø the geometry is eclipsed Ø [Re 2 Cl 8]2 - ≡ 2 Re 3+ (d 4) + 8 Cl- Slide 7/20
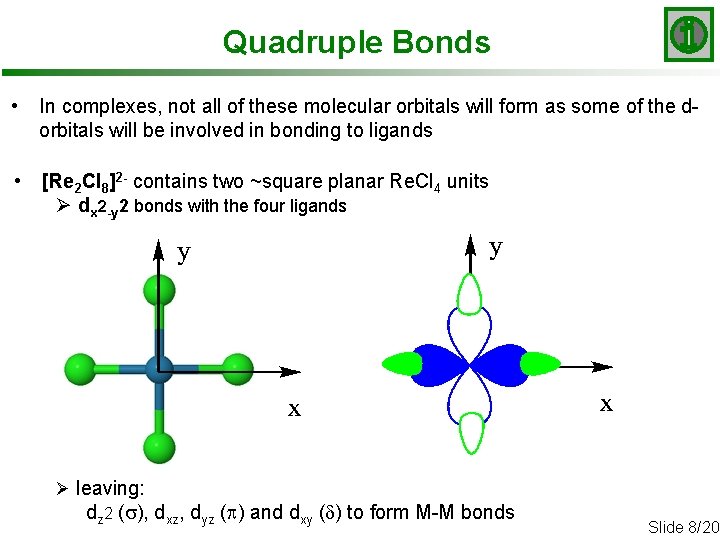
Quadruple Bonds • In complexes, not all of these molecular orbitals will form as some of the dorbitals will be involved in bonding to ligands • [Re 2 Cl 8]2 - contains two ~square planar Re. Cl 4 units Ø dx 2 -y 2 bonds with the four ligands Ø leaving: dz 2 (s), dxz, dyz (p) and dxy (d) to form M-M bonds Slide 8/20
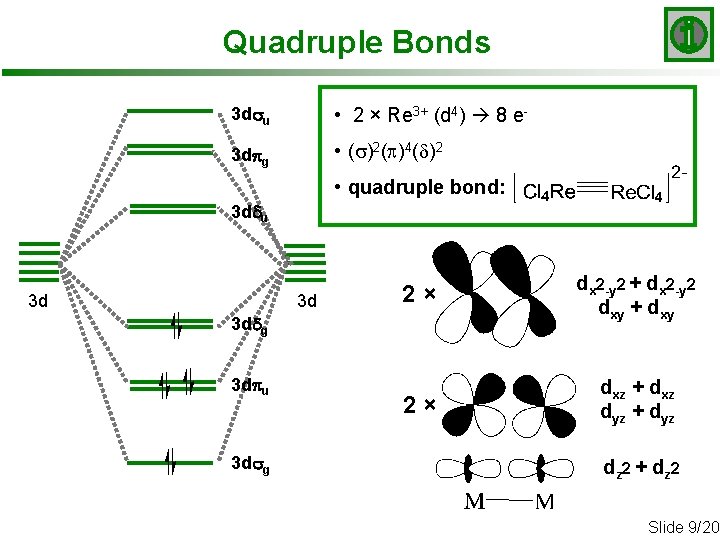
Quadruple Bonds 3 dsu • 2 × Re 3+ (d 4) 8 e- 3 dpg • (s)2(p)4(d)2 • quadruple bond: 3 ddu 3 d 3 d 2× dx 2 -y 2 + dx 2 -y 2 dxy + dxy 2× dxz + dxz dyz + dyz 3 ddg 3 dpu 3 dsg dz 2 + d z 2 Slide 9/20
![Quadruple Bonds • [Re 2 Cl 8]2 - quaduple bond: Ø (s)2(p)4(d)2: a s-bond, Quadruple Bonds • [Re 2 Cl 8]2 - quaduple bond: Ø (s)2(p)4(d)2: a s-bond,](http://slidetodoc.com/presentation_image_h/301eed13d3a2b57b5d1c1f8a319a72f5/image-10.jpg)
Quadruple Bonds • [Re 2 Cl 8]2 - quaduple bond: Ø (s)2(p)4(d)2: a s-bond, two p-bonds and one d-bond Ø the sterically unfavourable eclipsed geometry is due to the d-bond Slide 10/20
![Quadruple Bonds • • [Re 2 Cl 8]2Ø [Re 2 Cl 8]2 - ≡ Quadruple Bonds • • [Re 2 Cl 8]2Ø [Re 2 Cl 8]2 - ≡](http://slidetodoc.com/presentation_image_h/301eed13d3a2b57b5d1c1f8a319a72f5/image-11.jpg)
Quadruple Bonds • • [Re 2 Cl 8]2Ø [Re 2 Cl 8]2 - ≡ 2 Re 3+ (d 4) + 8 ClØ 2 × Re 3+ (d 4) 8 eØ (s)2(p)4(d)2 Ø bond order = 4 (quadruple bond) Ø d-bond: eclipsed geometry [Re 2 Cl 8]4Ø [Re 2 Cl 8]4 - ≡ 2 Re 2+ (d 5) + 8 ClØ 2 × Re 2+ (d 5) 10 eØ (s)2(p)4(d)2(d*)2 Ø bond order = 3 (triple bond) Ø no d-bond: staggered geometry 3 dsu 3 dpg 3 ddu 3 ddg 3 dpu 3 dsg Slide 11/20
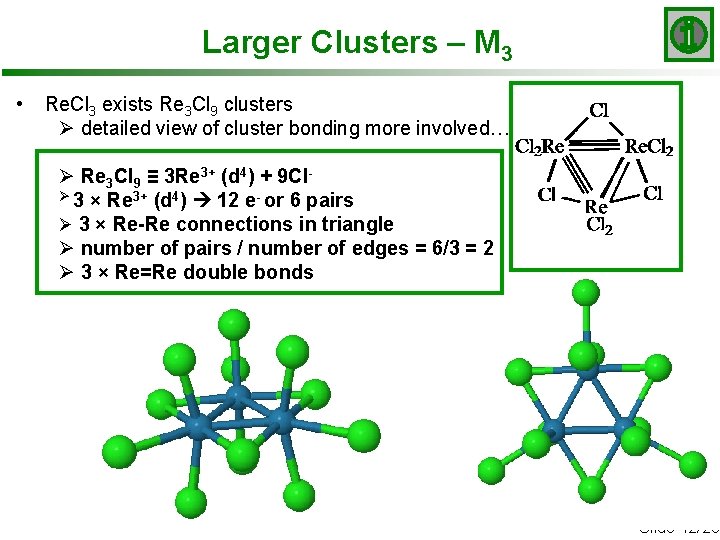
Larger Clusters – M 3 • Re. Cl 3 exists Re 3 Cl 9 clusters Ø detailed view of cluster bonding more involved… Ø Re 3 Cl 9 ≡ 3 Re 3+ (d 4) + 9 ClØ 3 × Re 3+ (d 4) 12 e- or 6 pairs Ø 3 × Re-Re connections in triangle Ø number of pairs / number of edges = 6/3 = 2 Ø 3 × Re=Re double bonds Slide 12/20
![Larger Clusters – M 6 • [Mo 6 Cl 14]2Ø Mo 6 octahedron with Larger Clusters – M 6 • [Mo 6 Cl 14]2Ø Mo 6 octahedron with](http://slidetodoc.com/presentation_image_h/301eed13d3a2b57b5d1c1f8a319a72f5/image-13.jpg)
Larger Clusters – M 6 • [Mo 6 Cl 14]2Ø Mo 6 octahedron with 8 Cl capping faces and 6 Cl on edges Ø [Mo 6 Cl 14]2 - ≡ 6 Mo 2+ (d 4) + 14 ClØ 6 × Mo 2+ (d 4) 24 e- or 12 pairs Ø 12 × Mo-Mo connections in octahedron Ø number of pairs / number of edges = 12/12 = 1 Ø 12 × Mo-Mo single bonds Slide 13/20
![Larger Clusters – M 6 • [Nb 6 Cl 12]2+ Ø Nb 6 octahedron Larger Clusters – M 6 • [Nb 6 Cl 12]2+ Ø Nb 6 octahedron](http://slidetodoc.com/presentation_image_h/301eed13d3a2b57b5d1c1f8a319a72f5/image-14.jpg)
Larger Clusters – M 6 • [Nb 6 Cl 12]2+ Ø Nb 6 octahedron with 12 Cl on edges Ø [Nb 6 Cl 12]2+ ≡ [Nb 6]14+ + 12 ClØ 6 × Nb (d 5) 30 eØ [Nb ]14+ so… 6 number for M-M bonding is 30 – 14 = 16 e- or 8 pairs Ø 12 × Nb-Nb connections in octahedron Ø number of pairs / number of edges = 8/12 = 2/3 Ø 12 × Nb-Nb bonds with bond order = ⅔ Slide 14/20
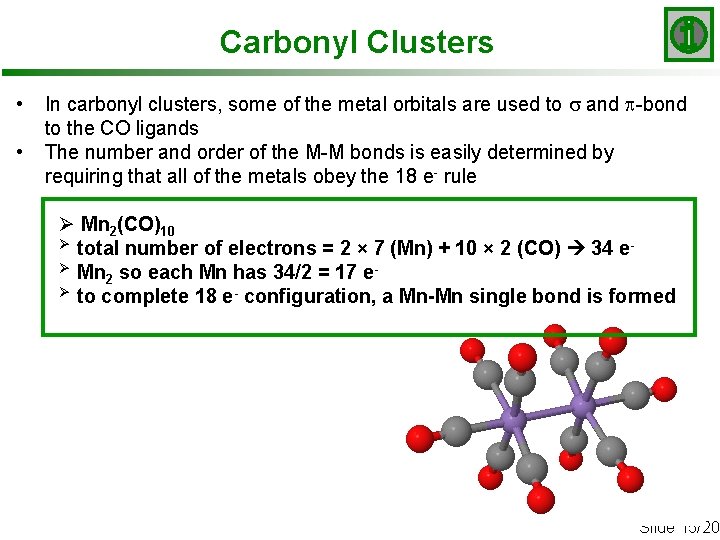
Carbonyl Clusters • • In carbonyl clusters, some of the metal orbitals are used to s and p-bond to the CO ligands The number and order of the M-M bonds is easily determined by requiring that all of the metals obey the 18 e- rule Ø Mn 2(CO)10 Ø total number of electrons = 2 × 7 (Mn) + 10 × 2 (CO) 34 eØ Mn so each Mn has 34/2 = 17 e 2 Ø to complete 18 e- configuration, a Mn-Mn single bond is formed Slide 15/20
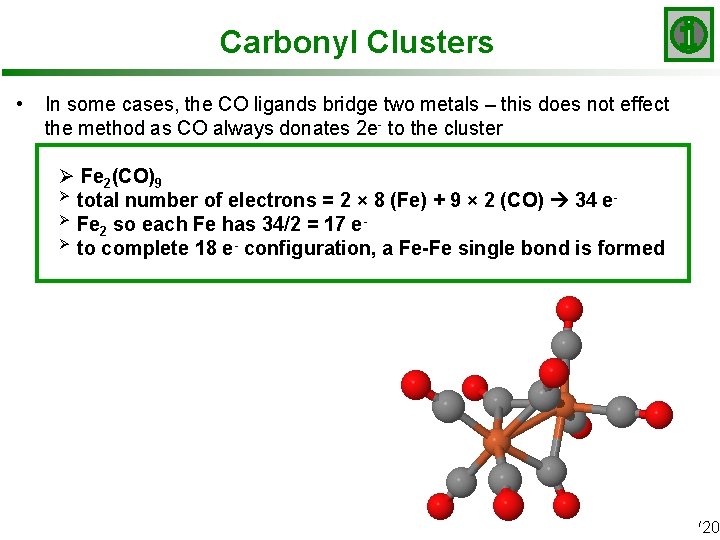
Carbonyl Clusters • In some cases, the CO ligands bridge two metals – this does not effect the method as CO always donates 2 e- to the cluster Ø Fe 2(CO)9 Ø total number of electrons = 2 × 8 (Fe) + 9 × 2 (CO) 34 eØ Fe so each Fe has 34/2 = 17 e 2 Ø to complete 18 e- configuration, a Fe-Fe single bond is formed Slide 16/20
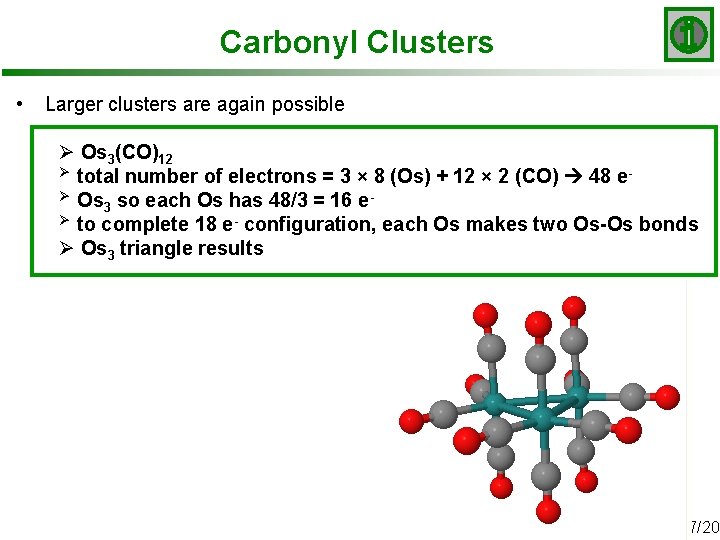
Carbonyl Clusters • Larger clusters are again possible Ø Os 3(CO)12 Ø total number of electrons = 3 × 8 (Os) + 12 × 2 (CO) 48 eØ Os so each Os has 48/3 = 16 e 3 Ø to complete 18 e- configuration, each Os makes two Os-Os bonds Ø Os 3 triangle results Slide 17/20

Summary By now you should be able to • Draw out the MO diagram for L 4 ML 4 complexes • Complete this diagram by filling in the appropriate number of electrons • Explain the appearance of eclipsed geometries due to dbonding • In larger clusters, use the total number of pairs of metal electrons and number of metal-metal connections to work out the bond order • In carbonyl clusters, use the 18 e- rule to work out how many bonds have to be formed Next lecture • Ligand-substitution reactions Slide 18/20

Practice Slide 19/20
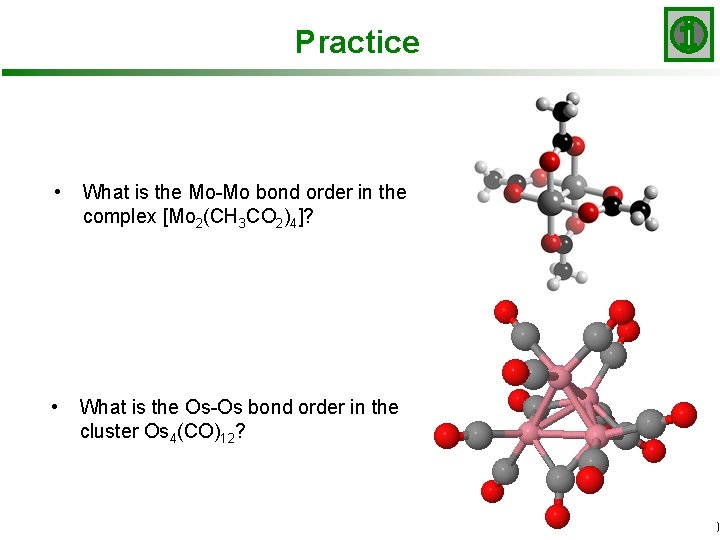
Practice • What is the Mo-Mo bond order in the complex [Mo 2(CH 3 CO 2)4]? • What is the Os-Os bond order in the cluster Os 4(CO)12? Slide 20/20
- Slides: 20