Sample Preparation Enzymatic Digestion Trypsin Fractionation Single Stage











- Slides: 29

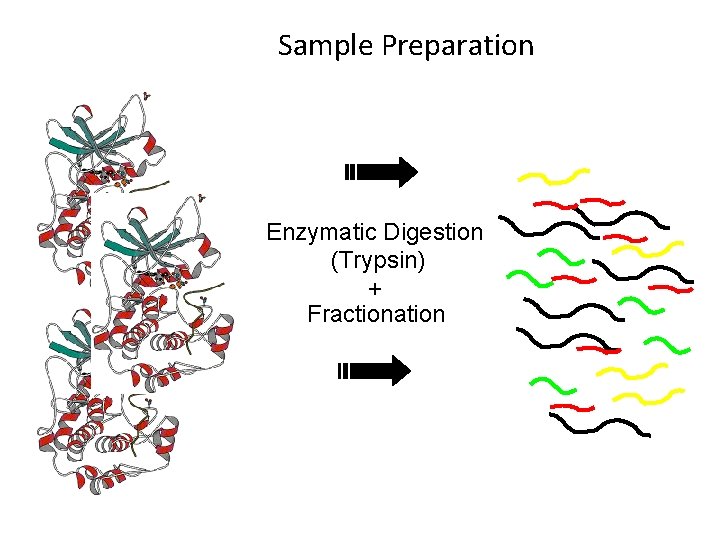
Sample Preparation Enzymatic Digestion (Trypsin) + Fractionation

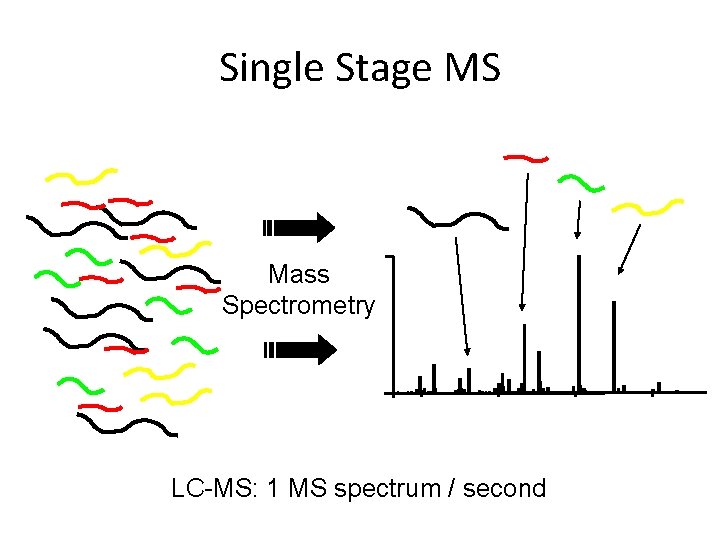
Single Stage MS Mass Spectrometry LC-MS: 1 MS spectrum / second

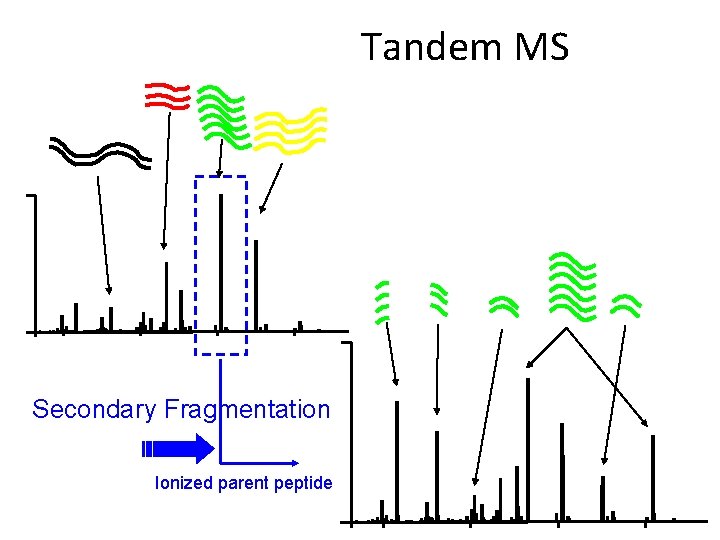
Tandem MS Secondary Fragmentation Ionized parent peptide

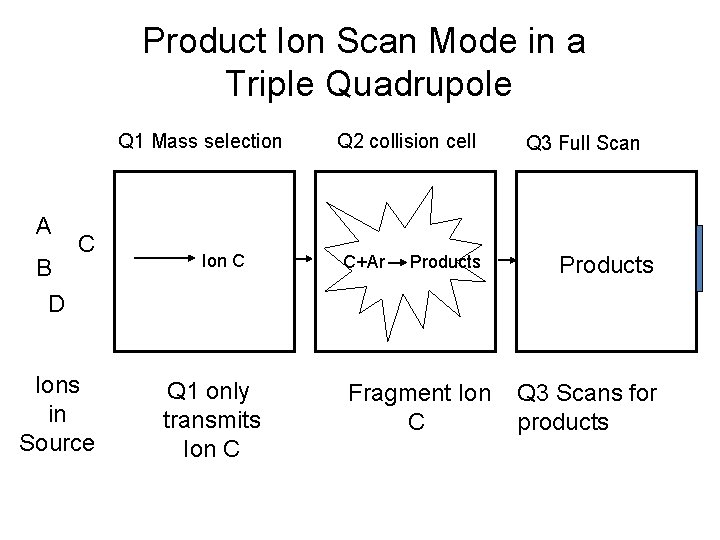
Product Ion Scan Mode in a Triple Quadrupole Q 1 Mass selection A B D C Ions in Source Ion C Q 1 only transmits Ion C Q 2 collision cell C+Ar Products Fragment Ion C Q 3 Full Scan Products Q 3 Scans for products

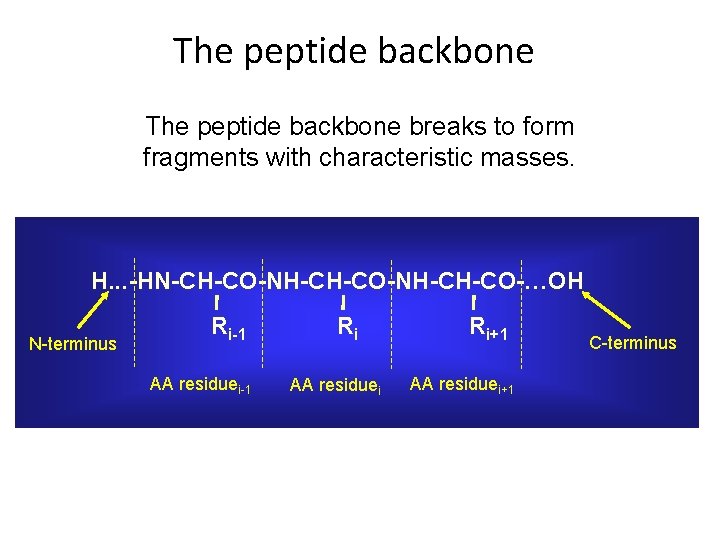
The peptide backbone breaks to form fragments with characteristic masses. H. . . -HN-CH-CO-NH-CH-CO-…OH N-terminus Ri-1 AA residuei-1 Ri AA residuei Ri+1 AA residuei+1 C-terminus

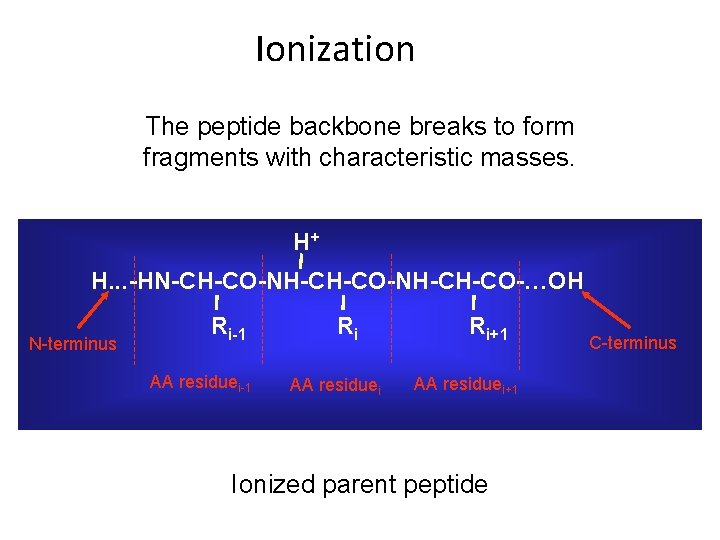
Ionization The peptide backbone breaks to form fragments with characteristic masses. H+ H. . . -HN-CH-CO-NH-CH-CO-…OH N-terminus Ri-1 AA residuei-1 Ri AA residuei Ri+1 AA residuei+1 Ionized parent peptide C-terminus

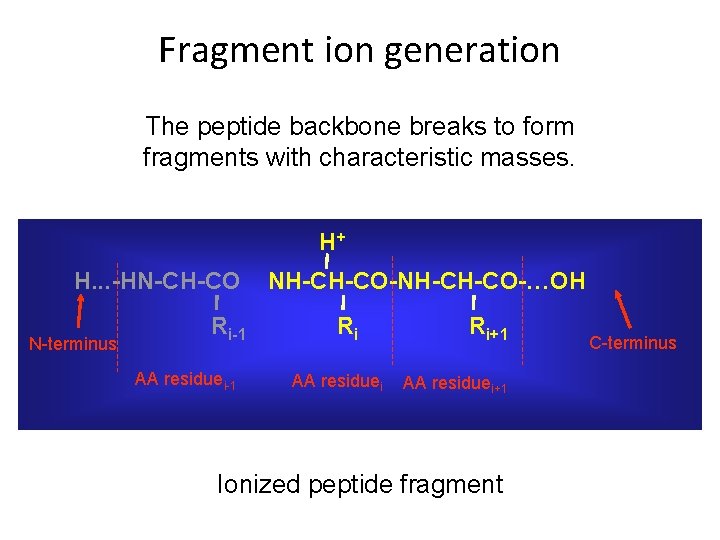
Fragment ion generation The peptide backbone breaks to form fragments with characteristic masses. H+ H. . . -HN-CH-CO N-terminus Ri-1 AA residuei-1 NH-CH-CO-…OH Ri AA residuei Ri+1 AA residuei+1 Ionized peptide fragment C-terminus

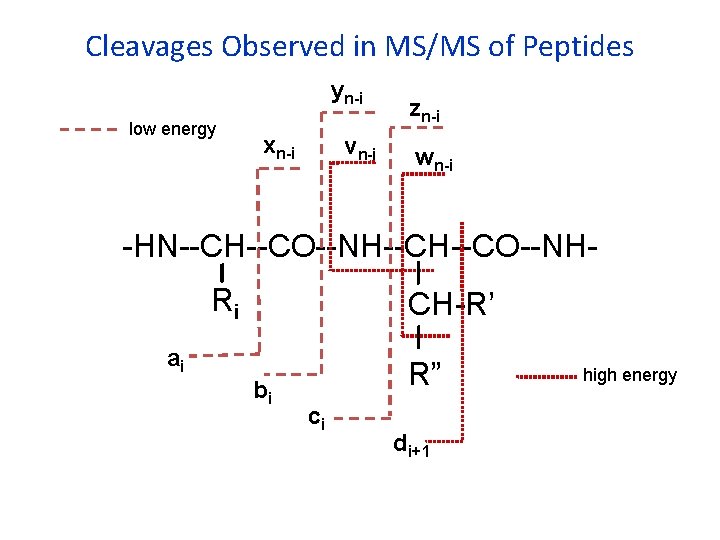
Cleavages Observed in MS/MS of Peptides yn-i low energy xn-i vn-i zn-i wn-i -HN--CH--CO--NHRi CH-R’ ai bi R” ci di+1 high energy

Different MS-MS Instruments Yield Different Spectra • A typical QTOF or triple quad MS-MS spectrum of a tryptic peptide contains a continuous series of ytype ions. The b-type ions are usually seen only at lower masses below the precursor m/z value • Ion trap CID data of tryptic peptides is different in that one often finds a continuous series of both btype and y-type ions throughout the spectrum

MS-MS for Protein ID • Proteins are isolated (from gel or HPLC) and subjected to tryptic digestion • Peptides are sent through ionizer and into a collision cell where the doubly charged ions are selected and fragmented through collision induced decay (CID) • The resulting singly charged ions (daughter ions) are analyzed to determine the sequence or to ID the parent peptide

Why Trypsin for MS-MS? • CID of peptides less than 2 -3 k. D is most reliable for MS-MS studies – The frequency of tryptic cleavage guarantees that most peptides will be of this size • Trypsin cleaves on the C-terminal side of arginine and lysine. By putting the basic residues at the Cterminus, peptides fragment in a more predictable manner throughout the length of the peptide

Why Double Charges? • Easiest spectra to interpret are those obtained from doubly-charged peptide precursors, where the resulting fragment ions are mostly singly-charged • Doubly-charged precursors also fragment such that most of the peptide bonds break with comparable frequency, such that one is more likely to derive a complete sequence

How the peptide sequencing works? • Use Tandem MS: two mass analyzer in series with a collision cell in between • Collision cell: a region where the ions collide with a gas (He, Ne, Ar) resulting in fragmentation of the ion • Fragmentation of the peptides in the collision cell occur in a predictable fashion, mainly at the peptide bonds (also phosphoester bonds) • The resulting daughter ions have masses that are consistent with known molecular weights of dipeptides, tripeptides, tetrapeptides… Ser-Glu-Leu-Ile-Arg-Trp Collision Cell Ser-Glu-Leu-Ile-Arg Ser-Glu-Leu-Ile Ser-Glu-Leu Etc…

MS-MS & Peptide Fragments • When peptides are proteins are admitted to a collision cell the peptide usually fragments at the weakest bond (the peptide bond, but some CHNH and CH-CO breakage also occurs) • Collision conditions have to be optimized for each peptide • Two main types of daughter ions are produced -“b” ions and “y” ions

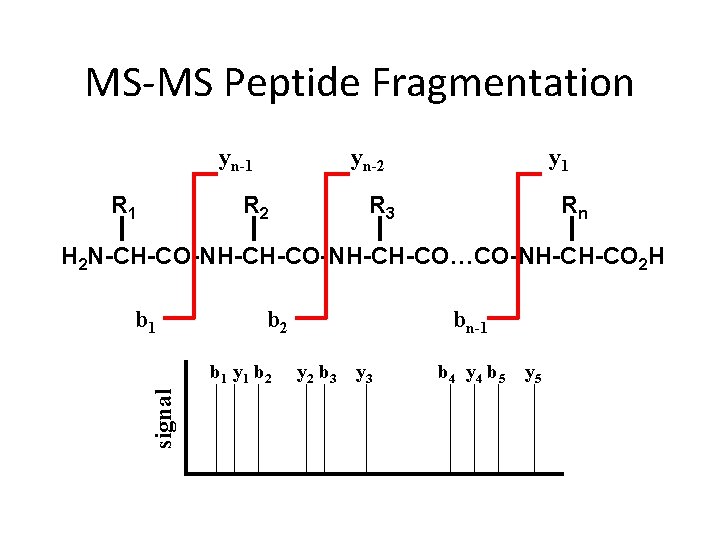
MS-MS Peptide Fragmentation yn-1 R 1 yn-2 R 2 y 1 R 3 Rn H 2 N-CH-CO-NH-CH-CO…CO-NH-CH-CO 2 H b 1 b 2 signal b 1 y 1 b 2 bn-1 y 2 b 3 y 3 b 4 y 4 b 5 y 5

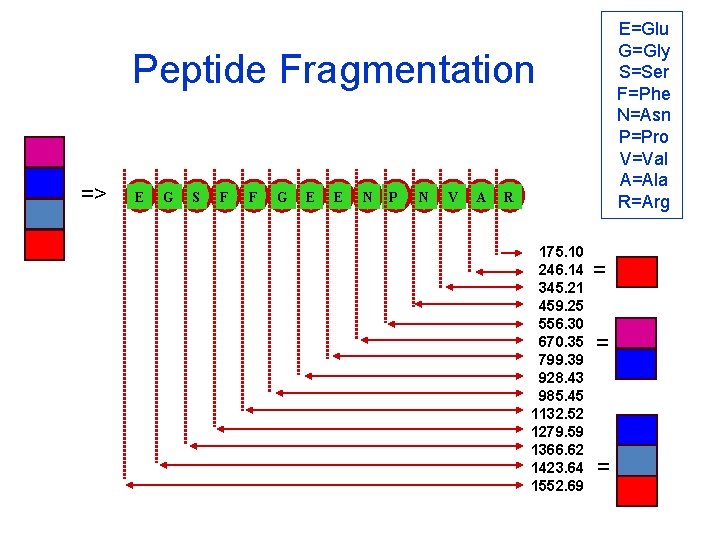
E=Glu G=Gly S=Ser F=Phe N=Asn P=Pro V=Val A=Ala R=Arg Peptide Fragmentation => E G S F F G E E N P N V A R 175. 10 246. 14 345. 21 459. 25 556. 30 670. 35 799. 39 928. 43 985. 45 1132. 52 1279. 59 1366. 62 1423. 64 1552. 69 = = =

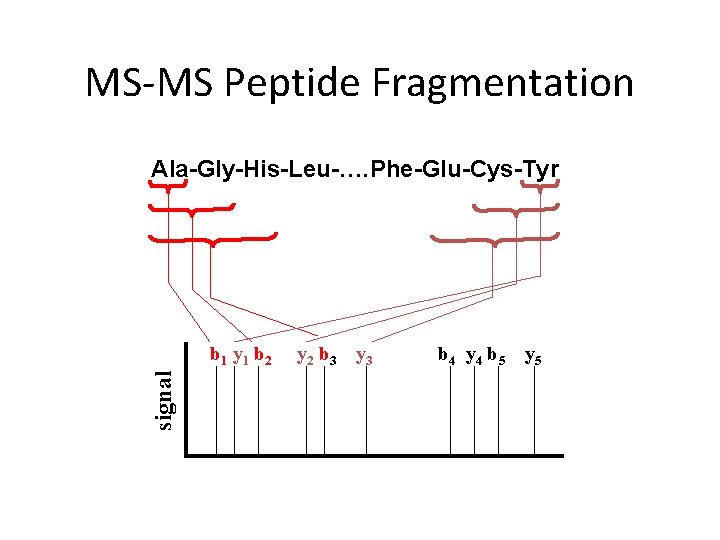
MS-MS Peptide Fragmentation Ala-Gly-His-Leu-…. Phe-Glu-Cys-Tyr signal b 1 y 1 b 2 y 2 b 3 y 3 b 4 y 4 b 5 y 5

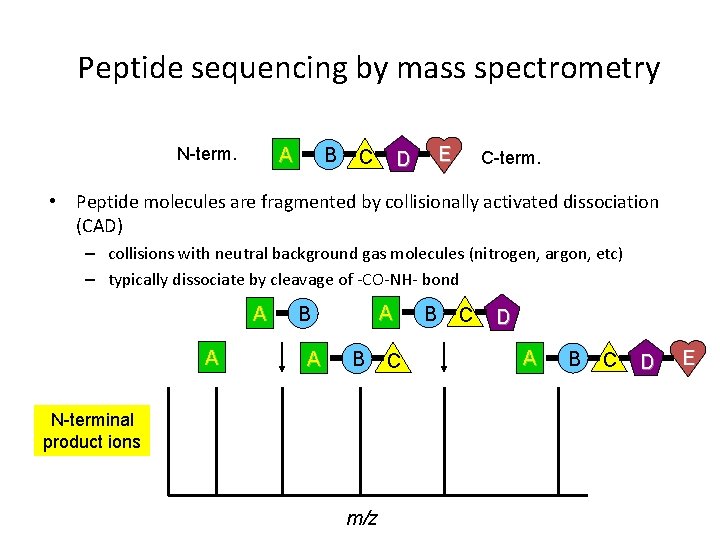
Peptide sequencing by mass spectrometry N-term. A B C E D C-term. • Peptide molecules are fragmented by collisionally activated dissociation (CAD) – collisions with neutral background gas molecules (nitrogen, argon, etc) – typically dissociate by cleavage of -CO-NH- bond A A A B N-terminal product ions m/z C B C D A B C D E

Peptide Sequencing

Protocols for MS-MS Sequencing • Usually can’t tell a “b” ion from a “y” ion • Assume the lowest mass visible in the spectrum is a lysine or arginine (this is the y 1 ion) this is because trypsin cuts after a lysine or arginine • This y 1 mass should be 147. 113 for lysine or 175. 119 for arginine {The y 1 ion is calculated by adding 19. 018 u (three hydrogens and one oxygen) to the residue masses of lysine and arginine}

MS-MS Sequencing • Use the remaining “unassigned” peaks to see if you can construct a “b” ion series • The highest mass peak corresponds to the parent ion or parent minus 147 (K) or 175 (R) • The “b” ions give the “normal” sequence • Both forward (b ion) and backward (y ion) sequences should be consistent • Use the resulting sequence tag to search the databases using BLAST (remember to use a high Expect value ~ 100) to see if the sequence matches something

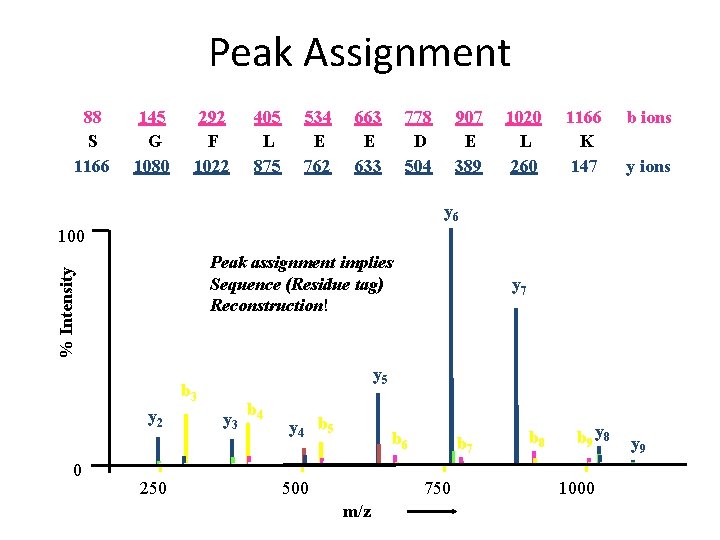
Peak Assignment 88 S 1166 145 G 1080 292 F 1022 405 L 875 534 E 762 663 E 633 778 D 504 907 E 389 1020 L 260 1166 K 147 b ions y 6 100 % Intensity Peak assignment implies Sequence (Residue tag) Reconstruction! [M+2 H]2+ y 5 b 3 y 2 0 250 y 7 y 3 b 4 y 4 b 5 b 6 500 b 7 750 m/z b 8 b 9 y 8 1000 y 9


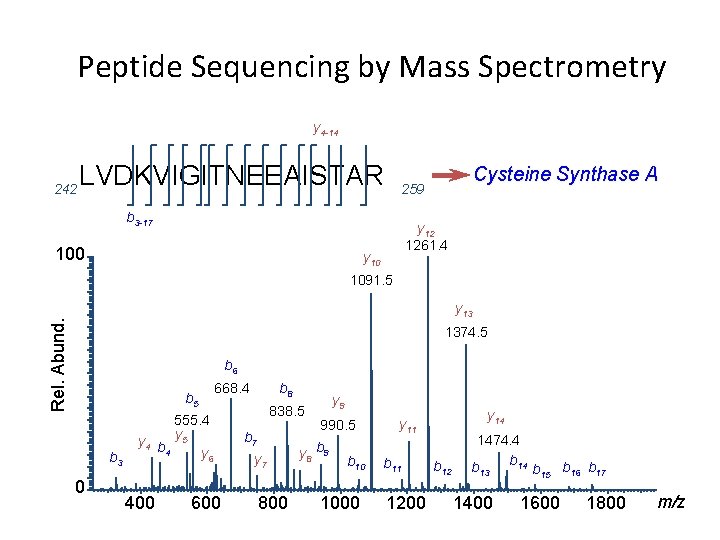
Peptide Sequencing by Mass Spectrometry y 4 -14 242 LVDKVIGITNEEAISTAR b 3 -17 100 Cysteine Synthase A 259 y 12 1261. 4 y 10 1091. 5 Rel. Abund. y 13 1374. 5 b 6 b 5 b 3 0 668. 4 555. 4 y 4 b y 5 4 y 6 400 600 b 8 838. 5 b 7 y 7 800 y 9 990. 5 y 8 b 9 b 10 1000 y 14 y 11 b 11 1200 b 12 1474. 4 b 13 1400 b 15 b 16 b 17 1600 1800 m/z

“+2” spectra

“bad” spectra

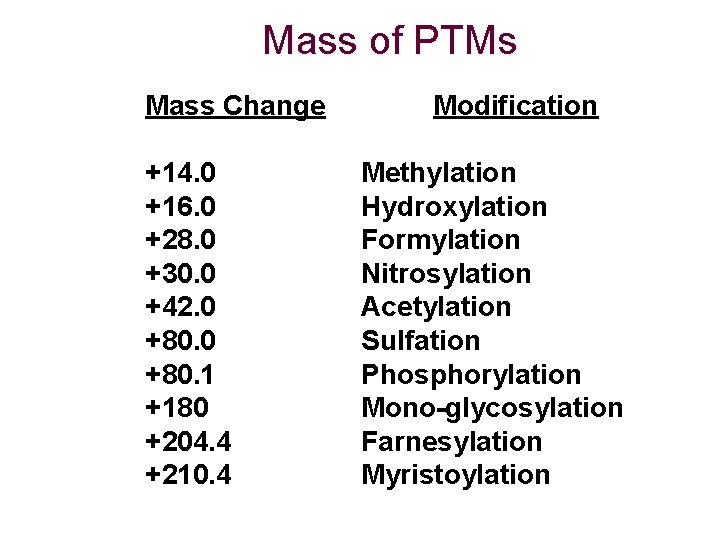
Modifications Some residues may be modified during the sample preparation procedure This introduces discrepancies in the expected and observed masses For example, Met residues are often oxidized

Mass of PTMs Mass Change +14. 0 +16. 0 +28. 0 +30. 0 +42. 0 +80. 1 +180 +204. 4 +210. 4 Modification Methylation Hydroxylation Formylation Nitrosylation Acetylation Sulfation Phosphorylation Mono-glycosylation Farnesylation Myristoylation

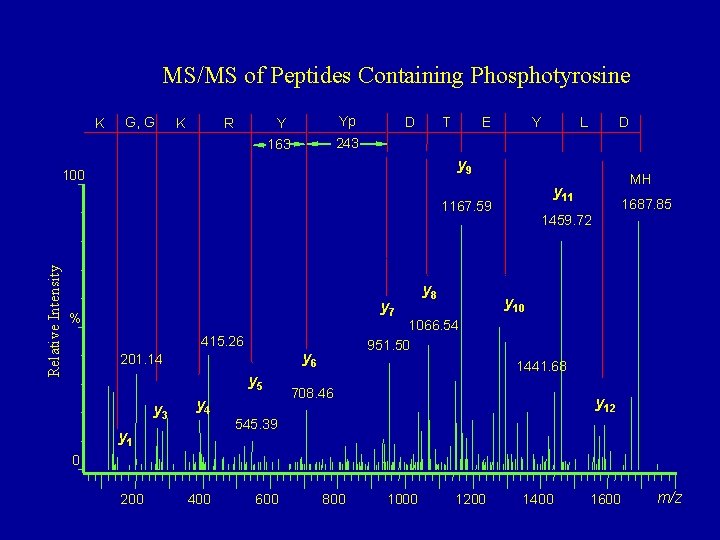
MS/MS of Peptides Containing Phosphotyrosine K G, G K R Y Yp 163 243 D T E Y D y 9 100 y 7 % 415. 26 y 5 y 3 y 4 y 1 1687. 85 1459. 72 y 10 1066. 54 951. 50 y 6 201. 14 y 8 MH y 11 1167. 59 Relative Intensity L 1441. 68 708. 46 y 12 545. 39 0 200 400 600 800 1000 1200 1400 1600 m/z