Rosetta Energy Function Glenn Butterfoss Rosetta Energy Function

Rosetta Energy Function Glenn Butterfoss

Rosetta Energy Function Major Classes: 1. Low resolution: Reduced atom representation Simple energy function Aggressively search conformational space 2. High resolution: Full atom More sophisticated energy function “Local” search of conformational (and sequence) space

Low resolution: Atom Model centroid reduction of side chains Energy function terms van der Waals repulsion “pair” terms (electrostatics) residue environment (prob of burial) 2º structure pairing terms (H-bonds) radius of gyration packing density Implicit terms fragments (local interactions) Rosetta Energy Function

Rosetta Energy Function Low resolution: Atom Model centroid reduction of side chains Energy function terms van der Waals repulsion In general … Weighted linear combination Energy = w 1*term 1 + w 2*term 2 + … Pair-wise decomposable “pair” terms (electrostatics) residue environment (prob of burial) 2º structure pairing terms (H-bonds) radius of gyration packing density Implicit terms fragments (local interactions) Heavily trained on PDB statistics Discriminate “near native” vs “non native” No single low resolution score Several functions with different weights
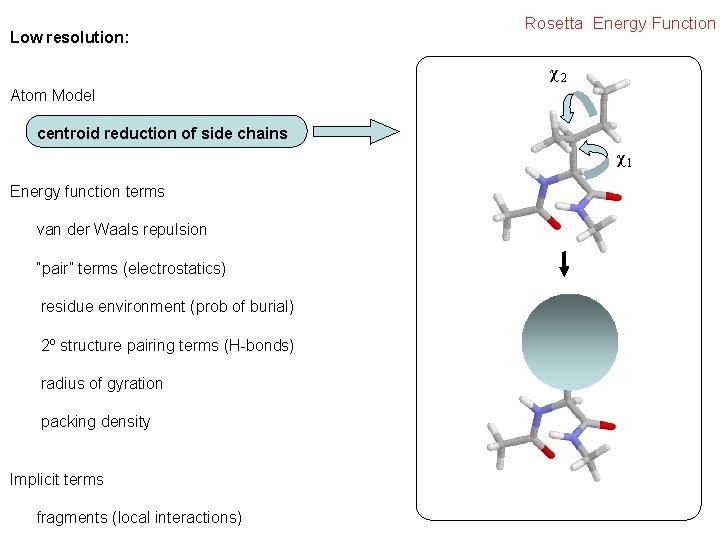
Low resolution: Rosetta Energy Function χ2 Atom Model centroid reduction of side chains χ1 Energy function terms van der Waals repulsion “pair” terms (electrostatics) residue environment (prob of burial) 2º structure pairing terms (H-bonds) radius of gyration packing density Implicit terms fragments (local interactions)
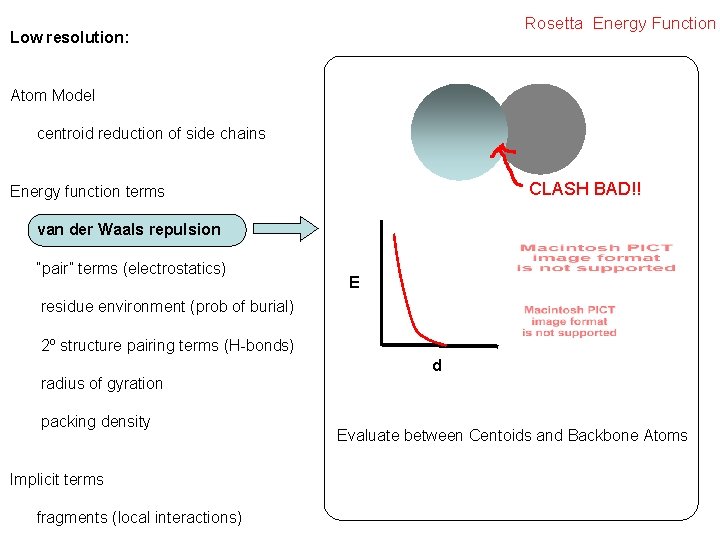
Rosetta Energy Function Low resolution: Atom Model centroid reduction of side chains CLASH BAD!! Energy function terms van der Waals repulsion “pair” terms (electrostatics) E residue environment (prob of burial) 2º structure pairing terms (H-bonds) d radius of gyration packing density Implicit terms fragments (local interactions) Evaluate between Centoids and Backbone Atoms
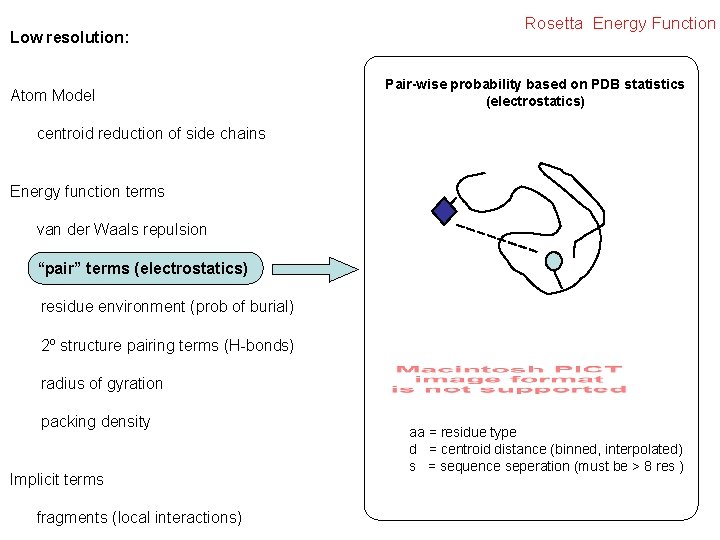
Low resolution: Atom Model Rosetta Energy Function Pair-wise probability based on PDB statistics (electrostatics) centroid reduction of side chains Energy function terms van der Waals repulsion “pair” terms (electrostatics) residue environment (prob of burial) 2º structure pairing terms (H-bonds) radius of gyration packing density Implicit terms fragments (local interactions) aa = residue type d = centroid distance (binned, interpolated) s = sequence seperation (must be > 8 res )
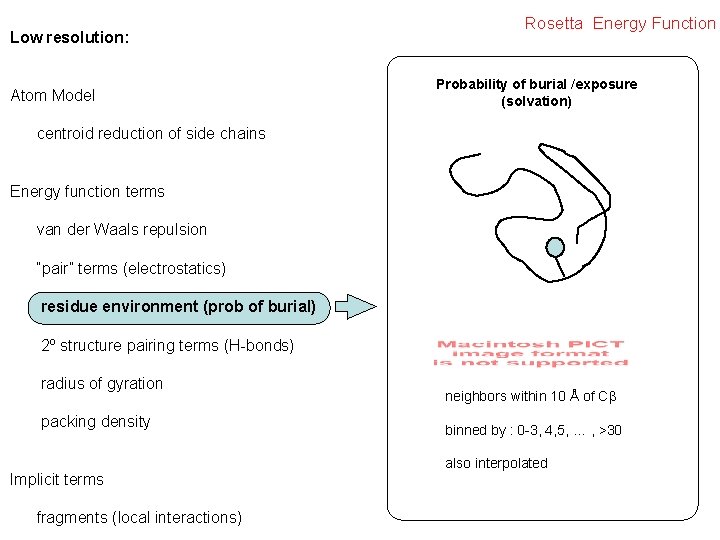
Low resolution: Atom Model Rosetta Energy Function Probability of burial /exposure (solvation) centroid reduction of side chains Energy function terms van der Waals repulsion “pair” terms (electrostatics) residue environment (prob of burial) 2º structure pairing terms (H-bonds) radius of gyration packing density Implicit terms fragments (local interactions) neighbors within 10 Å of C binned by : 0 -3, 4, 5, … , >30 also interpolated
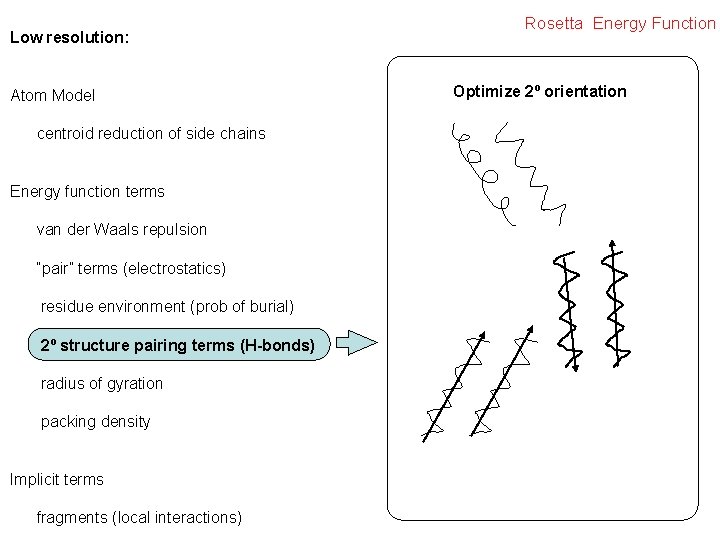
Low resolution: Atom Model centroid reduction of side chains Energy function terms van der Waals repulsion “pair” terms (electrostatics) residue environment (prob of burial) 2º structure pairing terms (H-bonds) radius of gyration packing density Implicit terms fragments (local interactions) Rosetta Energy Function Optimize 2º orientation
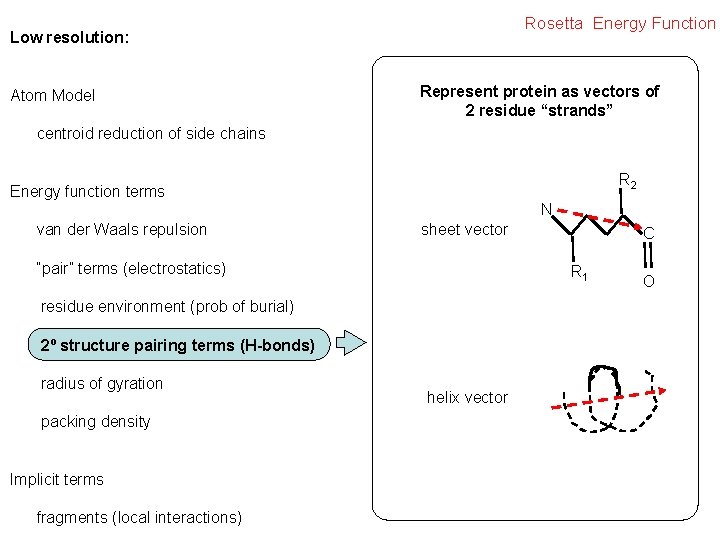
Rosetta Energy Function Low resolution: Atom Model Represent protein as vectors of 2 residue “strands” centroid reduction of side chains R 2 Energy function terms van der Waals repulsion N sheet vector “pair” terms (electrostatics) R 1 residue environment (prob of burial) 2º structure pairing terms (H-bonds) radius of gyration packing density Implicit terms fragments (local interactions) C helix vector O
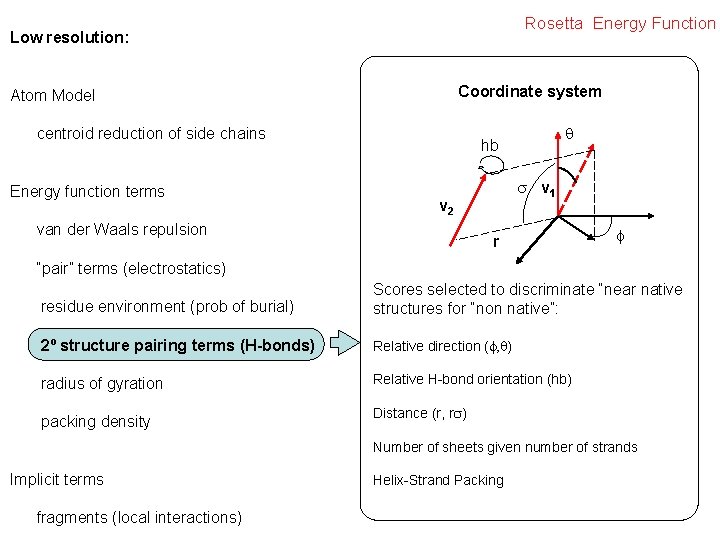
Rosetta Energy Function Low resolution: Coordinate system Atom Model centroid reduction of side chains Energy function terms hb v 1 v 2 van der Waals repulsion r “pair” terms (electrostatics) residue environment (prob of burial) Scores selected to discriminate “near native structures for “non native”: 2º structure pairing terms (H-bonds) Relative direction ( ) radius of gyration Relative H-bond orientation (hb) packing density Distance (r, r Number of sheets given number of strands Implicit terms fragments (local interactions) Helix-Strand Packing
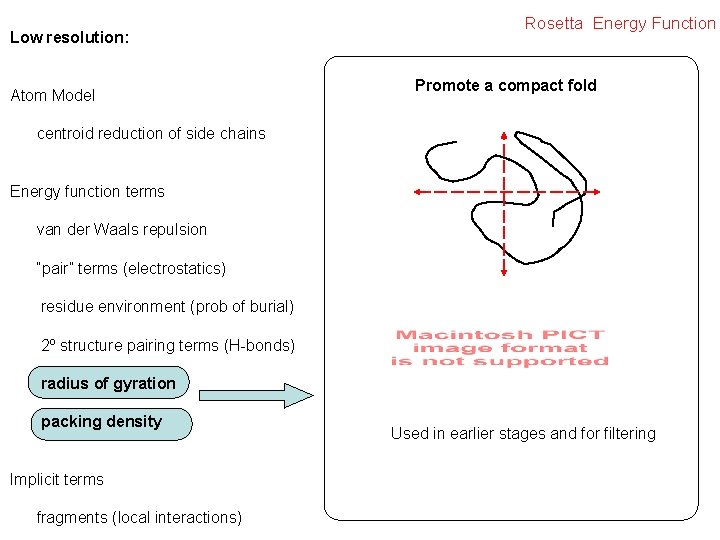
Low resolution: Atom Model Rosetta Energy Function Promote a compact fold centroid reduction of side chains Energy function terms van der Waals repulsion “pair” terms (electrostatics) residue environment (prob of burial) 2º structure pairing terms (H-bonds) radius of gyration packing density Implicit terms fragments (local interactions) Used in earlier stages and for filtering
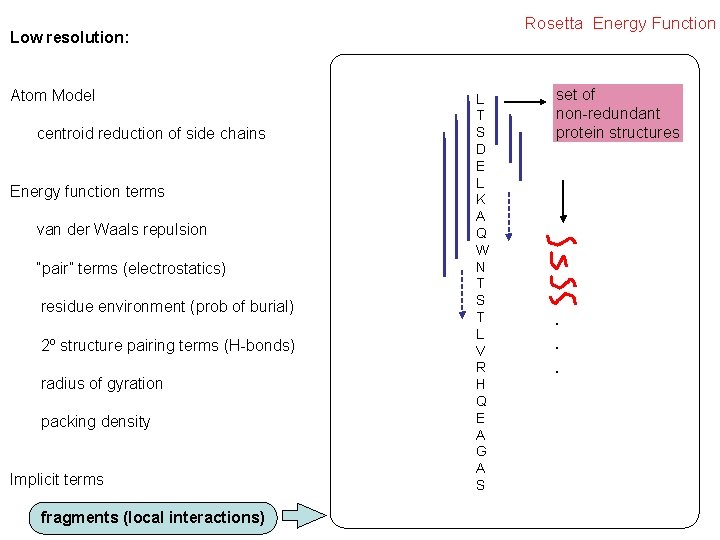
Rosetta Energy Function Low resolution: Atom Model centroid reduction of side chains Energy function terms van der Waals repulsion “pair” terms (electrostatics) residue environment (prob of burial) 2º structure pairing terms (H-bonds) radius of gyration packing density Implicit terms fragments (local interactions) L T S D E L K A Q W N T S T L V R H Q E A G A S set of non-redundant protein structures . . .
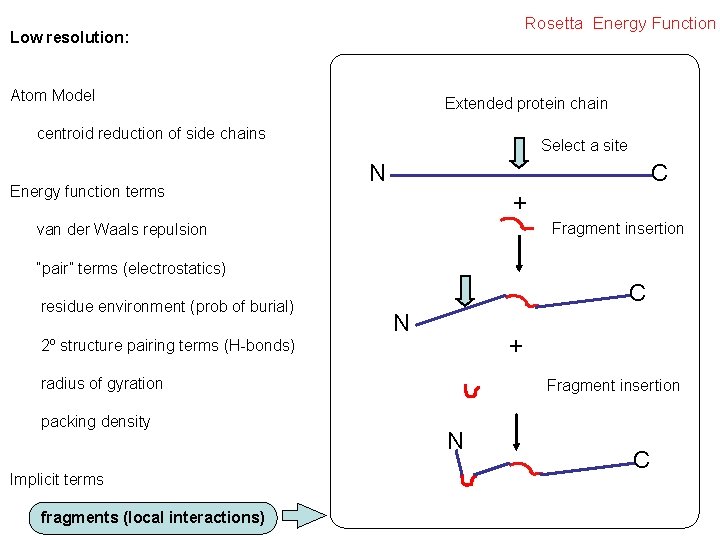
Rosetta Energy Function Low resolution: Atom Model Extended protein chain centroid reduction of side chains Energy function terms Select a site N C + Fragment insertion van der Waals repulsion “pair” terms (electrostatics) residue environment (prob of burial) C N + 2º structure pairing terms (H-bonds) radius of gyration packing density Implicit terms fragments (local interactions) Fragment insertion N C
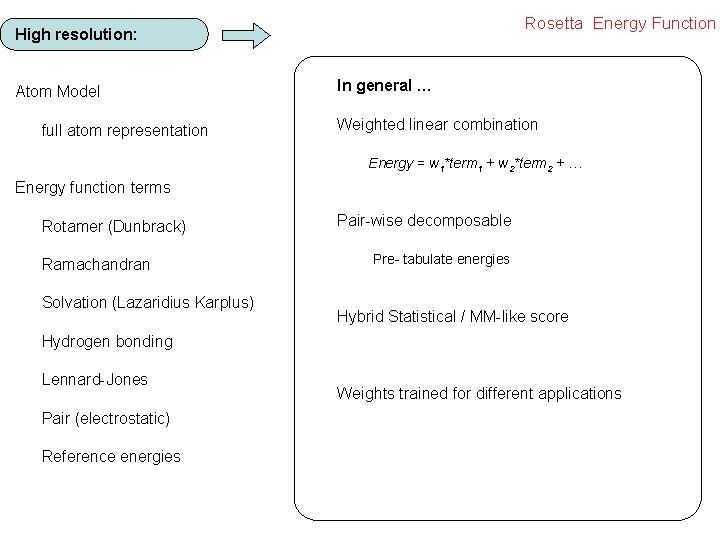
Rosetta Energy Function High resolution: Atom Model full atom representation In general … Weighted linear combination Energy = w 1*term 1 + w 2*term 2 + … Energy function terms Rotamer (Dunbrack) Ramachandran Solvation (Lazaridius Karplus) Pair-wise decomposable Pre- tabulate energies Hybrid Statistical / MM-like score Hydrogen bonding Lennard-Jones Pair (electrostatic) Reference energies Weights trained for different applications
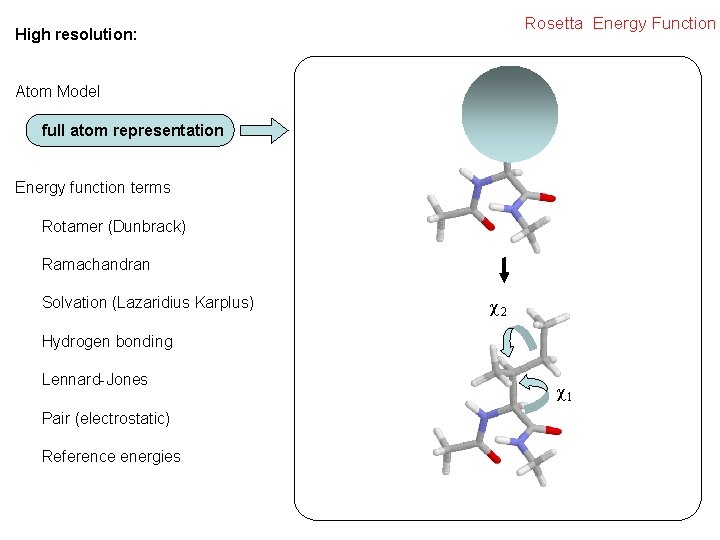
Rosetta Energy Function High resolution: Atom Model full atom representation Energy function terms Rotamer (Dunbrack) Ramachandran Solvation (Lazaridius Karplus) χ2 Hydrogen bonding Lennard-Jones Pair (electrostatic) Reference energies χ1
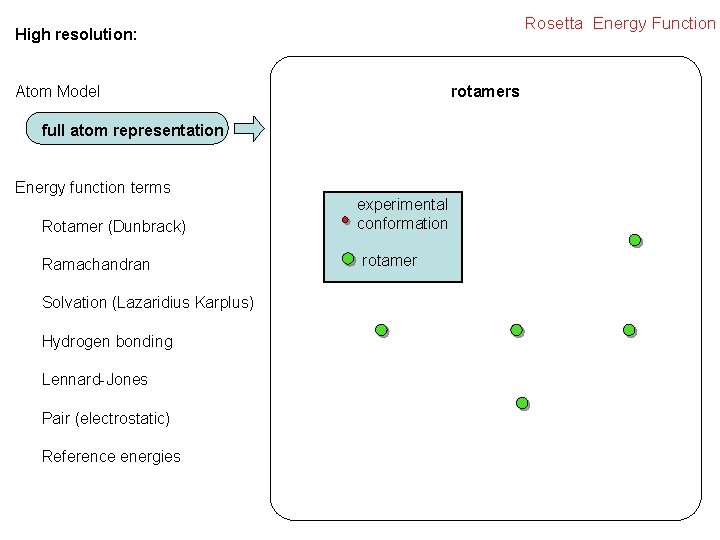
Rosetta Energy Function High resolution: rotamers Atom Model full atom representation Energy function terms Rotamer (Dunbrack) Ramachandran Solvation (Lazaridius Karplus) Hydrogen bonding Lennard-Jones Pair (electrostatic) Reference energies experimental conformation rotamer
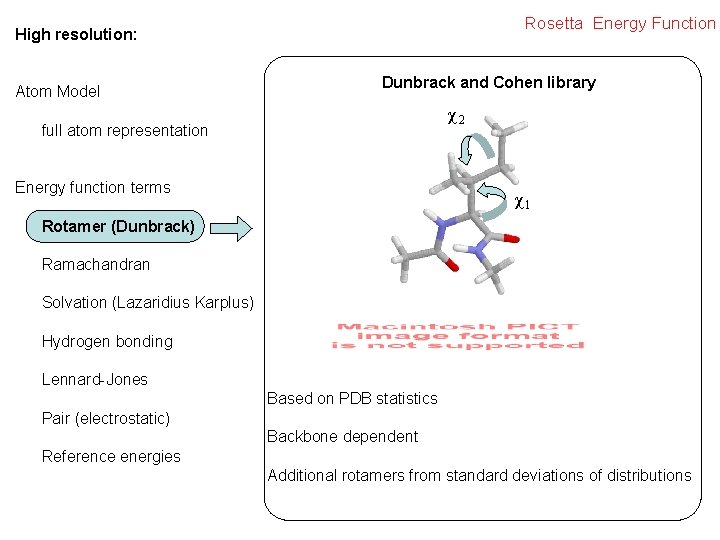
Rosetta Energy Function High resolution: Atom Model Dunbrack and Cohen library χ2 full atom representation Energy function terms χ1 Rotamer (Dunbrack) Ramachandran Solvation (Lazaridius Karplus) Hydrogen bonding Lennard-Jones Based on PDB statistics Pair (electrostatic) Backbone dependent Reference energies Additional rotamers from standard deviations of distributions
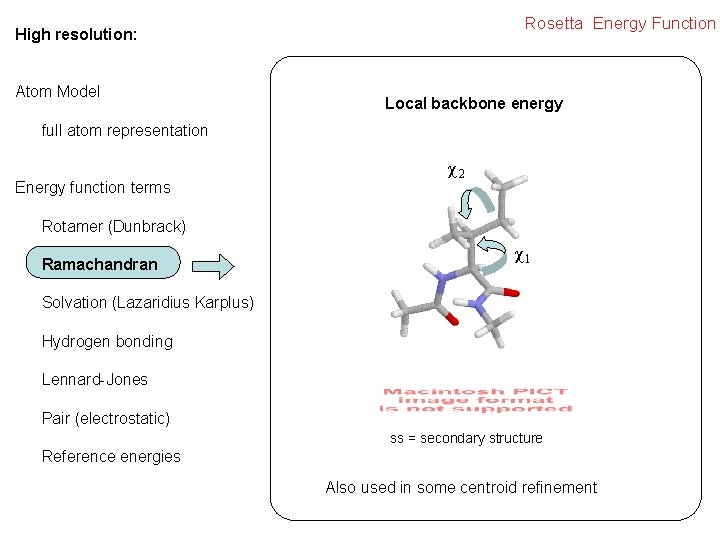
Rosetta Energy Function High resolution: Atom Model Local backbone energy full atom representation Energy function terms χ2 Rotamer (Dunbrack) Ramachandran χ1 Solvation (Lazaridius Karplus) Hydrogen bonding Lennard-Jones Pair (electrostatic) ss = secondary structure Reference energies Also used in some centroid refinement
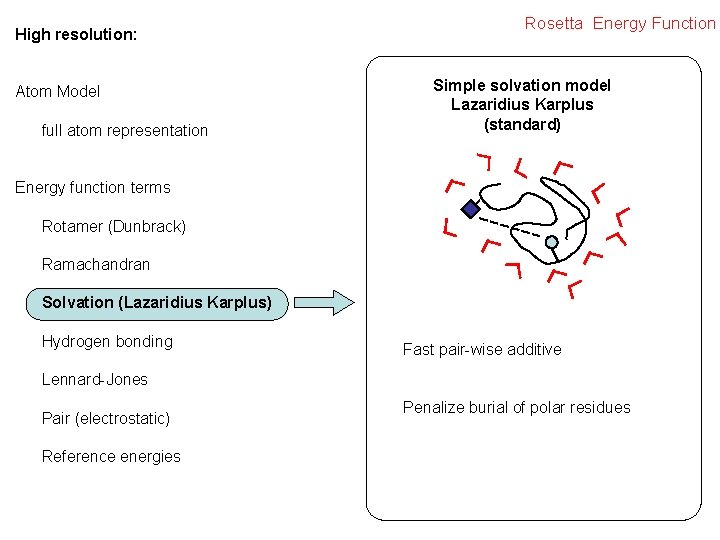
High resolution: Atom Model full atom representation Rosetta Energy Function Simple solvation model Lazaridius Karplus (standard) Energy function terms Rotamer (Dunbrack) Ramachandran Solvation (Lazaridius Karplus) Hydrogen bonding Fast pair-wise additive Lennard-Jones Pair (electrostatic) Reference energies Penalize burial of polar residues
High resolution: Atom Model Rosetta Energy Function Simple solvation model (Special cases) full atom representation Energy function terms Rotamer (Dunbrack) Ramachandran Solvation (Lazaridius Karplus) Hydrogen bonding Protein-DNA interactions: Generalized Born Lennard-Jones Pair (electrostatic) Reference energies Protein-Ligand: Coulomb
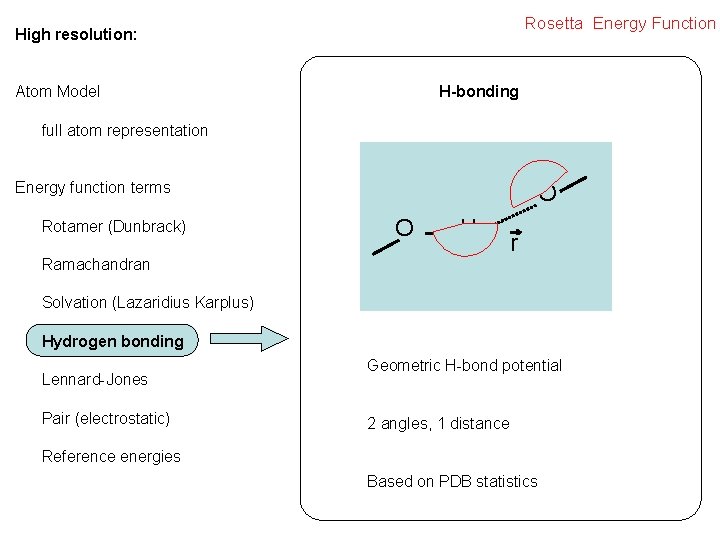
Rosetta Energy Function High resolution: H-bonding Atom Model full atom representation Energy function terms Rotamer (Dunbrack) O O H Ramachandran r Solvation (Lazaridius Karplus) Hydrogen bonding Lennard-Jones Pair (electrostatic) Geometric H-bond potential 2 angles, 1 distance Reference energies Based on PDB statistics
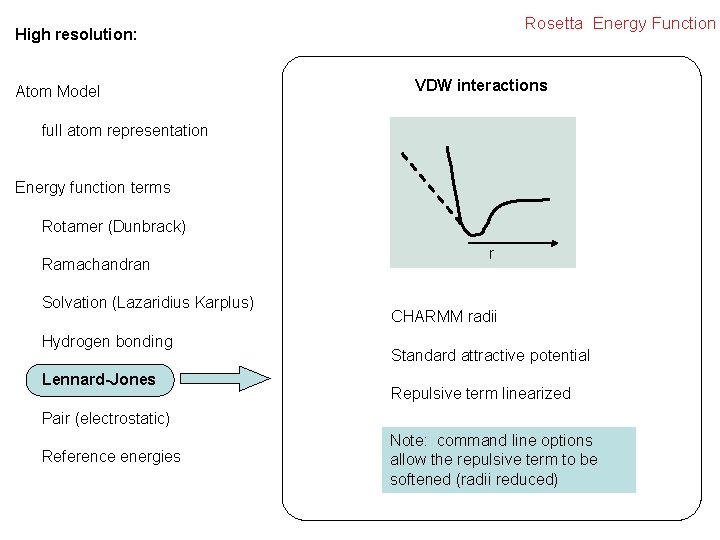
Rosetta Energy Function High resolution: Atom Model VDW interactions full atom representation Energy function terms Rotamer (Dunbrack) Ramachandran Solvation (Lazaridius Karplus) Hydrogen bonding Lennard-Jones r CHARMM radii Standard attractive potential Repulsive term linearized Pair (electrostatic) Reference energies Note: command line options allow the repulsive term to be softened (radii reduced)
Rosetta Energy Function High resolution: Atom Model Electrostatics full atom representation Energy function terms Rotamer (Dunbrack) Ramachandran Solvation (Lazaridius Karplus) Hydrogen bonding Probability of finding residue types at give in distance Lennard-Jones Pair (electrostatic) Reference energies Defined by C coordinates

High resolution: Atom Model Rosetta Energy Function Correction for “folding” full atom representation Energy function terms Rotamer (Dunbrack) Ramachandran Solvation (Lazaridius Karplus) Hydrogen bonding Lennard-Jones Pair (electrostatic) Reference energies Unique “cost” for designing in each residue type G for bringing residue type into folded protein Optimized with sequence recovery trials of folded protein structures
Rosetta Energy Function Xavier Thanks Rosetta Community
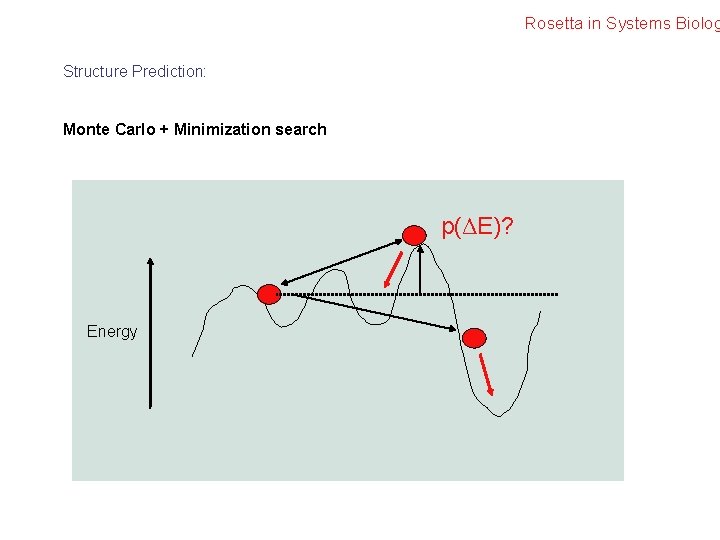
Rosetta in Systems Biolog Structure Prediction: Monte Carlo + Minimization search p(ΔE)? Energy

Rosetta in Systems Biolog Protein Design: Protocol: Packing: Pre-tabulate table of all pair-wise rotamer energies Monte Carlo search through rotamer / sequence space With docking and backbone movement: Iterate packing with (as above) with backbone / rigid body movements Possibly apply restraints docking, rmsd, disulfide, …
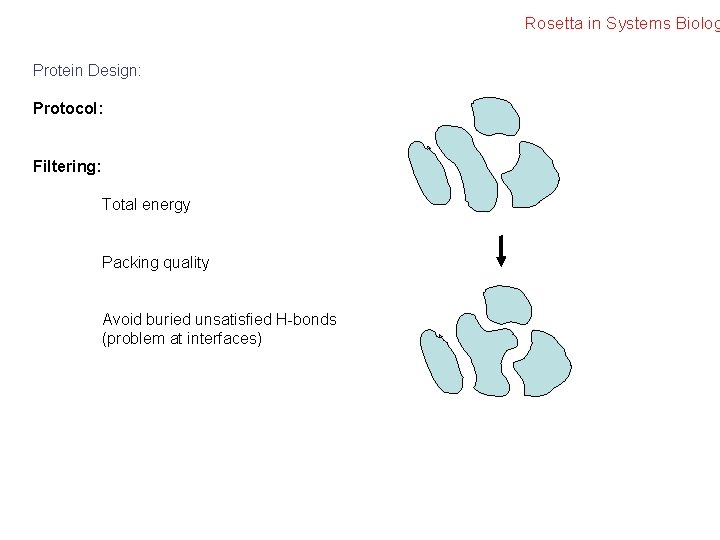
Rosetta in Systems Biolog Protein Design: Protocol: Filtering: Total energy Packing quality Avoid buried unsatisfied H-bonds (problem at interfaces)
Low resolution: Atom Model centroid reduction of side chains Energy function terms van der Waals repulsion “pair” terms (electrostatics) residue environment (prob of burial) 2º structure pairing terms (H-bonds) radius of gyration packing density Implicit terms fragments (local interactions) Rosetta Energy Function
- Slides: 30