Role of microRNAs in Atrial Fibrillation Amir Shaikh

Role of micro-RNAs in Atrial Fibrillation Amir Shaikh, MD; David D Mc. Manus, MD, Sc. M Assistant Professor, Department of Medicine University of Massachusetts Medical School, Worcester, MA, USA

Disclosures • David D Mc. Manus, MD, Sc. M has received research funding from: – – – – – US Department of Defense National Heart Lung and Blood Institute Worcester Polytechnic Institute (New Technology Development Grant) St. Jude Medical Philips Healthcare Sanofi Aventis Biotronik Otsuka Pharmaceuticals Astra Zeneca

University of Massachusetts Medical School
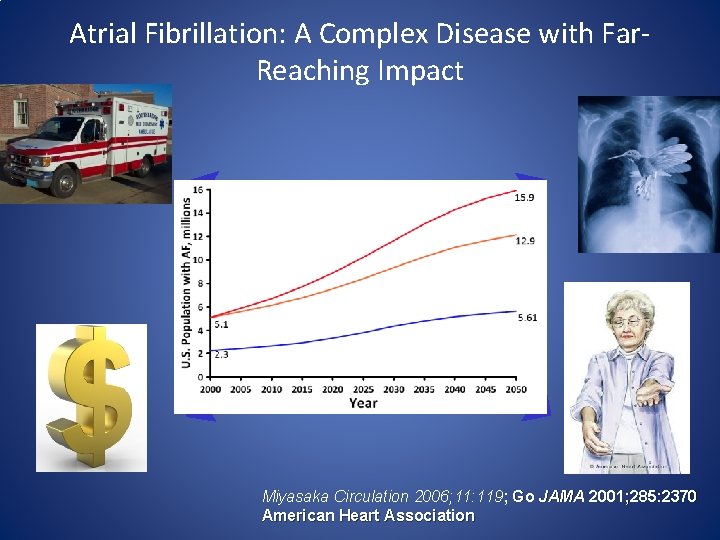
Atrial Fibrillation: A Complex Disease with Far. Reaching Impact Miyasaka Circulation 2006; 11: 119; Go JAMA 2001; 285: 2370 American Heart Association
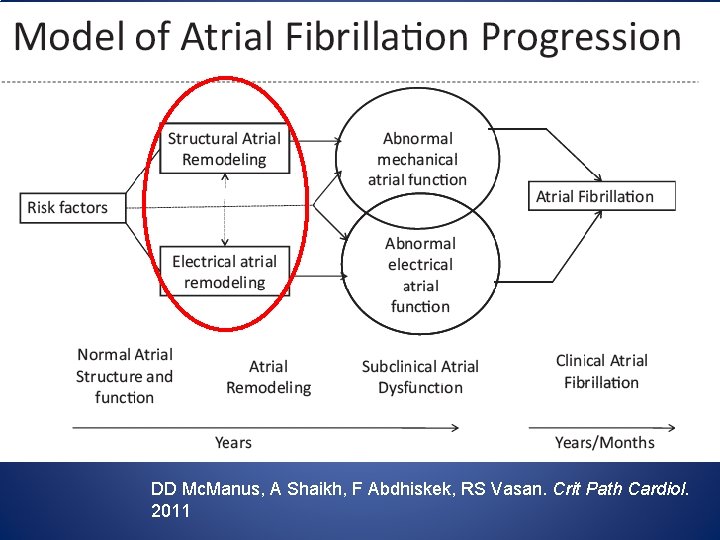
A useful phenotype to explore genetic and transcriptomic underpinnings of AF? DD Mc. Manus, A Shaikh, F Abdhiskek, RS Vasan. Crit Path Cardiol. 2011

Focal Triggers Initiate AF and Reentry Perpetuates It AF requires both a trigger and a vulnerable substrate Interplay between intrinsic susceptibility and exposures largely unknown Ding Sheng He, MD, Ph. D

Paroxysmal Persistent Permanent Although all are susceptible to AF, why do many Substrate individuals develop it early in life for AF with minimal (if any) exposures? Initiation substrate Why do some progress to more persistent forms. Triggers of the of AF arrhythmia? AF disease progression
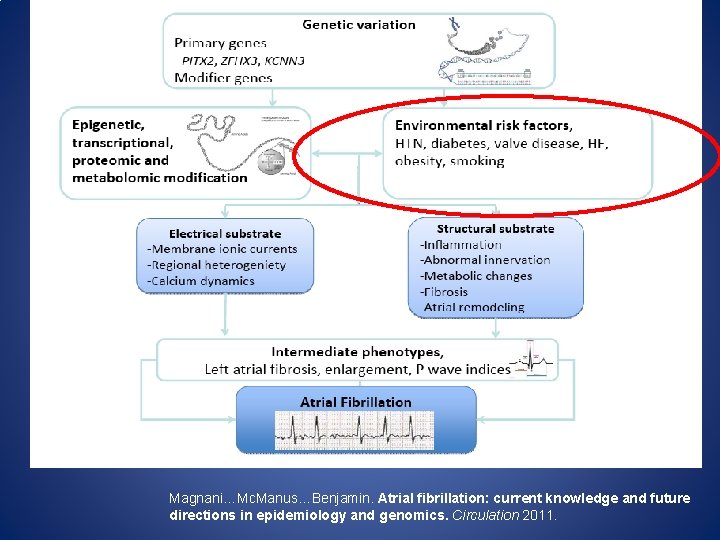
Magnani…Mc. Manus…Benjamin. Atrial fibrillation: current knowledge and future directions in epidemiology and genomics. Circulation 2011.
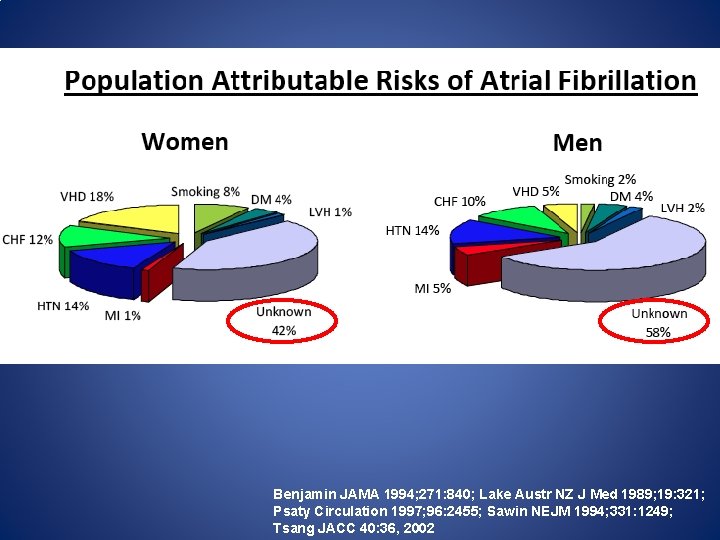
Benjamin JAMA 1994; 271: 840; Lake Austr NZ J Med 1989; 19: 321; Psaty Circulation 1997; 96: 2455; Sawin NEJM 1994; 331: 1249; Tsang JACC 40: 36, 2002
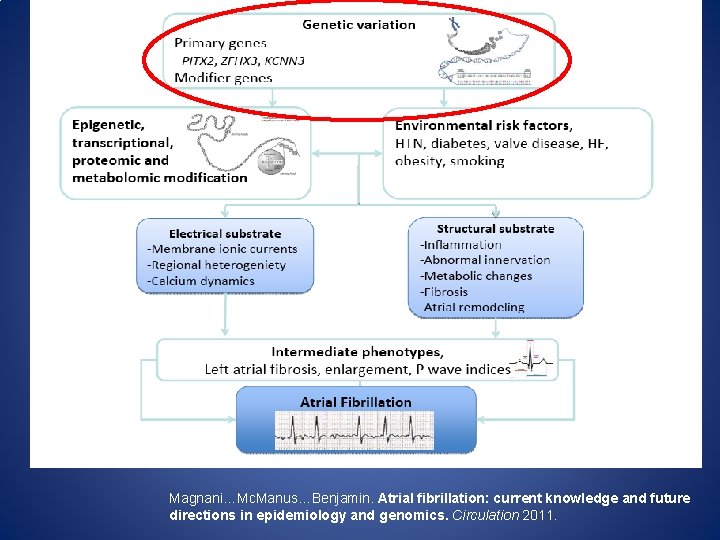
Magnani…Mc. Manus…Benjamin. Atrial fibrillation: current knowledge and future directions in epidemiology and genomics. Circulation 2011.
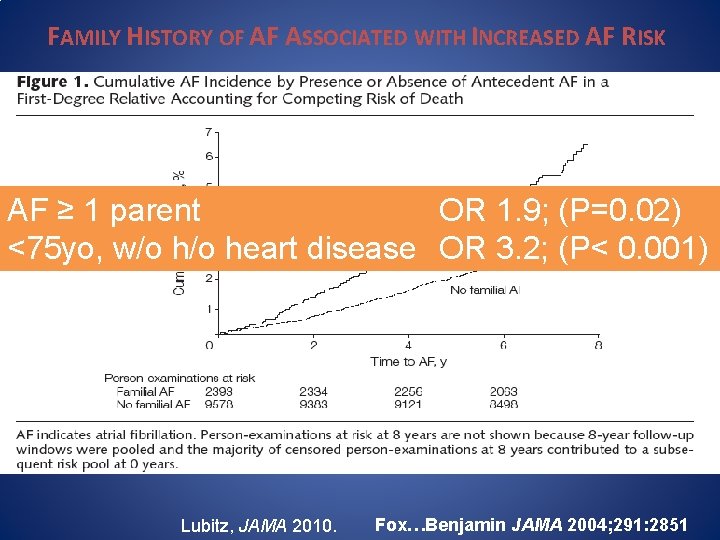
FAMILY HISTORY OF AF ASSOCIATED WITH INCREASED AF RISK Genetics of OR AF 1. 9; (P=0. 02) AF ≥ 1 parent <75 yo, w/o heart disease OR 3. 2; (P< 0. 001) • Association with Family History • Candidate Gene Studies • GWAS findings Lubitz, JAMA 2010. Fox…Benjamin JAMA 2004; 291: 2851
Genetics, Genomics and AF
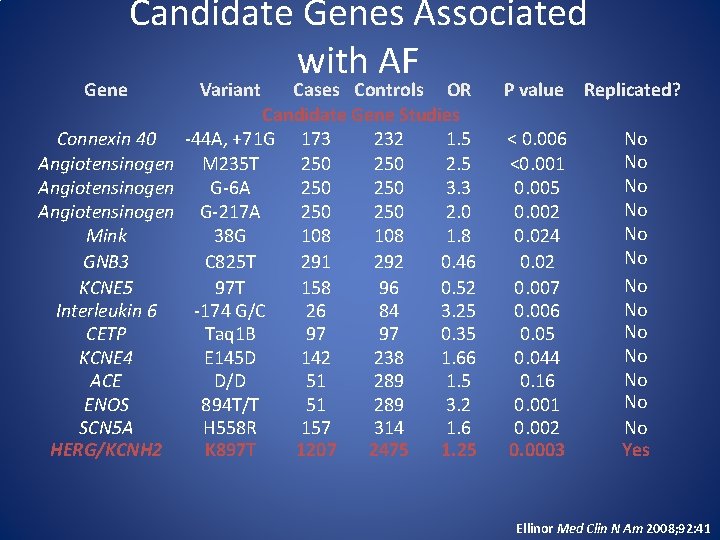
Gene Candidate Genes Associated with AF Variant Cases Controls OR Candidate Gene Studies Connexin 40 -44 A, +71 G 173 232 1. 5 Angiotensinogen M 235 T 250 2. 5 Angiotensinogen G-6 A 250 3. 3 Angiotensinogen G-217 A 250 2. 0 Mink 38 G 108 1. 8 GNB 3 C 825 T 291 292 0. 46 KCNE 5 97 T 158 96 0. 52 Interleukin 6 -174 G/C 26 84 3. 25 CETP Taq 1 B 97 97 0. 35 KCNE 4 E 145 D 142 238 1. 66 ACE D/D 51 289 1. 5 ENOS 894 T/T 51 289 3. 2 SCN 5 A H 558 R 157 314 1. 6 HERG/KCNH 2 K 897 T 1207 2475 1. 25 P value Replicated? < 0. 006 <0. 001 0. 005 0. 002 0. 024 0. 02 0. 007 0. 006 0. 05 0. 044 0. 16 0. 001 0. 002 0. 0003 No No No No Yes Ellinor Med Clin N Am 2008; 92: 41
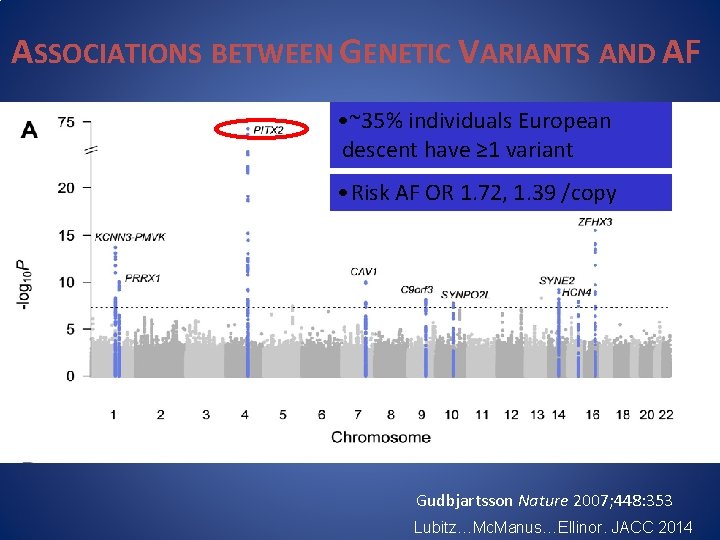
ASSOCIATIONS BETWEEN GENETIC VARIANTS AND AF • ~35% individuals European descent have ≥ 1 variant • Risk AF OR 1. 72, 1. 39 /copy Gudbjartsson Nature 2007; 448: 353 Lubitz…Mc. Manus…Ellinor. JACC 2014
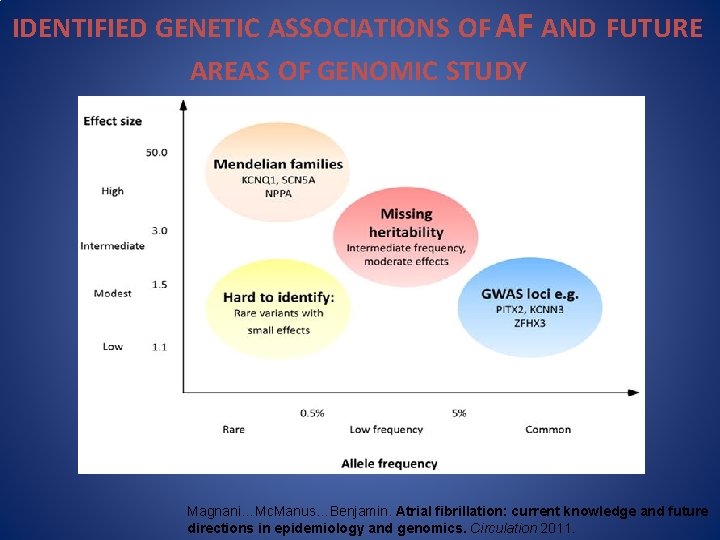
IDENTIFIED GENETIC ASSOCIATIONS OF AF AND FUTURE AREAS OF GENOMIC STUDY Magnani…Mc. Manus…Benjamin. Atrial fibrillation: current knowledge and future directions in epidemiology and genomics. Circulation 2011.
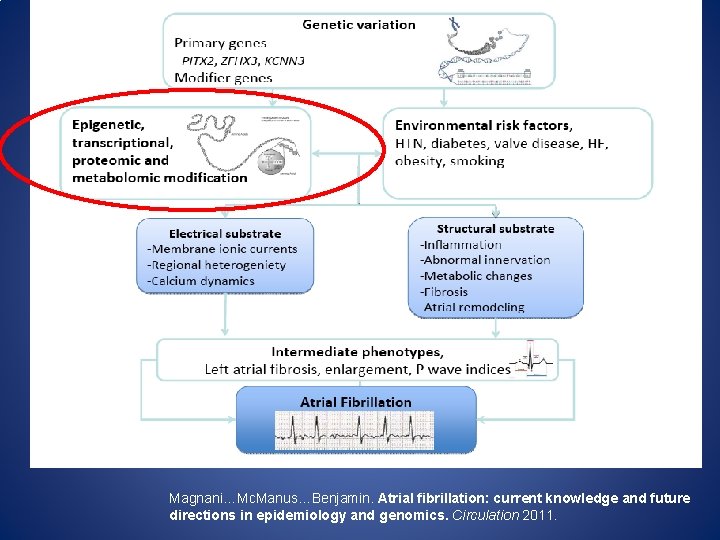
Magnani…Mc. Manus…Benjamin. Atrial fibrillation: current knowledge and future directions in epidemiology and genomics. Circulation 2011.

Heritability Gap in AF – Moving beyond GWAS Known unknowns in AF: • 40% AF risk unexplained by clinical CV Could variable gene expression in risk factors stress states explain heritability gap? • 2 -fold higher risk of AF in patients with family history of AF • 90+% of AF heritability unexplained by known SNPs and candidate gene studies • AF triggers contribute to altered atrial gene expression
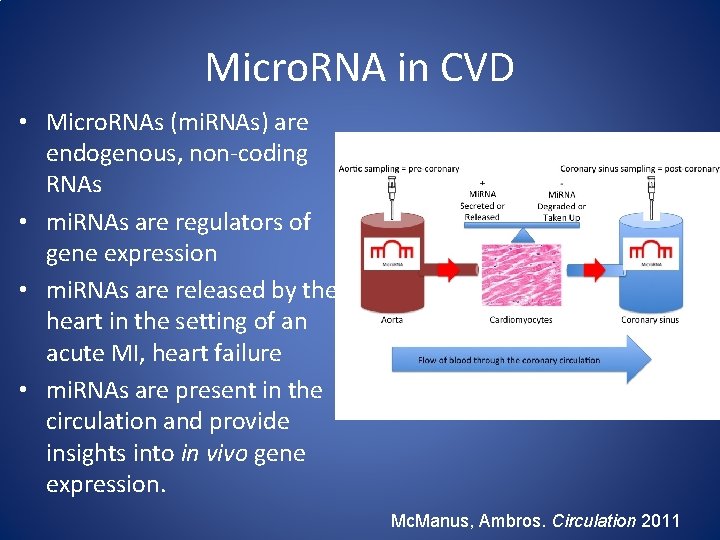
Micro. RNA in CVD • Micro. RNAs (mi. RNAs) are endogenous, non-coding RNAs • mi. RNAs are regulators of gene expression • mi. RNAs are released by the heart in the setting of an acute MI, heart failure • mi. RNAs are present in the circulation and provide insights into in vivo gene expression. Mc. Manus, Ambros. Circulation 2011
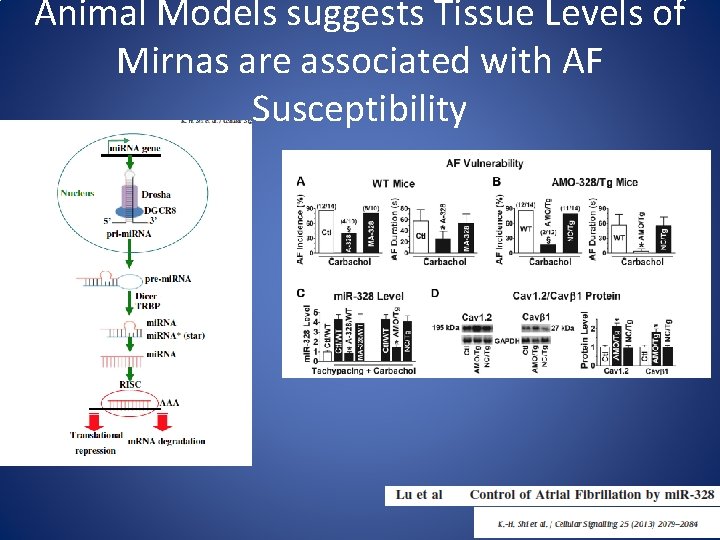
Animal Models suggests Tissue Levels of Mirnas are associated with AF Susceptibility
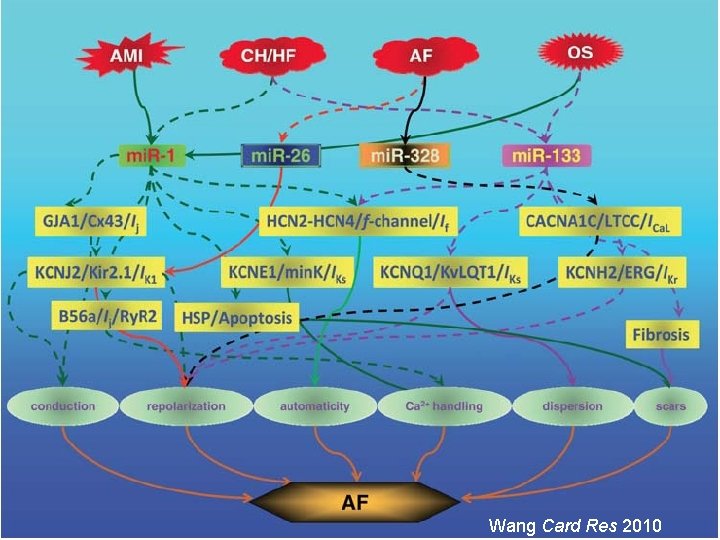
Wang Card Res 2010
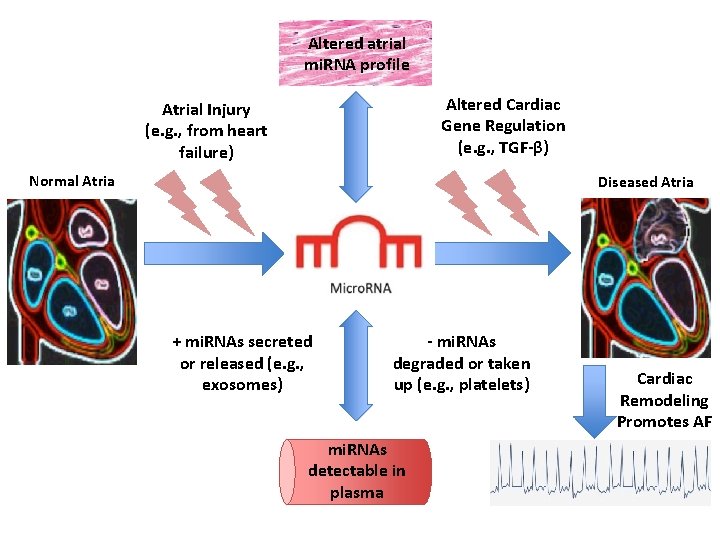
Altered atrial mi. RNA profile Altered Cardiac Gene Regulation (e. g. , TGF-β) Atrial Injury (e. g. , from heart failure) Normal Atria Diseased Atria + mi. RNAs secreted or released (e. g. , exosomes) - mi. RNAs degraded or taken up (e. g. , platelets) mi. RNAs detectable in plasma Cardiac Remodeling Promotes AF

High Throughput Technology exists to assess mi. RNA expression • High-throughput mi. RNA expression profiling systems allow rapid profiling of mi. RNAs from numerous samples • Use real-time PCR, or microarray • Primers correspond to over 1, 000 mi. RNAs • Accurate, specific and sensitive Courtesy, Jane Freedman, MD Kahraman Tanriverdi, Ph. D

• mi. R-328 is up-regulated in the atria of human subjects with AF • mi. R-328 regulates L-type Ca 2+ channel density, shortens the atrial effective refractory period • mi. R-328 enhances AF vulnerability in animal models Mc. Manus et al. Heart Rhythm 2014
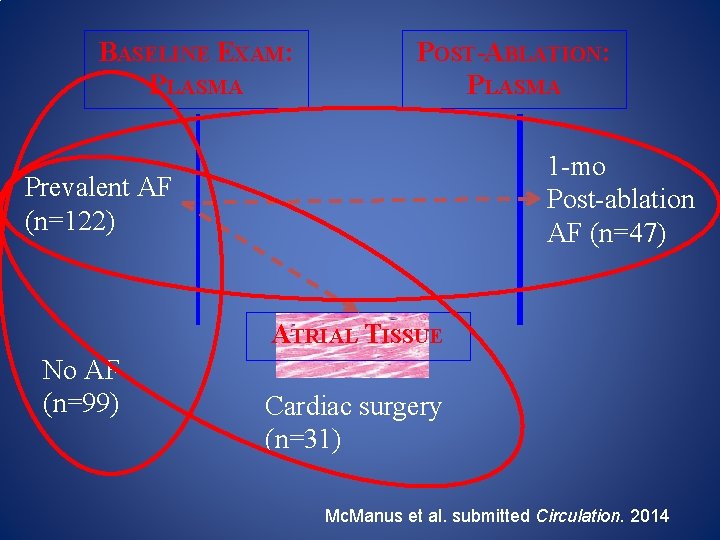
BASELINE EXAM: PLASMA POST-ABLATION: PLASMA 1 -mo Post-ablation AF (n=47) Prevalent AF (n=122) ATRIAL TISSUE No AF (n=99) Cardiac surgery (n=31) Mc. Manus et al. submitted Circulation. 2014
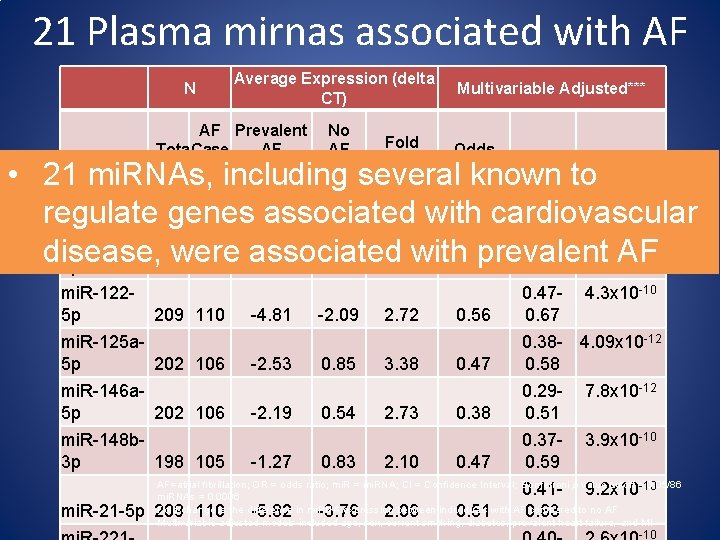
21 Plasma mirnas associated with AF mi. RNA Average Expression (delta Multivariable Adjusted*** CT) AF Prevalent No Fold AF AF Tota Case Odds Change l s (n=112) (n=99) Ratio 95% CI P-value* N • 21 mi. RNAs, including several known to mi. R-1500. 41 - 1. 5 x 10 5 p 206 107 -3. 26 -0. 96 2. 30 with 0. 51 cardiovascular 0. 63 regulate genes associated mi. R-1000. 33 - 3. 2 x 10 disease, were associated with prevalent AF 5 p 205 109 -1. 61 1. 45 3. 07 0. 42 0. 54 -10 -12 mi. R-1225 p 209 110 mi. R-125 a 5 p 202 106 mi. R-146 a 5 p 202 106 mi. R-148 b 3 p 198 105 -4. 81 -2. 53 -2. 19 -1. 27 -2. 09 0. 85 0. 54 0. 83 2. 72 3. 38 2. 73 2. 10 0. 56 0. 470. 67 4. 3 x 10 -10 0. 47 0. 38 - 4. 09 x 10 -12 0. 58 7. 8 x 10 -12 0. 38 0. 290. 51 3. 9 x 10 -10 0. 47 0. 370. 59 AF=atrial fibrillation; OR = odds ratio; mi. R = mi. RNA; CI = Confidence Interval; Bonferroni p value cutoff =-10 0. 05/86 mi. RNAs = 0. 0006 Fold-change is the difference in mi. RNA expression between individuals with AF compared to no AF Multivariable adjusted models included age, sex, current smoking, diabetes, prevalent heart failure, and MI mi. R-21 -5 p 209 110 -5. 82 -3. 76 2. 06 0. 51 0. 410. 63 9. 2 x 10 -10
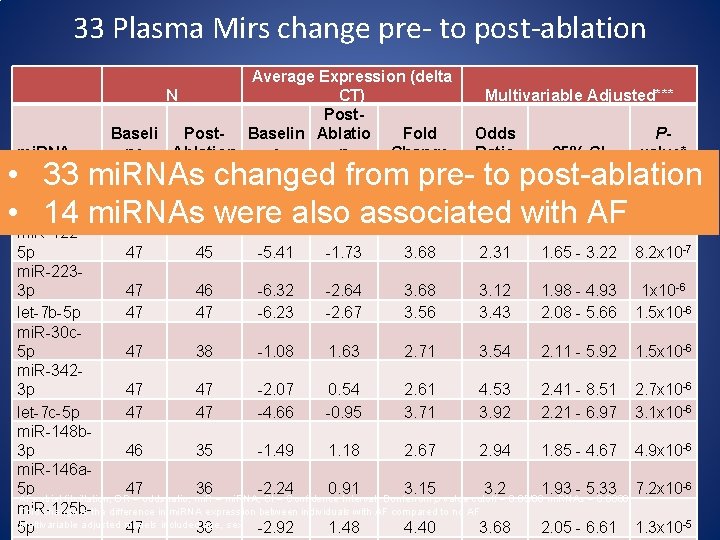
33 Plasma Mirs change pre- to post-ablation Average Expression (delta N CT) Post. Baseli Post- Baselin Ablatio Fold ne Ablation e n Change Multivariable Adjusted*** Odds Ratio mi. RNA 95% CI mi. R-1505 p 47 45 -3. 75 -0. 69 3. 06 2. 71 1. 85 - 3. 98 mi. R-21 -5 p 47 47 -6. 09 -2. 65 3. 44 3. 07 1. 98 - 4. 76 mi. R-1225 p 47 45 -5. 41 -1. 73 3. 68 2. 31 1. 65 - 3. 22 mi. R-2233 p 47 46 -6. 32 -2. 64 3. 68 3. 12 1. 98 - 4. 93 let-7 b-5 p 47 47 -6. 23 -2. 67 3. 56 3. 43 2. 08 - 5. 66 mi. R-30 c 5 p 47 38 -1. 08 1. 63 2. 71 3. 54 2. 11 - 5. 92 mi. R-3423 p 47 47 -2. 07 0. 54 2. 61 4. 53 2. 41 - 8. 51 let-7 c-5 p 47 47 -4. 66 -0. 95 3. 71 3. 92 2. 21 - 6. 97 mi. R-148 b 3 p 46 35 -1. 49 1. 18 2. 67 2. 94 1. 85 - 4. 67 mi. R-146 a 5 p 47 36 -2. 24 0. 91 3. 15 3. 2 1. 93 - 5. 33 AF=atrial fibrillation; OR = odds ratio; mi. R = mi. RNA; CI = Confidence Interval; Bonferroni p value cutoff = 0. 05/86 mi. RNAs = 0. 0006 mi. R-125 b. Fold-change is the difference in mi. RNA expression between individuals with AF compared to no AF Multivariable adjusted models included age, sex 5 p 47 38 -2. 92 1. 48 4. 40 3. 68 2. 05 - 6. 61 Pvalue* • 33 mi. RNAs changed from pre- to post-ablation 3. 6 x 10 • 14 mi. RNAs were also associated with AF 5. 3 x 10 -7 -7 8. 2 x 10 -7 1 x 10 -6 1. 5 x 10 -6 2. 7 x 10 -6 3. 1 x 10 -6 4. 9 x 10 -6 7. 2 x 10 -6 1. 3 x 10 -5
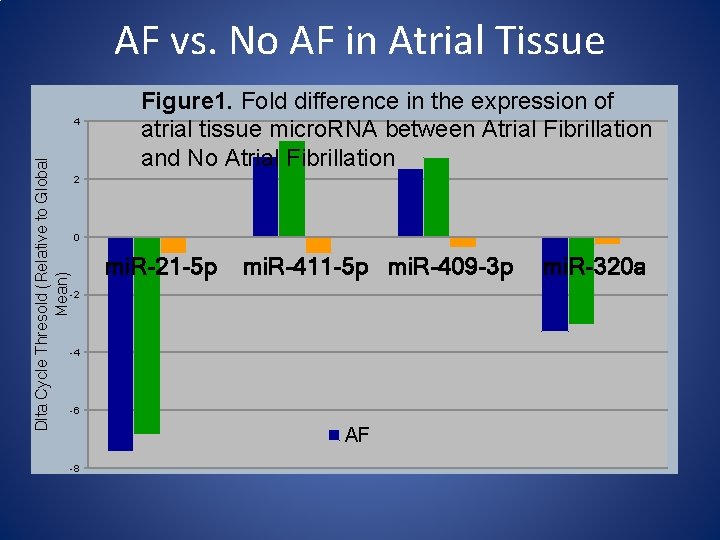
AF vs. No AF in Atrial Tissue Dlta Cycle Thresold (Relative to Global Mean) 4 Figure 1. Fold difference in the expression of atrial tissue micro. RNA between Atrial Fibrillation and No Atrial Fibrillation 2 0 mi. R-21 -5 p mi. R-411 -5 p mi. R-409 -3 p -2 -4 -6 AF -8 mi. R-320 a
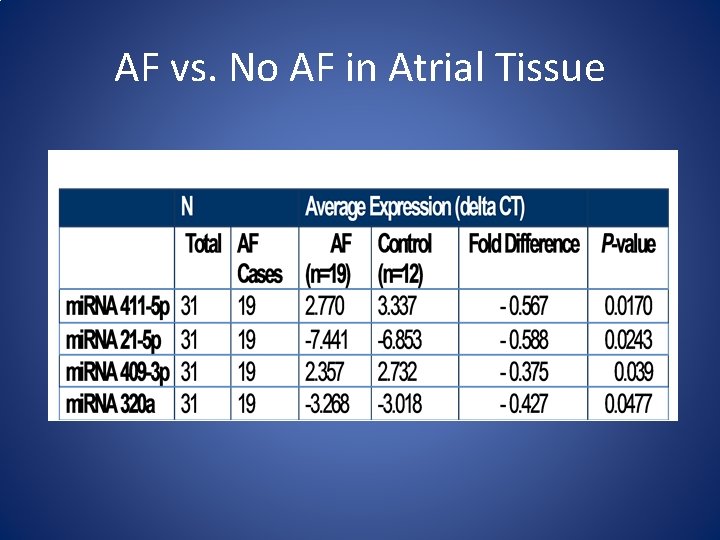
AF vs. No AF in Atrial Tissue
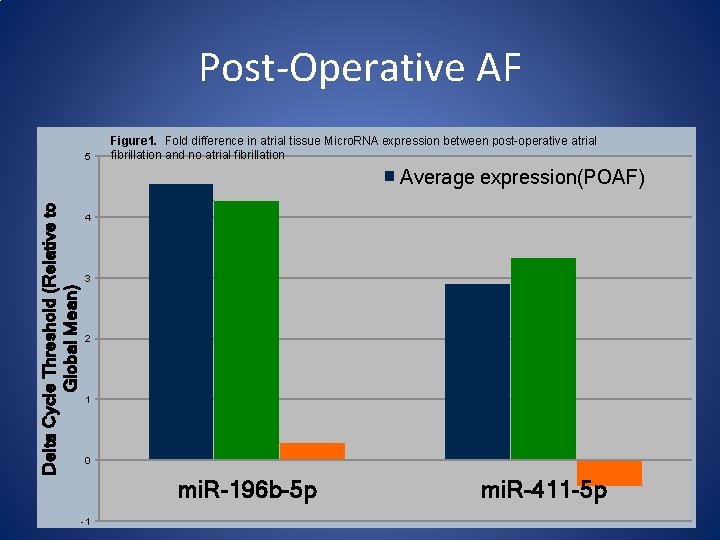
Post-Operative AF 5 Figure 1. Fold difference in atrial tissue Micro. RNA expression between post-operative atrial fibrillation and no atrial fibrillation Delta Cycle Threshold (Relative to Global Mean) Average expression(POAF) 4 3 2 1 0 mi. R-196 b-5 p -1 mi. R-411 -5 p

CONSIDERABLE OVERLAP IN HIGHLY VARIANT MIRS AND THOSE ASSOCIATED WITH AF AF vs. No AF mi. R-10 b-5 p mi. R-24 -3 p mi. R-29 a-3 p mi. R-99 b-5 p mi. R-221 -3 p mi. R-375 mi. R-411 -5 p Pre vs. Post. Ablation mi. R-21 -5 p mi. R-30 c-5 p mi. R-100 -5 p mi. R-122 -5 p mi. R-125 a-5 p mi. R-125 b-5 p mi. R-126 -3 p mi. R-146 a-5 p mi. R-148 b-3 p mi. R-150 -5 p mi. R-223 -3 p mi. R-342 -3 p let-7 b-5 p let-7 c-5 p mi. R-7 -5 p mi. R-221 -3 p mi. R-10 b-5 p mi. R-320 a mi. R-19 a-3 p mi. R-451 a mi. R-20 a-5 p mi. R-144 -3 p mi. R-24 -3 p mi. R-146 b-5 p mi. R-25 -3 p mi. R-29 b-3 p mi. R-26 a-5 p mi. R-29 a-3 p mi. R-30 a-5 p mi. R-92 a-3 p mi. R-106 b-5 p let-7 f-5 p let-7 g-5 p
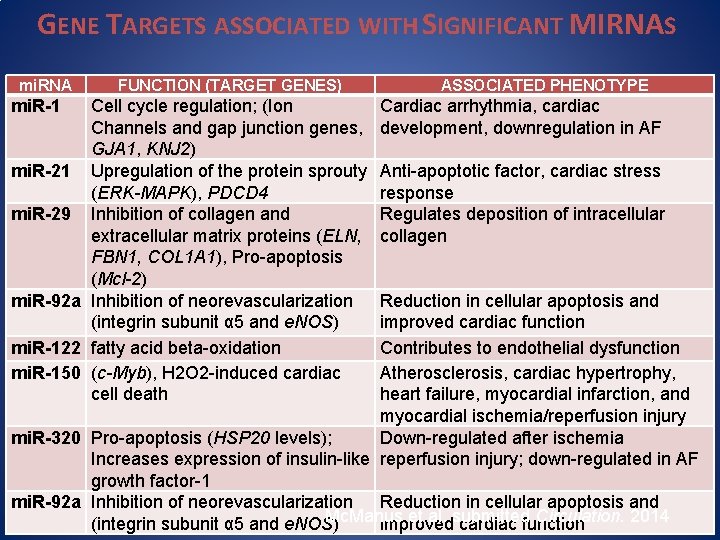
GENE TARGETS ASSOCIATED WITH SIGNIFICANT MIRNAS mi. RNA mi. R-1 FUNCTION (TARGET GENES) Cell cycle regulation; (Ion Channels and gap junction genes, GJA 1, KNJ 2) mi. R-21 Upregulation of the protein sprouty (ERK-MAPK), PDCD 4 mi. R-29 Inhibition of collagen and extracellular matrix proteins (ELN, FBN 1, COL 1 A 1), Pro-apoptosis (Mcl-2) mi. R-92 a Inhibition of neorevascularization (integrin subunit α 5 and e. NOS) mi. R-122 fatty acid beta-oxidation mi. R-150 (c-Myb), H 2 O 2 -induced cardiac cell death ASSOCIATED PHENOTYPE Cardiac arrhythmia, cardiac development, downregulation in AF Anti-apoptotic factor, cardiac stress response Regulates deposition of intracellular collagen Reduction in cellular apoptosis and improved cardiac function Contributes to endothelial dysfunction Atherosclerosis, cardiac hypertrophy, heart failure, myocardial infarction, and myocardial ischemia/reperfusion injury mi. R-320 Pro-apoptosis (HSP 20 levels); Down-regulated after ischemia Increases expression of insulin-like reperfusion injury; down-regulated in AF growth factor-1 mi. R-92 a Inhibition of neorevascularization Reduction in cellular apoptosis and Mc. Manus et al. submitted Circulation. 2014 (integrin subunit α 5 and e. NOS) improved cardiac function
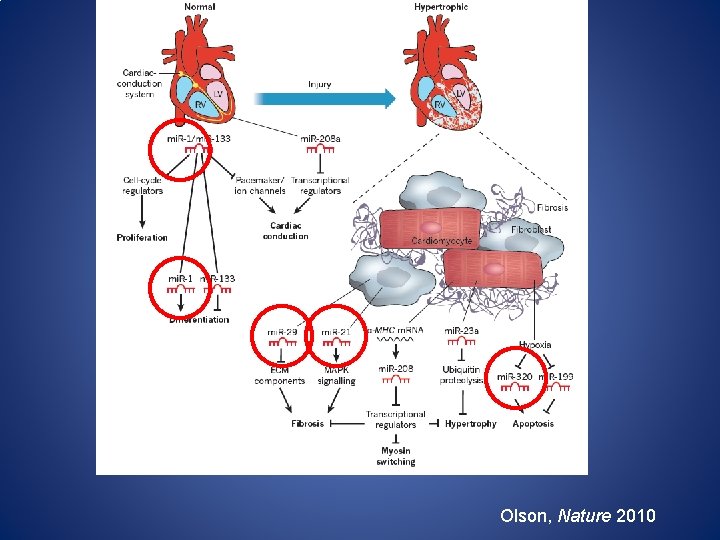
Olson, Nature 2010

Mi. Rhythm Findings • We observed associations between AF and plasma mi. RNAs linked to gene regulatory pathways responsible for cardiac remodeling • Overlap was observed between plasma mi. RNAs associated with AF and those changing after ablation • Studies are needed to explore gene regulatory pathways implicated in susceptibility to AF and to examine the role of mi. RNAs as circulating biomarkers of diagnostic or prognostic importance in AF Mc. Manus et al. submitted Circulation. 2014

Future Directions • Exploring functional significance of mi. RNA dysregulation in animal models of AF • Complete echocardiographic phenotyping of LA structure in FHS and look at genomic and transcriptomic profiles of LA-EF, LAVI • Leverage AF Registry and Biobank

A special thank you to the 650+ UMMS -Nada Esa, MD AF patients who have entrusted -Raghava Velagaleti, MD BU/FHS -John Keaney MD us and participated in -Vasan Ramachandran MDtheir care to -Robert Goldberg Ph. D -Emelia Benjamin MD, Sc. Mthe Umass-Victor Ambros, Ph. D AF Registry, AF -Jane Freedman, MD -Jared Magnani, MD, MPHBiobank, and In. Rhythm! -Kahraman Tanriverdi, Ph. D -Shuxia Fan -Susan Cheng, MD MS -Honghuan Lin, MD MGH -Patrick Ellinor MD, Ph. D -Steven Lubitz, MD -Rosalind Lee, BS -Jeanine Ward, MD Ph. D -Iryna Nieto, MD -Divakar Mandapati, MD -Stanley Tam, MD MBA -Okike N. Okike, MD -Timothy Fitzgibbons, MD -Donna Suter, RN -Amir Shaikh, MD -Menhel Kinno, MD -EP Colleagues

Thank you for your attention!
- Slides: 36