Roland Knispel Business Analyst Biologics and Plexus Suite

Roland Knispel Business Analyst, Biologics and Plexus Suite WHEN IS A LARGE BIOMOLECULE A SMALL MOLECULE? COMPOUND DATABASES WITH A T w i s t

COMPANY FINGERPRINT CHEMAXON provides chemical software development platforms and solutions for the biotechnology and pharmaceutical industries, which are successfully used in publishing, flavors and fragrances research, petroleum and fine chemicals development areas as well. Our innovation targets are to become leaders in web-based data management solutions, to bridge chemistry and biology together, and to offer out-of-the-box solutions for cloud.

Company fingerprint HQ: Budapest, Hungary 18+ 3 130+ 100% 50+ 800+ 200 k+ years of experience offices employees private company implementation partners clients academic users

REAL WORLD SCENARIOS
Customer A A happy and satisfied user of Chem. Axon‘s Compound Registration said „I want to report a performance issue: a single submission takes > 12 min to register“ That‘s unusual, could you send us the structure? Here it is:


Customer A A happy and satisfied user of Chem. Axon‘s Compound Registration I want to report a performance issue: a single submission takes > 12 min to register That‘s unusual, could you send us the structure? Here it is: We accelerated our tautomer check for proteins, protein registration now works in a few seconds. But really try to find a better alternative!
Customer B Another happy and satisfied user of Chem. Axon‘s Compound Registration I want to report a performance issue: registration and search times on our system take minutes. That‘s unusual, could we investigate your DB?

Customer B

Customer B Another happy and satisfied user of Chem. Axon‘s Compound Registration I want to report a performance issue: registration and search times on our system take minutes. That‘s unusual, could we investigate your DB? You have > 50 k proteins in it, stored as a single star atom (*) with the sequence as atom attribute. Our JChem technology is not really optimized for that. We setup a custom pre-filter for you, but it would be better to find an alternative!

Customer C Several of our peptide chemists are drawing their structures in our ELN using Chem. Draw, from where it gets automatically submitted to our Chemical Registration. They receive a registration notification but sometimes cannot locate it in the registry DB afterwards. Turns out a registrar has re-directed some of them to another bespoke registration system. Could you help us resolve this issue? Lost your peptide?

Lost your peptide? Looks like, we‘ve got just the tools you need! We‘ll integrate them into your environment to make the life of your registrars and scientists easier.
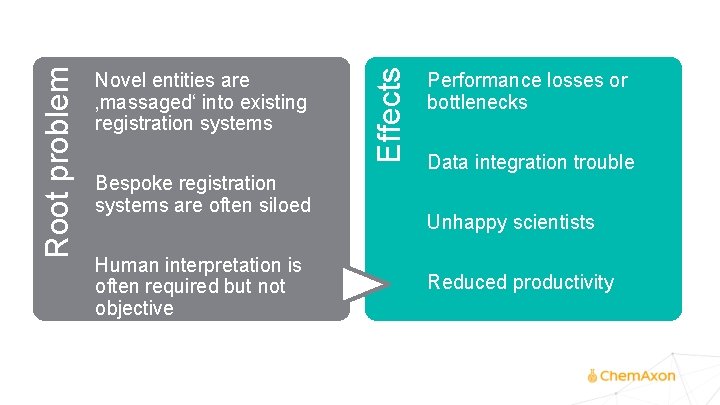
Bespoke registration systems are often siloed Human interpretation is often required but not objective Effects Root problem Novel entities are ‚massaged‘ into existing registration systems Performance losses or bottlenecks Data integration trouble Unhappy scientists Reduced productivity

PLEASE LAUNCH: POLL QUESTION 1

Customer C Several of our peptide chemists are drawing their structures in our ELN using Chem. Draw, from where it gets automatically submitted to our Chemical Registration. They get a registration notification but sometimes cannot always locate it in the registry DB afterwards. Turns out a registrar has re-directed some of them to another bespoke registration system. Could you help us resolve this? Looks like, we have got just the tools you need! We‘ll integrate them into your environment to make the life of registrars and scientists easier.

WHAT‘S IN A STRUCTURE
![Canonical SMILES Cyclosporin A CC[C@@H]1 NC(=O)[C@H](O)[C@ H](C)CC=CC)N(C)C(=O)[C@H](C(C)C)N(C )C(=O)[C@H](CC(C)C)N(C)C(=O)[C@H](C C(C)C)N(C)C(=O)[C@@H](C)NC(=O)[C@H] IUPAC name (chemical) (C)NC(=O)[C@H](CC(C)C)N(C)C(=O)[C@ (3 Canonical SMILES Cyclosporin A CC[C@@H]1 NC(=O)[C@H](O)[C@ H](C)CC=CC)N(C)C(=O)[C@H](C(C)C)N(C )C(=O)[C@H](CC(C)C)N(C)C(=O)[C@H](C C(C)C)N(C)C(=O)[C@@H](C)NC(=O)[C@H] IUPAC name (chemical) (C)NC(=O)[C@H](CC(C)C)N(C)C(=O)[C@ (3](http://slidetodoc.com/presentation_image_h/085b9206db6a6802cc50c96f096817af/image-17.jpg)
Canonical SMILES Cyclosporin A CC[C@@H]1 NC(=O)[C@H](O)[C@ H](C)CC=CC)N(C)C(=O)[C@H](C(C)C)N(C )C(=O)[C@H](CC(C)C)N(C)C(=O)[C@H](C C(C)C)N(C)C(=O)[C@@H](C)NC(=O)[C@H] IUPAC name (chemical) (C)NC(=O)[C@H](CC(C)C)N(C)C(=O)[C@ (3 S, 6 S, 9 S, 12 R, 15 S, 18 S, 21 S, 24 S, 30 S, 33 S)-30 -ethyl-33 -[(E, 1 R, 2 R)-1 -hydroxy- @H](NC(=O)[C@H](CC(C)C)N(C)C(=O)CN( 2 -methylhex-4 -enyl]-1, 4, 7, 10, 12, 15, 19, 25, 28 -nonamethyl-6, 9, 18, 24 -tetrakis(2 - C)C 1=O)C(C)C methylpropyl)-3, 21 -di(propan-2 -yl)-1, 4, 7, 10, 13, 16, 19, 22, 25, 28, 31 Closest natural undecazacyclotritriacontane-2, 5, 8, 11, 14, 17, 20, 23, 26, 29, 32 -undecone AALLVTAGLVL Common name In. Ch. I Cyclosporin A CAS number 59865 -13 -3 Sequence-based depiction IUPAC name (biological) cyclo[((2 S)-2 -aminobutyryl)-sarcosyl-Nmethyl-L-leucyl-L-valyl-N-methyl-L-leucyl. L-alanyl-D-alanyl-N-methyl-L-leucyl-N-methyl-L-valyl-Nmethyl-(4 R)-4 -[(E)-but-2 -enyl]-4 -methyl-L Canonical HELM* PEPTIDE 1{A. [d. A]. [me. L]. [me. V]. [B -threonyl] MT]. [Abu]. [Sar]. [me. L]. V. [me. L]}$PEPTIDE 1, P EPTIDE 1, 11: R 2 -1: R 1$$$ Ref: Pub. Chem CID 5284373 *Chem. Axon generated sequence* In. Ch. I=1 S/C 62 H 111 N 11 O 12/c 1 -25 -27 -2840(15)52(75)51 -56(79)65 -43(262)58(81)67(18)33 -48(74)68(19)44(2934(3)4)55(78)6649(38(11)12)61(84)69(20)45(3035(5)6)54(77)63 -41(16)53(76)6442(17)57(80)70(21)46(3136(7)8)59(82)71(22)47(3237(9)10)60(83)72(23)50(39(13)14)62(85)73 (51)24/h 25, 27, 34 -47, 49 -52, 75 H, 26, 2833 H 2, 124 H 3, (H, 63, 77)(H, 64, 76)(H, 65, 79)(H, 66, 78) /b 27 -25+/t 40 -, 41+, 42, 43+, 44+, 45+, 46+, 47+, 49+, 50+, 51+, 52/m 1/s 1 In. Ch. I key PMATZTZNYRCHOR-CGLBZJNRSA-N
![Cyclosporin A Canonical SMILES CC[C@@H]1 NC(=O)[C@H](O)[C@ H](C)CC=CC)N(C)C(=O)[C@H](C(C)C)N(C )C(=O)[C@H](CC(C)C)N(C)C(=O)[C@H](C C(C)C)N(C)C(=O)[C@@H](C)NC(=O)[C@H] IUPAC name (chemical) (C)NC(=O)[C@H](CC(C)C)N(C)C(=O)[C@ (3 Cyclosporin A Canonical SMILES CC[C@@H]1 NC(=O)[C@H](O)[C@ H](C)CC=CC)N(C)C(=O)[C@H](C(C)C)N(C )C(=O)[C@H](CC(C)C)N(C)C(=O)[C@H](C C(C)C)N(C)C(=O)[C@@H](C)NC(=O)[C@H] IUPAC name (chemical) (C)NC(=O)[C@H](CC(C)C)N(C)C(=O)[C@ (3](http://slidetodoc.com/presentation_image_h/085b9206db6a6802cc50c96f096817af/image-18.jpg)
Cyclosporin A Canonical SMILES CC[C@@H]1 NC(=O)[C@H](O)[C@ H](C)CC=CC)N(C)C(=O)[C@H](C(C)C)N(C )C(=O)[C@H](CC(C)C)N(C)C(=O)[C@H](C C(C)C)N(C)C(=O)[C@@H](C)NC(=O)[C@H] IUPAC name (chemical) (C)NC(=O)[C@H](CC(C)C)N(C)C(=O)[C@ (3 S, 6 S, 9 S, 12 R, 15 S, 18 S, 21 S, 24 S, 30 S, 33 S)-30 -ethyl-33 -[(E, 1 R, 2 R)-1 -hydroxy- @H](NC(=O)[C@H](CC(C)C)N(C)C(=O)CN( 2 -methylhex-4 -enyl]-1, 4, 7, 10, 12, 15, 19, 25, 28 -nonamethyl-6, 9, 18, 24 -tetrakis(2 - C)C 1=O)C(C)C methylpropyl)-3, 21 -di(propan-2 -yl)-1, 4, 7, 10, 13, 16, 19, 22, 25, 28, 31 Closest natural undecazacyclotritriacontane-2, 5, 8, 11, 14, 17, 20, 23, 26, 29, 32 -undecone AALLVTAGLVL Common name In. Ch. I Cyclosporin A CAS number 59865 -13 -3 IUPAC name (biological) cyclo[((2 S)-2 -aminobutyryl)-sarcosyl-Nmethyl-L-leucyl-L-valyl-N-methyl-L-leucyl. L-alanyl-D-alanyl-N-methyl-L-leucyl-N-methyl-L-valyl-Nmethyl-(4 R)-4 -[(E)-but-2 -enyl]-4 -methyl-L Canonical HELM* PEPTIDE 1{A. [d. A]. [me. L]. [me. V]. [B -threonyl] MT]. [Abu]. [Sar]. [me. L]. V. [me. L]}$PEPTIDE 1, P EPTIDE 1, 11: R 2 -1: R 1$$$ Ref: Pub. Chem CID 5284373 *Chem. Axon generated sequence* 2 D structure (Marvin. JS) In. Ch. I=1 S/C 62 H 111 N 11 O 12/c 1 -25 -27 -2840(15)52(75)51 -56(79)65 -43(262)58(81)67(18)33 -48(74)68(19)44(2934(3)4)55(78)6649(38(11)12)61(84)69(20)45(3035(5)6)54(77)63 -41(16)53(76)6442(17)57(80)70(21)46(3136(7)8)59(82)71(22)47(3237(9)10)60(83)72(23)50(39(13)14)62(85)73 (51)24/h 25, 27, 34 -47, 49 -52, 75 H, 26, 2833 H 2, 124 H 3, (H, 63, 77)(H, 64, 76)(H, 65, 79)(H, 66, 78) /b 27 -25+/t 40 -, 41+, 42, 43+, 44+, 45+, 46+, 47+, 49+, 50+, 51+, 52/m 1/s 1 In. Ch. I key PMATZTZNYRCHOR-CGLBZJNRSA-N
![Cyclosporin A Canonical SMILES CC[C@@H]1 NC(=O)[C@H](O)[C@ H](C)CC=CC)N(C)C(=O)[C@H](C(C)C)N(C )C(=O)[C@H](CC(C)C)N(C)C(=O)[C@H](C C(C)C)N(C)C(=O)[C@@H](C)NC(=O)[C@H] IUPAC name (chemical) (C)NC(=O)[C@H](CC(C)C)N(C)C(=O)[C@ (3 Cyclosporin A Canonical SMILES CC[C@@H]1 NC(=O)[C@H](O)[C@ H](C)CC=CC)N(C)C(=O)[C@H](C(C)C)N(C )C(=O)[C@H](CC(C)C)N(C)C(=O)[C@H](C C(C)C)N(C)C(=O)[C@@H](C)NC(=O)[C@H] IUPAC name (chemical) (C)NC(=O)[C@H](CC(C)C)N(C)C(=O)[C@ (3](http://slidetodoc.com/presentation_image_h/085b9206db6a6802cc50c96f096817af/image-19.jpg)
Cyclosporin A Canonical SMILES CC[C@@H]1 NC(=O)[C@H](O)[C@ H](C)CC=CC)N(C)C(=O)[C@H](C(C)C)N(C )C(=O)[C@H](CC(C)C)N(C)C(=O)[C@H](C C(C)C)N(C)C(=O)[C@@H](C)NC(=O)[C@H] IUPAC name (chemical) (C)NC(=O)[C@H](CC(C)C)N(C)C(=O)[C@ (3 S, 6 S, 9 S, 12 R, 15 S, 18 S, 21 S, 24 S, 30 S, 33 S)-30 -ethyl-33 -[(E, 1 R, 2 R)-1 -hydroxy- @H](NC(=O)[C@H](CC(C)C)N(C)C(=O)CN( 2 -methylhex-4 -enyl]-1, 4, 7, 10, 12, 15, 19, 25, 28 -nonamethyl-6, 9, 18, 24 -tetrakis(2 - C)C 1=O)C(C)C methylpropyl)-3, 21 -di(propan-2 -yl)-1, 4, 7, 10, 13, 16, 19, 22, 25, 28, 31 Closest natural undecazacyclotritriacontane-2, 5, 8, 11, 14, 17, 20, 23, 26, 29, 32 -undecone AALLVTAGLVL Common name In. Ch. I Cyclosporin A CAS number 59865 -13 -3 IUPAC name (biological) cyclo[((2 S)-2 -aminobutyryl)-sarcosyl-Nmethyl-L-leucyl-L-valyl-N-methyl-L-leucyl. L-alanyl-D-alanyl-N-methyl-L-leucyl-N-methyl-L-valyl-Nmethyl-(4 R)-4 -[(E)-but-2 -enyl]-4 -methyl-L Canonical HELM* PEPTIDE 1{A. [d. A]. [me. L]. [me. V]. [B -threonyl] MT]. [Abu]. [Sar]. [me. L]. V. [me. L]}$PEPTIDE 1, P EPTIDE 1, 11: R 2 -1: R 1$$$ Ref: Pub. Chem CID 5284373 *Chem. Axon generated sequence* Sequence Bio. Eddie In. Ch. I=1 S/C 62 H 111 N 11 O 12/c 1 -25 -27 -2840(15)52(75)51 -56(79)65 -43(262)58(81)67(18)33 -48(74)68(19)44(2934(3)4)55(78)6649(38(11)12)61(84)69(20)45(3035(5)6)54(77)63 -41(16)53(76)6442(17)57(80)70(21)46(3136(7)8)59(82)71(22)47(3237(9)10)60(83)72(23)50(39(13)14)62(85)73 (51)24/h 25, 27, 34 -47, 49 -52, 75 H, 26, 2833 H 2, 124 H 3, (H, 63, 77)(H, 64, 76)(H, 65, 79)(H, 66, 78) /b 27 -25+/t 40 -, 41+, 42, 43+, 44+, 45+, 46+, 47+, 49+, 50+, 51+, 52/m 1/s 1 In. Ch. I key PMATZTZNYRCHOR-CGLBZJNRSA-N
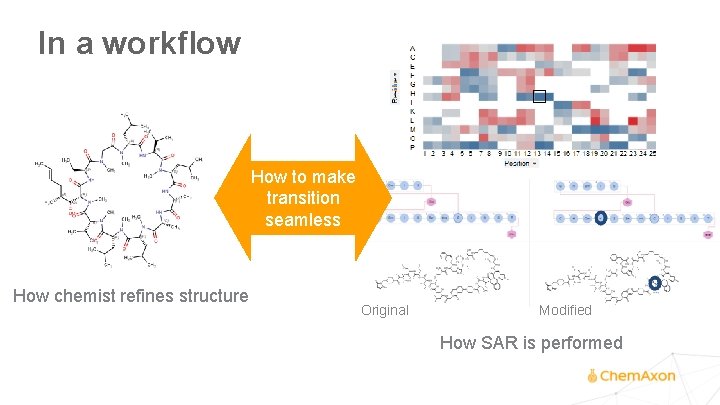
In a workflow How to make transition seamless How chemist refines structure Original Modified How SAR is performed

CASE STUDY: CHEMBL V 21 Data migration, enrichment and curation

CHEMBL v 21 • 19773 peptide biotherapeutics with associated activity data • Molecules served as HELM or chemical structure file • Ch. EMBL interface: searchable by Compound. ID, chemical structure or select metadata
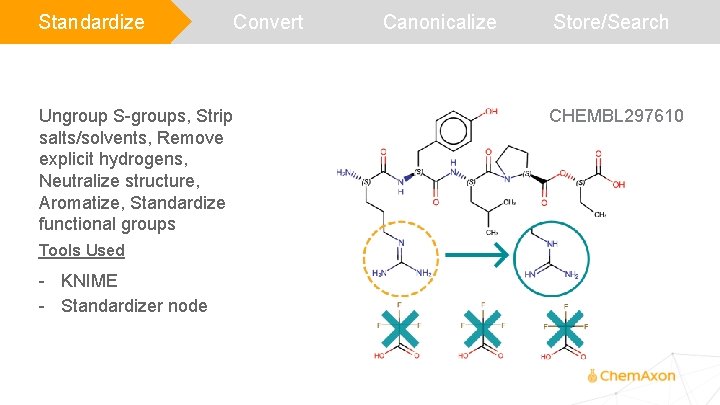
Standardize Ungroup S-groups, Strip salts/solvents, Remove explicit hydrogens, Neutralize structure, Aromatize, Standardize functional groups Tools Used - KNIME - Standardizer node Convert Canonicalize Store/Search CHEMBL 297610
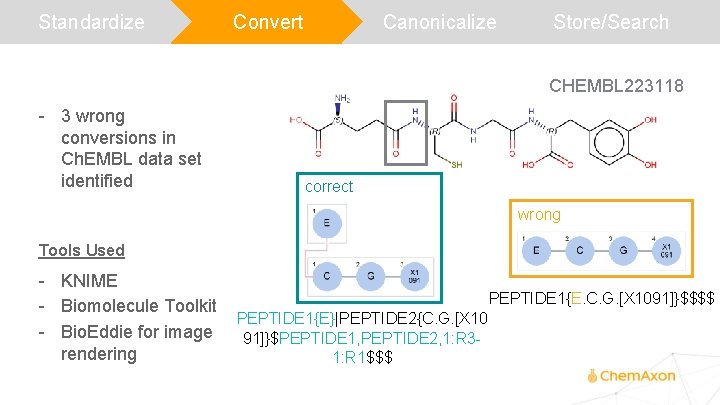
Standardize Convert Canonicalize Store/Search CHEMBL 223118 - 3 wrong conversions in Ch. EMBL data set identified correct wrong Tools Used - KNIME - Biomolecule Toolkit - Bio. Eddie for image rendering PEPTIDE 1{E. C. G. [X 1091]}$$$$ PEPTIDE 1{E}|PEPTIDE 2{C. G. [X 10 91]}$PEPTIDE 1, PEPTIDE 2, 1: R 31: R 1$$$
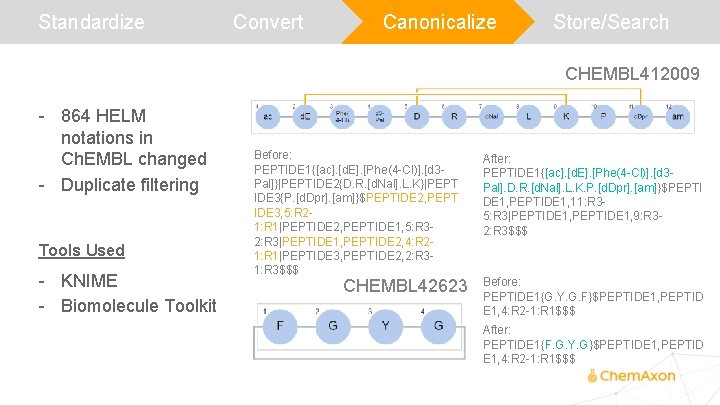
Standardize Convert Canonicalize Store/Search CHEMBL 412009 - 864 HELM notations in Ch. EMBL changed - Duplicate filtering Tools Used - KNIME - Biomolecule Toolkit Before: PEPTIDE 1{[ac]. [d. E]. [Phe(4 -Cl)]. [d 3 Pal]}|PEPTIDE 2{D. R. [d. Nal]. L. K}|PEPT IDE 3{P. [d. Dpr]. [am]}$PEPTIDE 2, PEPT IDE 3, 5: R 21: R 1|PEPTIDE 2, PEPTIDE 1, 5: R 32: R 3|PEPTIDE 1, PEPTIDE 2, 4: R 21: R 1|PEPTIDE 3, PEPTIDE 2, 2: R 31: R 3$$$ CHEMBL 42623 After: PEPTIDE 1{[ac]. [d. E]. [Phe(4 -Cl)]. [d 3 Pal]. D. R. [d. Nal]. L. K. P. [d. Dpr]. [am]}$PEPTI DE 1, PEPTIDE 1, 11: R 35: R 3|PEPTIDE 1, 9: R 32: R 3$$$ Before: PEPTIDE 1{G. Y. G. F}$PEPTIDE 1, PEPTID E 1, 4: R 2 -1: R 1$$$ After: PEPTIDE 1{F. G. Y. G}$PEPTIDE 1, PEPTID E 1, 4: R 2 -1: R 1$$$

Standardize Search by: - Sequence (incl. wildcards) - Chemical structure - Modifications - Metadata Tools Used - Biomolecule Toolkit Convert Canonicalize Store/Search Query # hits Molecules containing „Oxytocin“ in name field 2 Molecules with the natural analogue sequence of Oxytocin 18 Oxytocin-like sequences with non-standard amino acids 17 Oxytocin derivatives containing the chemical structure of penicillamine 3 Oxytocin derivatives containing the Lpenicillamine monomer 2

Bio. Eddie: Oxytocin from Sequence or MOL File

BIOMOLECULE TOOLKIT AND BIOEDDIE JChem and Marvin analogues for large molecule informatics

Biomolecule Toolkit • API (Java and REST-ful) for – – Native HELM support (HELM, HELM 2, x. HELM) Standardization Centralized DB storage Registration of entities and batches with custom business logic – Search by sequence/chemical structure/metadata – Conversion to/from Mol/FASTA/HELM – Property calculations • Integrated in – Instant. JChem, Texelia Bio. Scity, IDBS E-Workbook
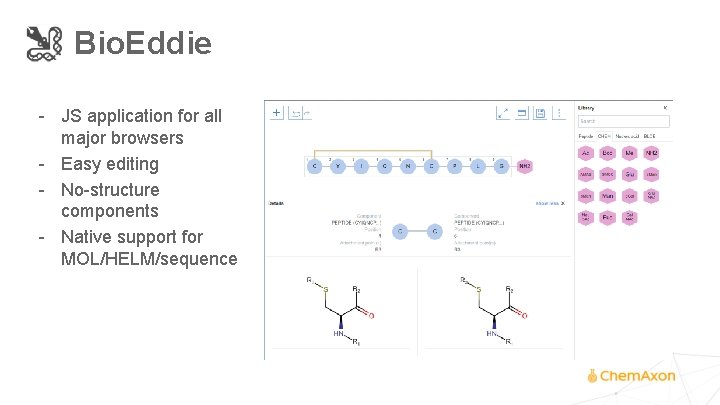
Bio. Eddie - JS application for all major browsers - Easy editing - No-structure components - Native support for MOL/HELM/sequence

Bio. Eddie - JS application for all major browsers - Easy editing - No-structure components - Native support for MOL/HELM/sequence - Customizable views - Multi-level annotations

Bio. Eddie - JS application for all major browsers - Easy editing - No-structure components - Native support for MOL/HELM/sequence - Customizable views - Multi-level annotations - NEW: sequence domain support (Abs)

SUPPORTED ENTITY TYPES
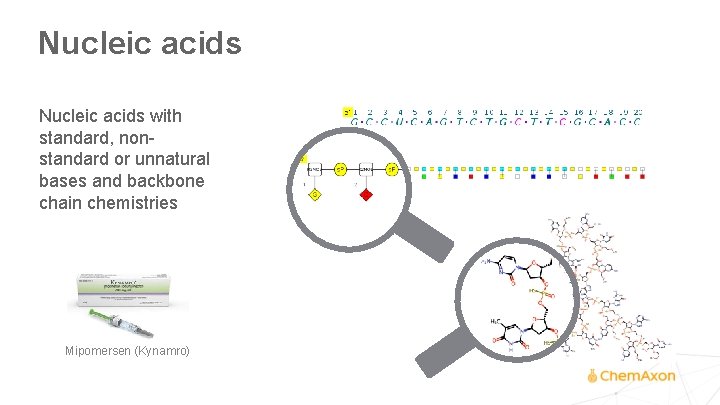
Nucleic acids with standard, nonstandard or unnatural bases and backbone chain chemistries Mipomersen (Kynamro)
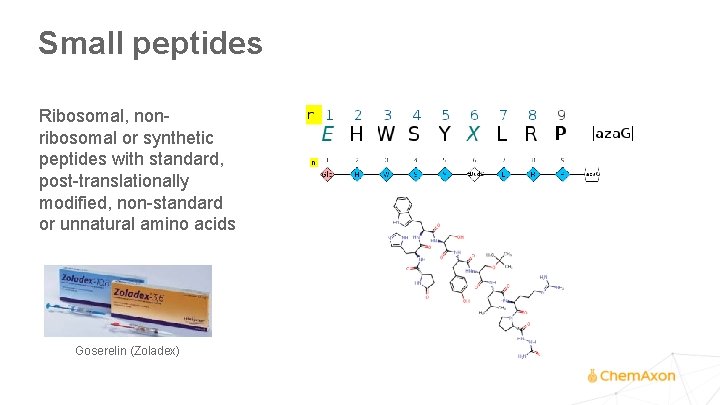
Small peptides Ribosomal, nonribosomal or synthetic peptides with standard, post-translationally modified, non-standard or unnatural amino acids Goserelin (Zoladex)

Large peptides (proteins) Protein sequences including posttranslationally modified residues, intrachain and interchain cross-links Trastuzumab (Herceptin) Monoclonal antibody
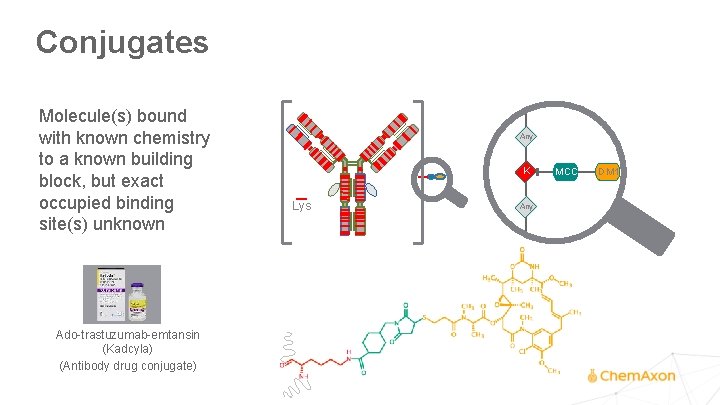
Conjugates Molecule(s) bound with known chemistry to a known building block, but exact occupied binding site(s) unknown Any K Lys Any ADR=3. 5 Ado-trastuzumab-emtansin (Kadcyla) (Antibody drug conjugate) MCC DM 1

PLEASE LAUNCH: POLL QUESTION 2

AGNOSTIC REGISTRATION
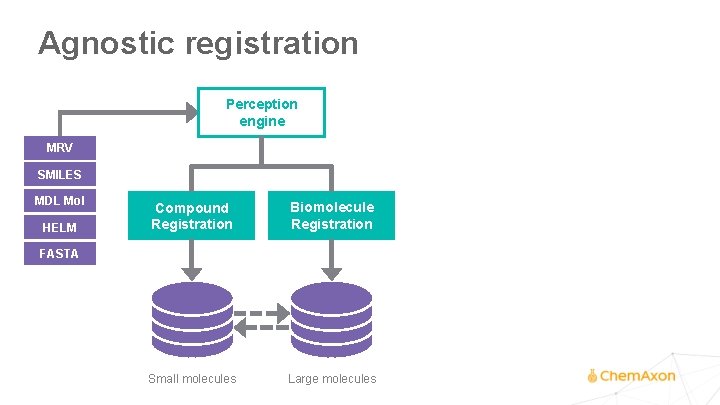
Agnostic registration Perception engine MRV SMILES MDL Mol HELM Compound Registration Biomolecule Registration Small molecules Large molecules FASTA
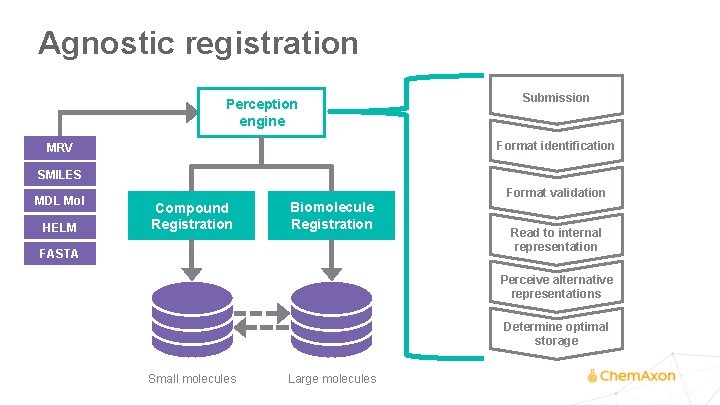
Agnostic registration Perception engine Submission Format identification MRV SMILES MDL Mol HELM Format validation Compound Registration Biomolecule Registration FASTA Read to internal representation Perceive alternative representations Determine optimal storage Small molecules Large molecules
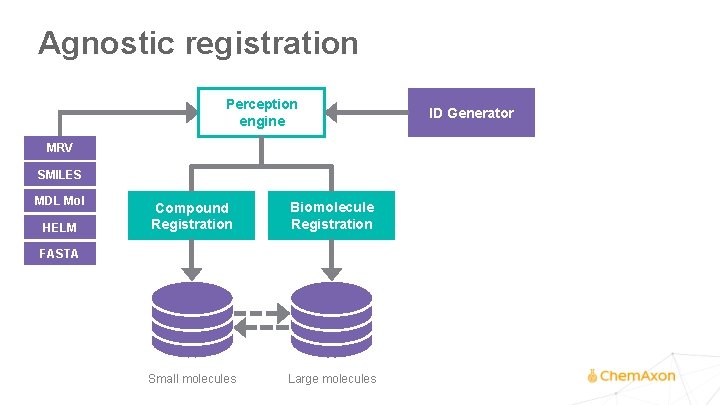
Agnostic registration Perception engine MRV SMILES MDL Mol HELM Compound Registration Biomolecule Registration Small molecules Large molecules FASTA ID Generator
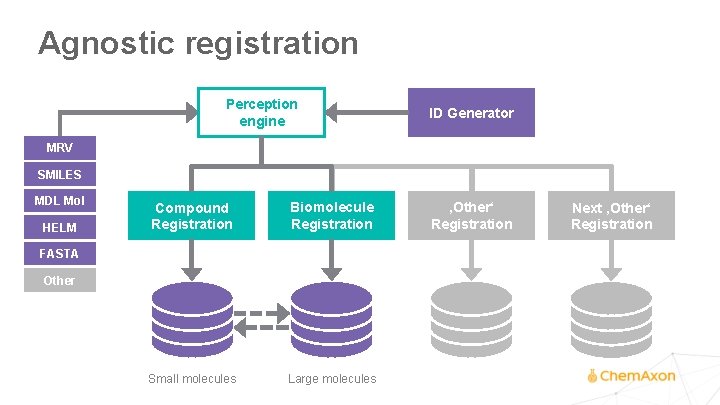
Agnostic registration Perception engine ID Generator MRV SMILES MDL Mol HELM Compound Registration Biomolecule Registration Small molecules Large molecules FASTA Other ‚Other‘ Registration Next ‚Other‘ Registration
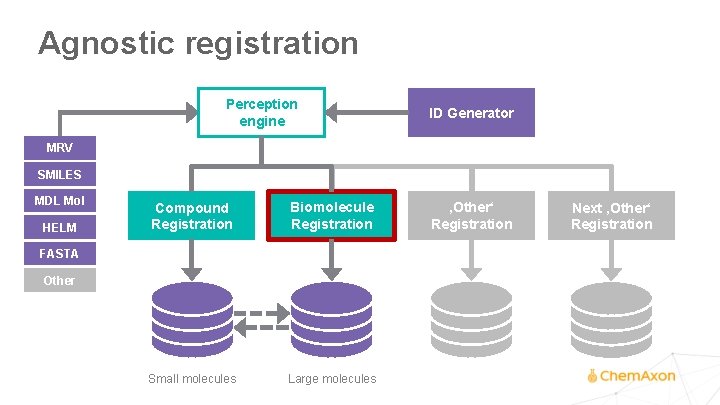
Agnostic registration Perception engine ID Generator MRV SMILES MDL Mol HELM Compound Registration Biomolecule Registration Small molecules Large molecules FASTA Other ‚Other‘ Registration Next ‚Other‘ Registration

SUMMARY
Bespoke registration systems are often siloed Human interpretation is often required but not objective Cure Root problem Novel entities are ‚massaged‘ into existing registration systems Bespoke registration systems ensure performance, accuracy and consistency A perception engine helps to integrate registration and other workflows Chem. Axon offers tools and services to implement

WANT TO KNOW MORE? GET IN TOUCH! Roland Knispel rknispel@chemaxon. com

Q&A Please feel free to ask away!
- Slides: 48