Rock lizards Gene flow and hybridization within species

Rock lizards - Gene flow and hybridization within species of "mixta" clade Sopiko Kurdadze Supervisor: Professor David Tarkhnishvili Master’s Program of Applied Genetics Ilia State University 12/7/2020

1 • Aim of the research 2 • Overview 3 • Materials and Methods 4 • Results and interpretation 5 • Conclusion 12/7/2020

Aim of the research q To check a gene flow between Darevskia mixta and Darevkia derjugini where their areas overlap q To study population structure and genetic diversity of both species by using microsatellite loci 12/7/2020
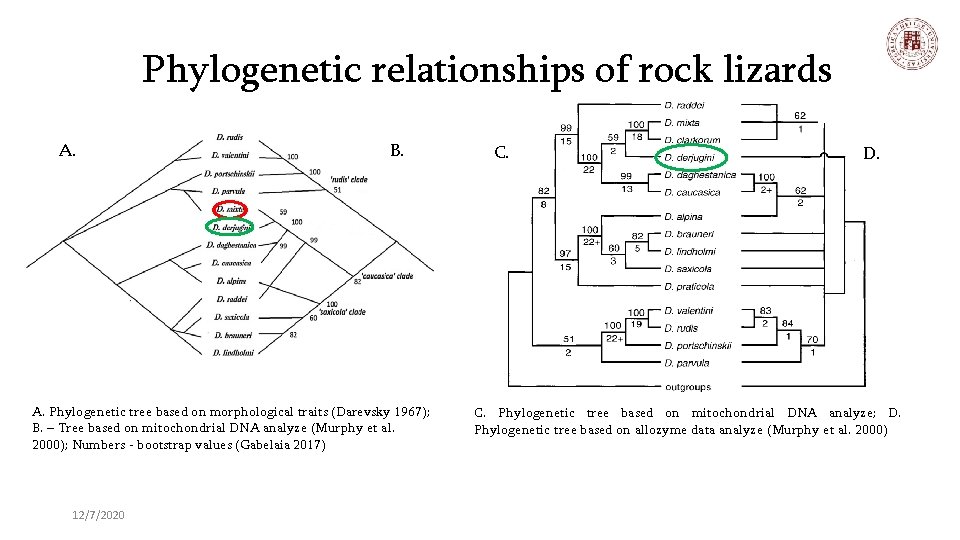
Phylogenetic relationships of rock lizards A. B. A. Phylogenetic tree based on morphological traits (Darevsky 1967); B. – Tree based on mitochondrial DNA analyze (Murphy et al. 2000); Numbers - bootstrap values (Gabelaia 2017) 12/7/2020 C. D. C. Phylogenetic tree based on mitochondrial DNA analyze; D. Phylogenetic tree based on allozyme data analyze (Murphy et al. 2000)

Hypothesis about hybridization between different species v. D. derjugini + D. parvula = D. mixta (Darevky 1967) v. D. mixta + D. derjugini=? ? ? (Murphy et al. 2000) v. D. derjugini and D. mixta had an identical mitochondrial haplogroups Gabelaia (2013) 12/7/2020

Sample collection 12/7/2020
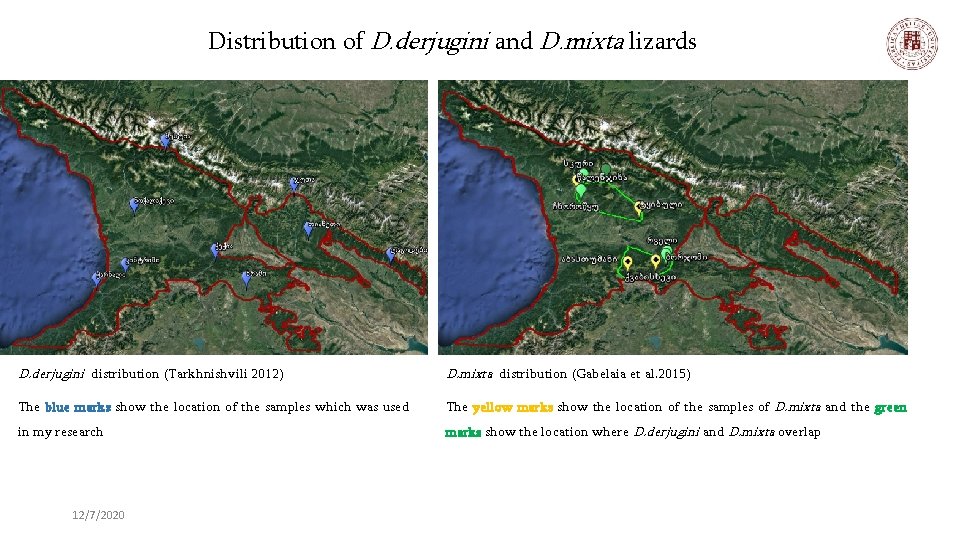
Distribution of D. derjugini and D. mixta lizards D. derjugini distribution (Tarkhnishvili 2012) D. mixta distribution (Gabelaia et al. 2015) The blue marks show the location of the samples which was used The yellow marks show the location of the samples of D. mixta and the green in my research marks show the location where D. derjugini and D. mixta overlap 12/7/2020

Methods q DNA extraction (QIAGEN®, 2007) q PCR (Polymerase Chain Reaction) of nuclear microsatellite loci: Du 161, Du 183, Du 255, Du 231, Du 363, Du 47, Du 215, Du 418, Du 281 and. Du 323 (Korchagin et al. 2003; 2007; Malysheva 2012) q Electrophoresis (ABI 3130 Genetic Analyzer) q Visualization of Alleles Gene. Mapper. TM V 5 (Applied Biosystems) q Statistical methods Arlequin V 3. 5. 2. 2 (Excoffier & Lischer 2010) and Structure V 2. 3. 4 (Pritchard et al. 2000) 12/7/2020
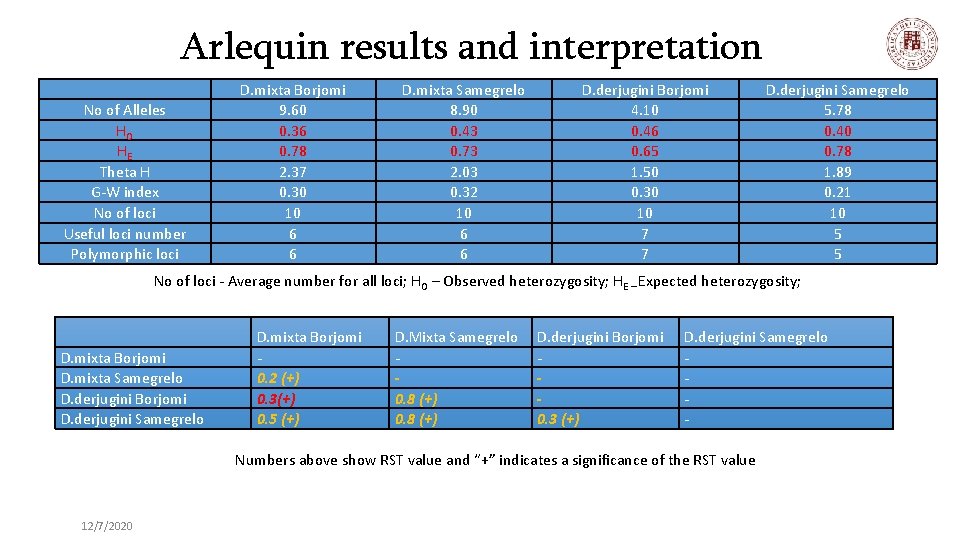
Arlequin results and interpretation No of Alleles HO HE Theta H G-W index No of loci Useful loci number Polymorphic loci D. mixta Borjomi 9. 60 0. 36 0. 78 2. 37 0. 30 10 6 6 D. mixta Samegrelo 8. 90 0. 43 0. 73 2. 03 0. 32 10 6 6 D. derjugini Borjomi 4. 10 0. 46 0. 65 1. 50 0. 30 10 7 7 D. derjugini Samegrelo 5. 78 0. 40 0. 78 1. 89 0. 21 10 5 5 No of loci - Average number for all loci; HO – Observed heterozygosity; HE – Expected heterozygosity; D. mixta Borjomi D. mixta Samegrelo D. derjugini Borjomi D. derjugini Samegrelo D. mixta Borjomi 0. 2 (+) 0. 3(+) 0. 5 (+) D. Mixta Samegrelo 0. 8 (+) D. derjugini Borjomi 0. 3 (+) D. derjugini Samegrelo - Numbers above show RST value and “+” indicates a significance of the RST value 12/7/2020
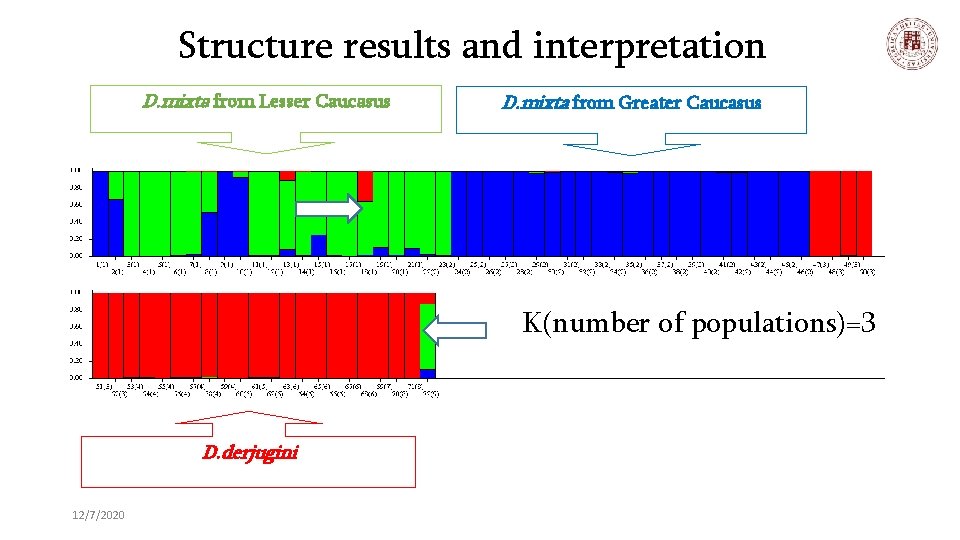
Structure results and interpretation D. mixta from Lesser Caucasus D. mixta from Greater Caucasus K(number of populations)=3 D. derjugini 12/7/2020
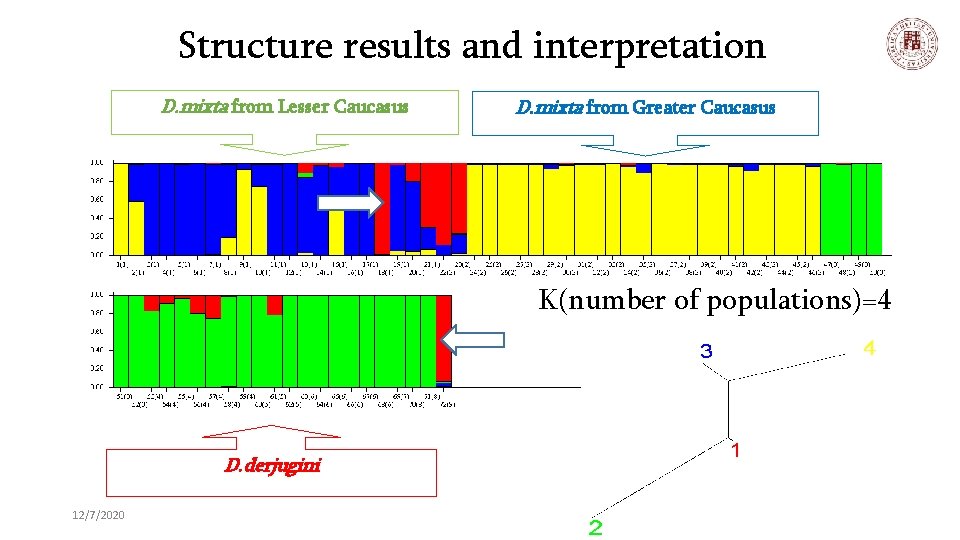
Structure results and interpretation D. mixta from Lesser Caucasus D. mixta from Greater Caucasus K(number of populations)=4 D. derjugini 12/7/2020

Conclusion q There is a gene flow between Lesser Caucasus D. mixta and D. derjugini where they overlap q D. mixta population of Lesser Caucasus is more diverse than the population of Greater Caucasus q Population of D. derjugini more likely tend to Hardy-Weinberg equilibrium than population of D. mixta 12/7/2020

Questions? 12/7/2020

Thank you for your attention! 12/7/2020
- Slides: 14