Robustness in system modeling Rdiger W Brause Traditional

Robustness in system modeling Rüdiger W. Brause

Traditional Modelling, e. g. modelling of biochemical pathways Traditional time dynamic modeling § fuzzy clustering stage § dynamical interaction of the clusters by linear differential equations based on the expression data of selected genes § selection criterion: most simple network But: after long evolutionary development, small genetic mutations will not cause fatal changes any more. Ni. SIS Workshop, Mallorca 2006 R. Brause: Nature-inspired Robustness sheet 2
Example plasmid replication Escherichia coli Col E 1 plasmids = short DNA loops • give resistance against toxics and antibiotics • replicated separately • segregated on cell division High plasmid replication: longer bacteria replication time, smaller fraction of population No plasmid replication: smaller fitness in the long run, not in the short. Modest plasmid replication regulation – how? Ni. SIS Workshop, Mallorca 2006 R. Brause: Nature-inspired Robustness sheet 3
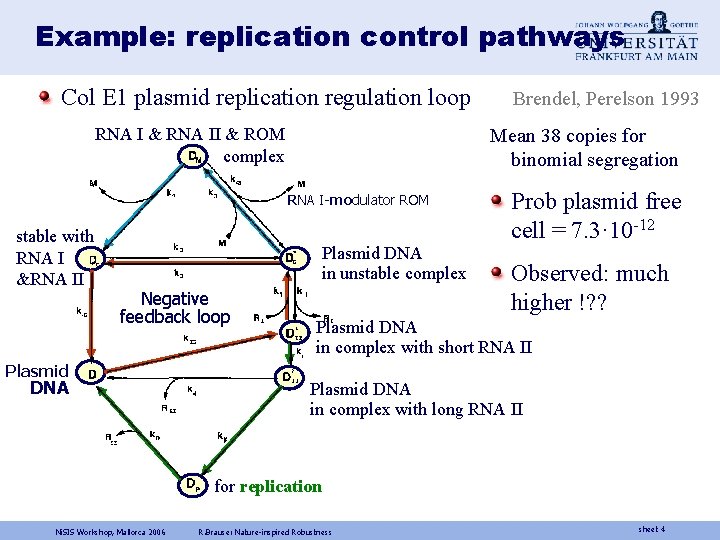
Example: replication control pathways Col E 1 plasmid replication regulation loop RNA I & RNA II & ROM complex Mean 38 copies for binomial segregation RNA I-modulator ROM stable with RNA I &RNA II Plasmid DNA in unstable complex Negative feedback loop Plasmid DNA Brendel, Perelson 1993 Prob plasmid free cell = 7. 3· 10 -12 Observed: much higher !? ? Plasmid DNA in complex with short RNA II Plasmid DNA in complex with long RNA II for replication Ni. SIS Workshop, Mallorca 2006 R. Brause: Nature-inspired Robustness sheet 4
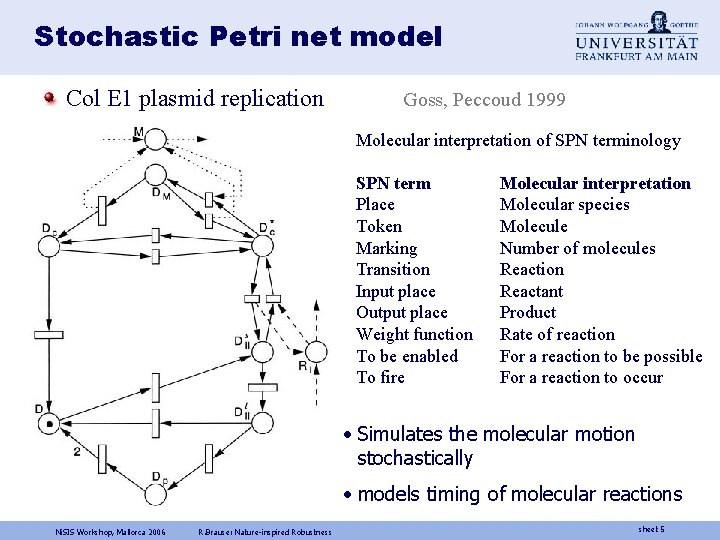
Stochastic Petri net model Col E 1 plasmid replication Goss, Peccoud 1999 Molecular interpretation of SPN terminology SPN term Place Token Marking Transition Input place Output place Weight function To be enabled To fire Molecular interpretation Molecular species Molecule Number of molecules Reaction Reactant Product Rate of reaction For a reaction to be possible For a reaction to occur • Simulates the molecular motion stochastically • models timing of molecular reactions Ni. SIS Workshop, Mallorca 2006 R. Brause: Nature-inspired Robustness sheet 5
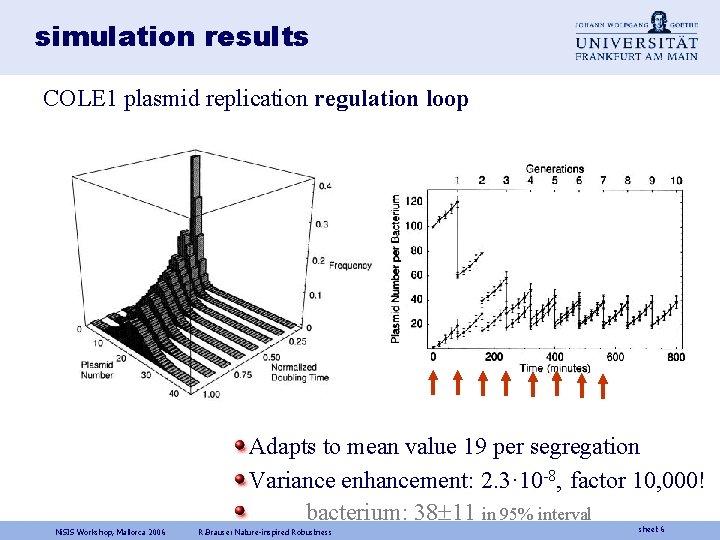
simulation results COLE 1 plasmid replication regulation loop Adapts to mean value 19 per segregation Variance enhancement: 2. 3· 10 -8, factor 10, 000! bacterium: 38 11 in 95% interval Ni. SIS Workshop, Mallorca 2006 R. Brause: Nature-inspired Robustness sheet 6

Simulation results No ROM protein double plasmids/bacterium Kinetic parameters adapted for same plasmid mean like wild type bigger variance of mutant 2 -6 fold plasmid loss ! No segregation variance assumed variance is due to timing Ni. SIS Workshop, Mallorca 2006 R. Brause: Nature-inspired Robustness sheet 7
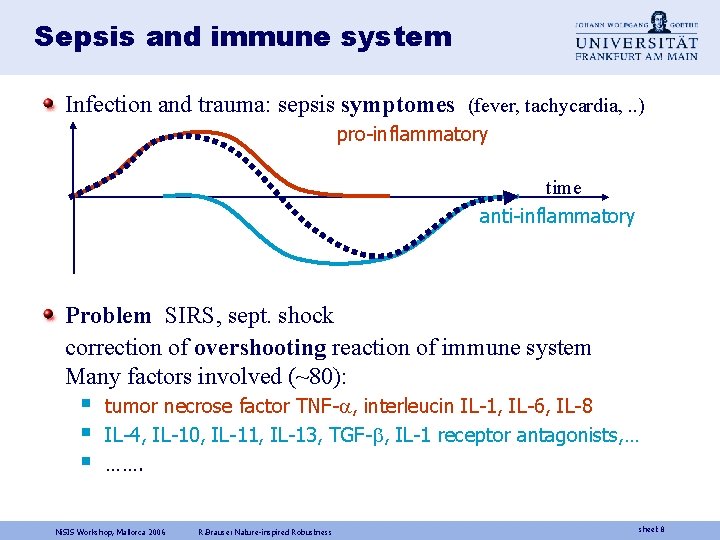
Sepsis and immune system Infection and trauma: sepsis symptomes (fever, tachycardia, . . ) pro-inflammatory time anti-inflammatory Problem SIRS, sept. shock correction of overshooting reaction of immune system Many factors involved (~80): § § § tumor necrose factor TNF- , interleucin IL-1, IL-6, IL-8 IL-4, IL-10, IL-11, IL-13, TGF- , IL-1 receptor antagonists, … ……. Ni. SIS Workshop, Mallorca 2006 R. Brause: Nature-inspired Robustness sheet 8
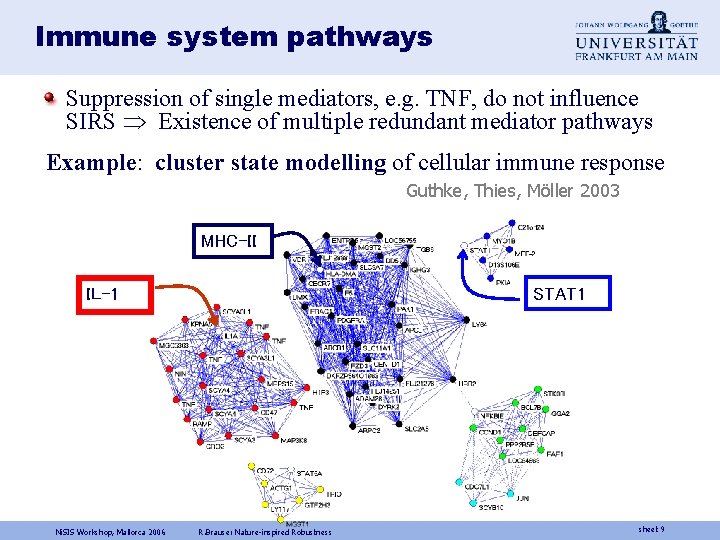
Immune system pathways Suppression of single mediators, e. g. TNF, do not influence SIRS Existence of multiple redundant mediator pathways Example: cluster state modelling of cellular immune response Guthke, Thies, Möller 2003 MHC-II STAT 1 IL-1 Ni. SIS Workshop, Mallorca 2006 R. Brause: Nature-inspired Robustness sheet 9

Differential equation modeling n clusters, n time series xj(t) of representative genes Setup n differential equations by Select variable xj which determines a set of measurements Select next variable xi which is only determined by xj By induction, continue the modeling by with u(t>0) = 1, u(t<0) = 0 and xi(0) = 0. Coupling coefficients wi, j are determined approximately Time delay r is included. Ni. SIS Workshop, Mallorca 2006 R. Brause: Nature-inspired Robustness sheet 10

Differential equation modeling Critics: The number of clusters, i. e. the number of variables is chosen according to some criteria which do not take possible interactions and redundancy into account. The dependency evaluation of the xi is taken by measuring correlations Also, the values of the coupling coefficients are due to some side conditions which do not include possible deviations of the computed parameter values. The criteria for modelling performance is the mean squared error, but probability distributions instead of the MSE of the results may better represent the resulting networks and give better probability estimations for redundant branches. Ni. SIS Workshop, Mallorca 2006 R. Brause: Nature-inspired Robustness sheet 11

Proposition 1: black box modeling Model causalities H(X) = entropy (average information) of variable X I(X; Y) = mutual information between X and Y if I(X; Y) = H(Y) and H(Y) < H(X) then „Y depends on X“ Black box modeling Dyi (t) = fi(x 1(t-1), x 2(t-1), . . . , xn(t-1) ) e. g. neural network approximation Select only those xi with I(Y; X 1, X 2, . . , Xn) > 90% (ranked list) Ni. SIS Workshop, Mallorca 2006 R. Brause: Nature-inspired Robustness sheet 12

Simple sepsis model Preferred modeling strategy: make it simple! make clusters choose clusters representatives model state dynamics between representatives as simple as possible Example protein interaction map of pheromone cell response B Steffe, Petti, Aach, D‘haeseleer, Church 2002 start signal protein target 70 proteins, 354 pathways Ni. SIS Workshop, Mallorca 2006 score > median R. Brause: Nature-inspired Robustness top 15 graph, node size =S path scores sheet 13

Proposition 2: robustness constraints Select robust pathway modeling predict new signalling pathways compared to literature identify previously unknown members of documented pathways identify relevant groups of interacting proteins Robustness Criteria ü fault tolerance: random faults should not propagate and impede essential system functions ü inherent stability: no system deviation by noise or random input, even by internal component change Ni. SIS Workshop, Mallorca 2006 R. Brause: Nature-inspired Robustness sheet 14
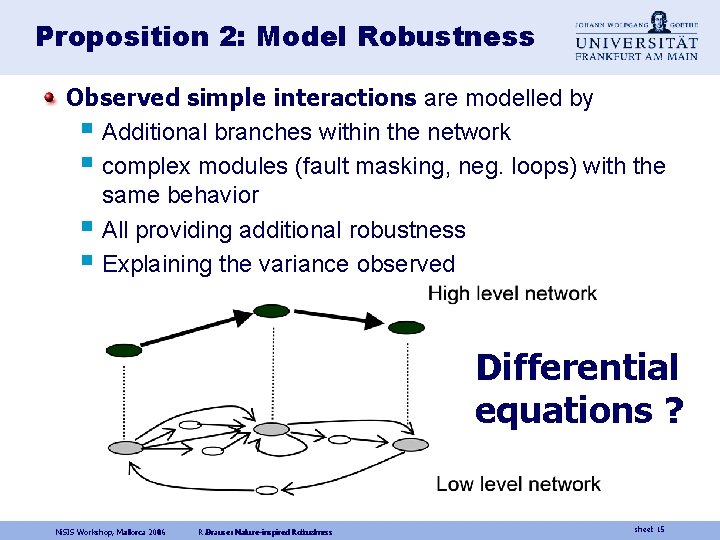
Proposition 2: Model Robustness Observed simple interactions are modelled by § Additional branches within the network § complex modules (fault masking, neg. loops) with the same behavior § All providing additional robustness § Explaining the variance observed Differential equations ? Ni. SIS Workshop, Mallorca 2006 R. Brause: Nature-inspired Robustness sheet 15
- Slides: 15