Robustness Analysis and Tuning of Synthetic Gene Networks

Robustness Analysis and Tuning of Synthetic Gene Networks Grégory Batt 1‡ Boyan Yordanov 1 Calin Belta 1 Ron Weiss 2 1 Centers for Information and Systems Engineering and for Bio. Dynamics Boston University 2 (‡ now at Departments of Molecular Biology and of Electrical Engineering Princeton University Towards Systems Biology 2007 )

Synthetic biology v Synthetic biology: application of engineering approaches to produce novel artificial devices using biological building blocks

Synthetic biology v Synthetic biology: application of engineering approaches to produce novel artificial devices using biological building blocks banana-smelling bacteria
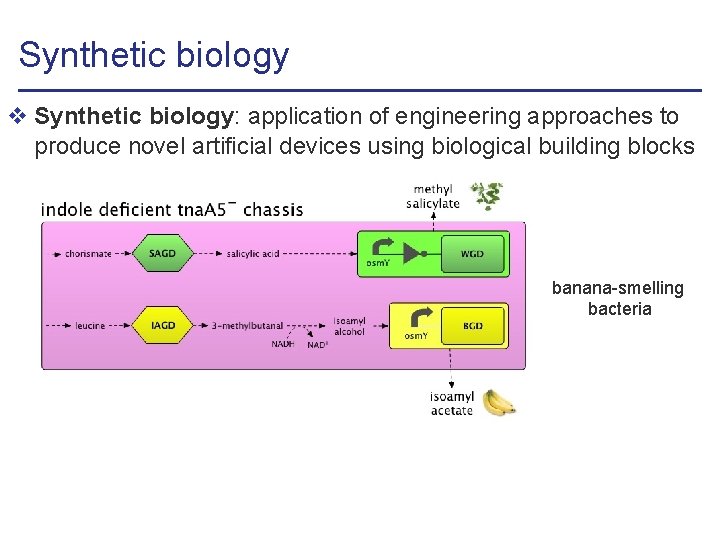
Synthetic biology v Synthetic biology: application of engineering approaches to produce novel artificial devices using biological building blocks banana-smelling bacteria
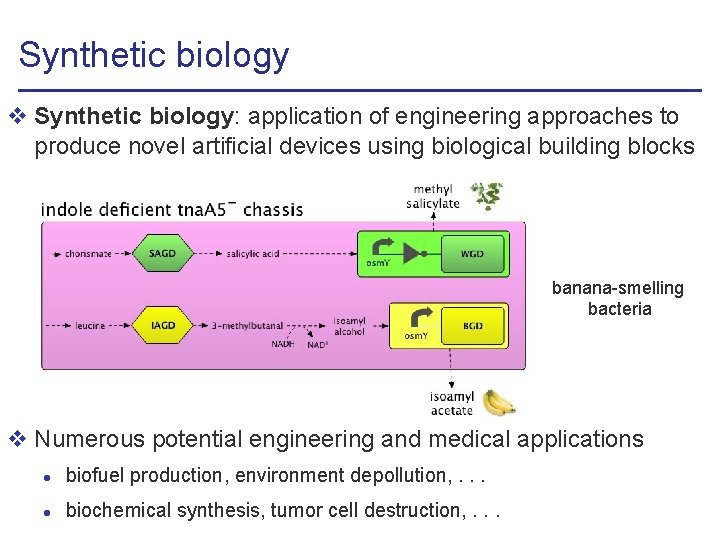
Synthetic biology v Synthetic biology: application of engineering approaches to produce novel artificial devices using biological building blocks banana-smelling bacteria v Numerous potential engineering and medical applications l biofuel production, environment depollution, . . . l biochemical synthesis, tumor cell destruction, . . .
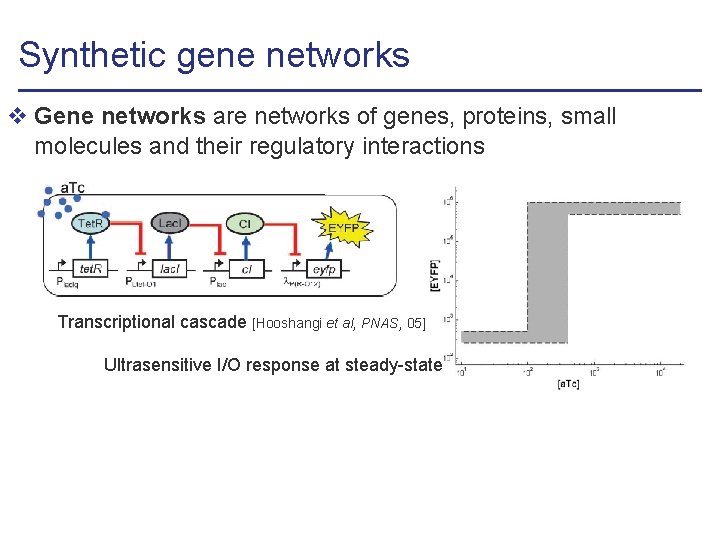
Synthetic gene networks v Gene networks are networks of genes, proteins, small molecules and their regulatory interactions Transcriptional cascade [Hooshangi et al, PNAS, 05] Ultrasensitive I/O response at steady-state
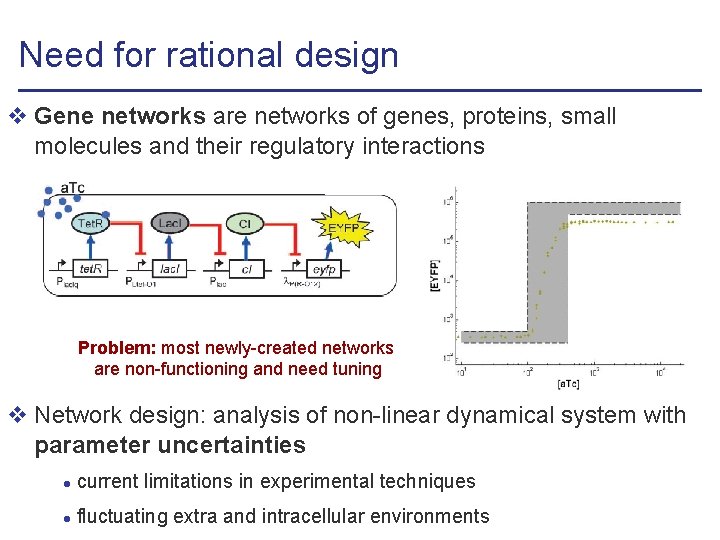
Need for rational design v Gene networks are networks of genes, proteins, small molecules and their regulatory interactions Problem: most newly-created networks are non-functioning and need tuning v Network design: analysis of non-linear dynamical system with parameter uncertainties l current limitations in experimental techniques l fluctuating extra and intracellular environments

Robustness analysis and tuning v Two problems of interest: l robustness analysis: check whether dynamical properties are satisfied for all parameters in a set l tuning: find parameter sets such that dynamical properties are satisfied for all parameters in the sets v Approach: l unknown parameters, initial conditions and inputs given by intervals l piecewise-multiaffine differential equations models of gene networks l dynamical properties specified in temporal logic (LTL) l adapt techniques from hybrid systems theory and model checking

Hybrid systems approach v Analysis of dynamical systems l Traditional view: fixed initial condition and fixed parameter l More interesting: set of initial conditions and set of parameters P 1 p 1 X 0 x 0 p 2 P 2

Hybrid systems approach v Analysis of dynamical systems l Traditional view: fixed initial condition and fixed parameter l More interesting: set of initial conditions and set of parameters P 1 p 1 X 0 x 0 p 2 P 2 v How to reason with infinite number of parameters and initial conditions ?
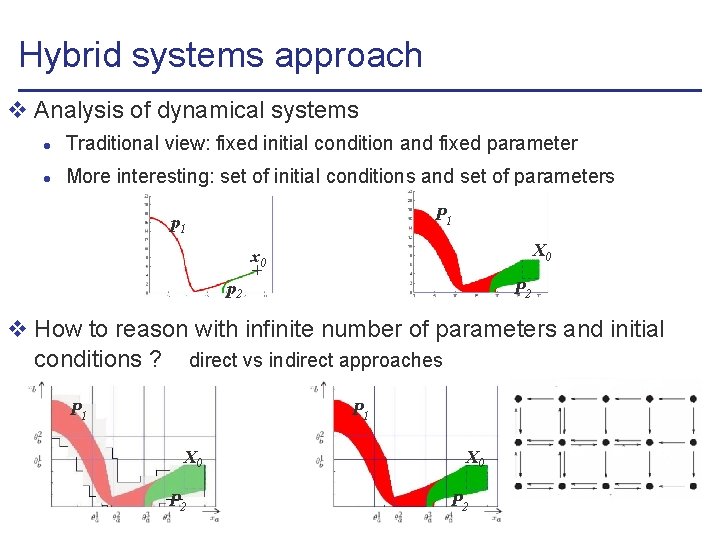
Hybrid systems approach v Analysis of dynamical systems l Traditional view: fixed initial condition and fixed parameter l More interesting: set of initial conditions and set of parameters P 1 p 1 X 0 x 0 P 2 p 2 v How to reason with infinite number of parameters and initial conditions ? direct vs indirect approaches P 1 X 0 P 2
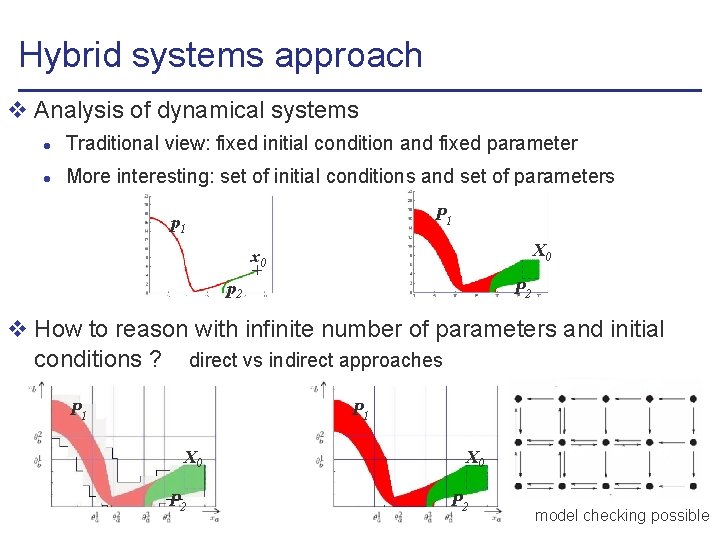
Hybrid systems approach v Analysis of dynamical systems l Traditional view: fixed initial condition and fixed parameter l More interesting: set of initial conditions and set of parameters P 1 p 1 X 0 x 0 P 2 p 2 v How to reason with infinite number of parameters and initial conditions ? direct vs indirect approaches P 1 X 0 P 2 model checking possible

Overview I. Introduction II. Problem definition III. Robust design of gene networks IV. Application: tuning a synthetic transcriptional cascade V. Discussion and conclusions

Overview I. Introduction II. Problem definition III. Robust design of gene networks IV. Application: tuning a synthetic transcriptional cascade V. Discussion and conclusions
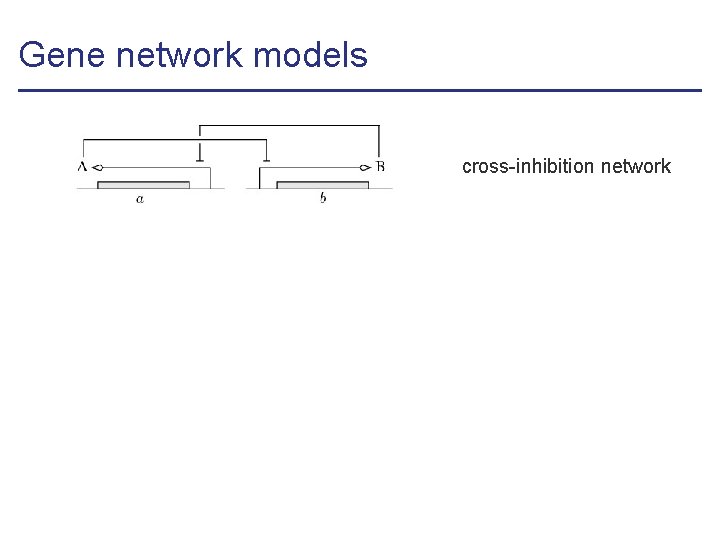
Gene network models cross-inhibition network
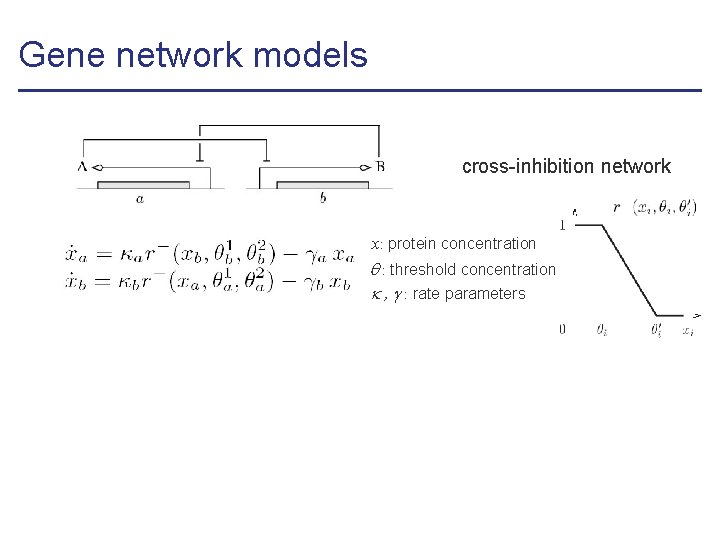
Gene network models cross-inhibition network x: protein concentration : threshold concentration , : rate parameters
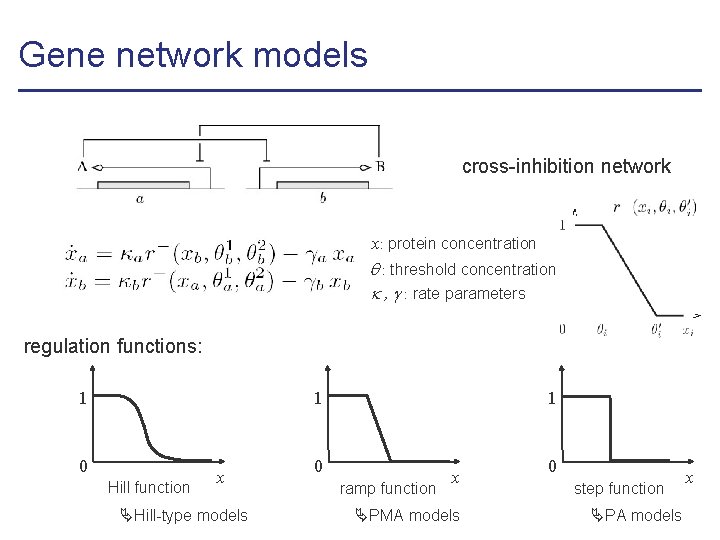
Gene network models cross-inhibition network x: protein concentration : threshold concentration , : rate parameters regulation functions: 1 1 0 Hill function x Hill-type models 1 0 ramp function x PMA models 0 step function PA models x
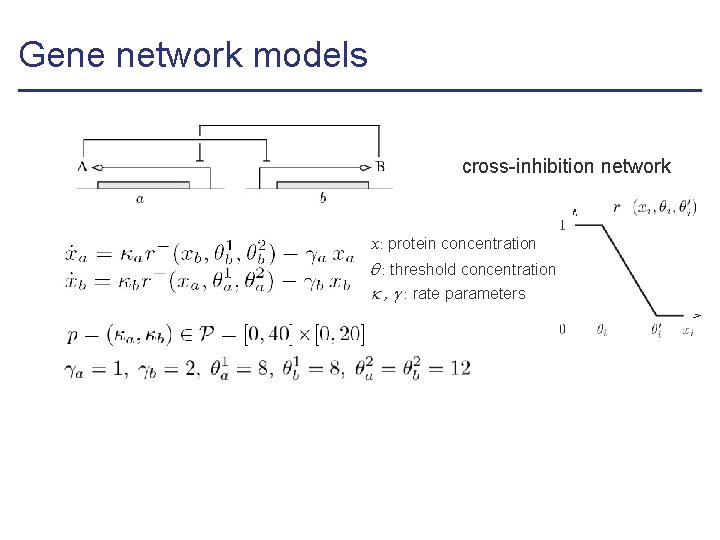
Gene network models cross-inhibition network x: protein concentration : threshold concentration , : rate parameters
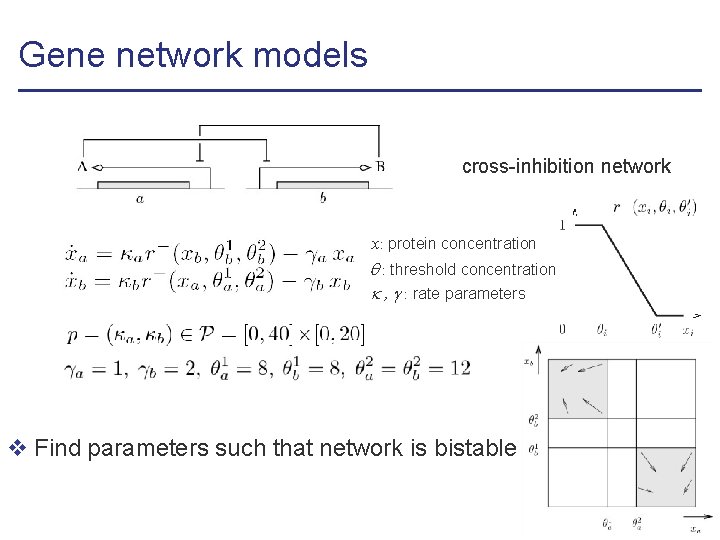
Gene network models cross-inhibition network x: protein concentration : threshold concentration , : rate parameters v Find parameters such that network is bistable
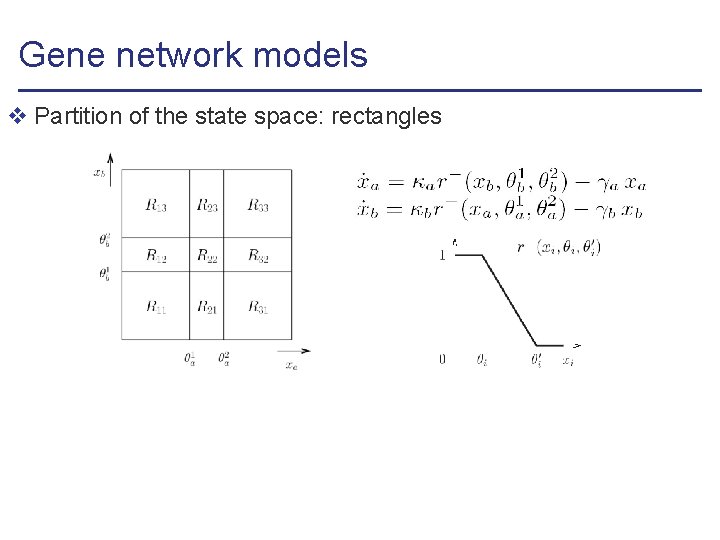
Gene network models v Partition of the state space: rectangles
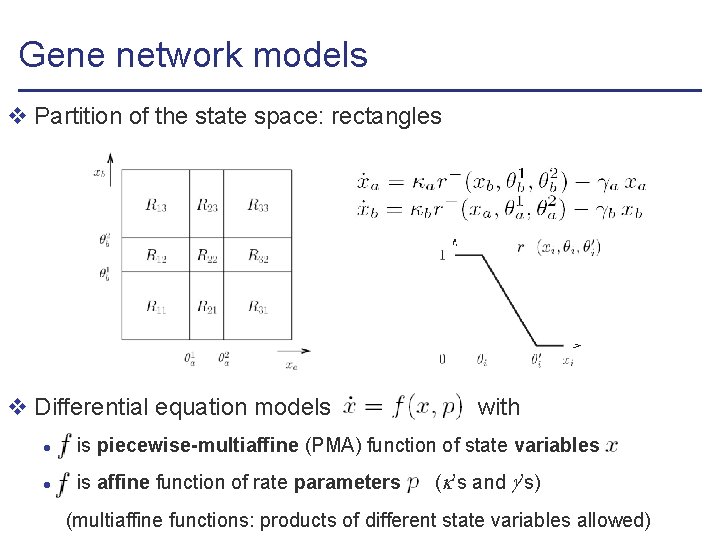
Gene network models v Partition of the state space: rectangles v Differential equation models , with l is piecewise-multiaffine (PMA) function of state variables l is affine function of rate parameters ( ’s and ’s) (multiaffine functions: products of different state variables allowed)

Specifications of dynamical properties v Dynamical properties expressed in temporal logic (LTL) l set of atomic proposition l usual logical operators l temporal operators ,

Specifications of dynamical properties v Dynamical properties expressed in temporal logic (LTL) l set of atomic proposition l usual logical operators l temporal operators , v Semantics of LTL formulas defined over executions of transition systems. .

Specifications of dynamical properties v Dynamical properties expressed in temporal logic (LTL) l set of atomic proposition l usual logical operators l temporal operators , v Semantics of LTL formulas defined over executions of transition systems. . Solution trajectories of PMA models are associated with executions of embedding transition system

Specifications of dynamical properties v Dynamical properties expressed in temporal logic (LTL) l set of atomic proposition l usual logical operators l temporal operators , v Semantics of LTL formulas defined over executions of transition systems bistability property:

Overview I. Introduction II. Problem definition III. Robust design of gene networks IV. Application: tuning a synthetic transcriptional cascade V. Discussion and conclusions
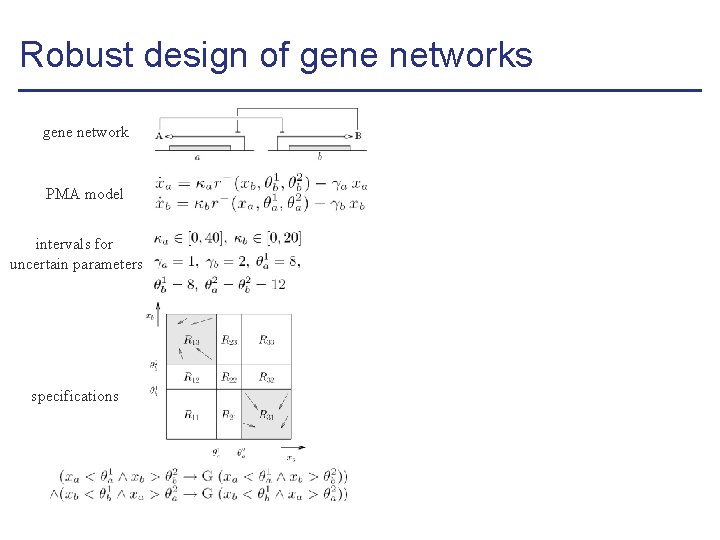
Robust design of gene networks gene network PMA model intervals for uncertain parameters specifications
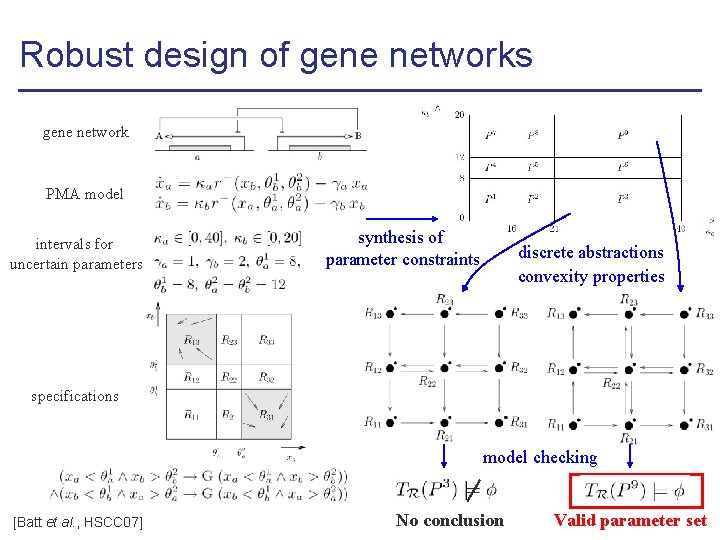
Robust design of gene networks gene network PMA model intervals for uncertain parameters synthesis of parameter constraints discrete abstractions convexity properties specifications model checking [Batt et al. , HSCC 07] No conclusion Valid parameter set

Computation of discrete abstraction Multiaffine function: in every rectangular region, the flow is a convex combination of its values at the vertices [Belta and Habets, Trans. Autom. Contr. , 06]

Computation of discrete abstraction Multiaffine function: in every rectangular region, the flow is a convex combination of its values at the vertices [Belta and Habets, Trans. Autom. Contr. , 06] v Transition between rectangles iff for some parameter, the flow at a common vertex agrees with relative position of rectangles

Computation of discrete abstraction Multiaffine function: in every rectangular region, the flow is a convex combination of its values at the vertices [Belta and Habets, Trans. Autom. Contr. , 06] v Transition between rectangles iff for some parameter, the flow at a common vertex agrees with relative position of rectangles v Transitions can be computed by polyhedral operations where (Because is a piecewise-multiaffine function of x and an affine function of p)

Ro. Ver. Ge. Ne v Approach implemented in publicly-available tool Ro. Ver. Ge. Ne Written in Matlab, exploits polyhedral operation toolbox MPT and model checker Nu. SMV http: //iasi. bu. edu/~batt

Overview I. Introduction II. Problem definition III. Robustness design of gene networks IV. Application: tuning a synthetic transcriptional cascade V. Discussion and conclusions
![Transcriptional cascade: approach Transcriptional cascade [Hooshangi et al, PNAS, 05] v Approach for robust Transcriptional cascade: approach Transcriptional cascade [Hooshangi et al, PNAS, 05] v Approach for robust](http://slidetodoc.com/presentation_image_h2/f07f1134b7ee98fb25b9a709d41a0cc6/image-34.jpg)
Transcriptional cascade: approach Transcriptional cascade [Hooshangi et al, PNAS, 05] v Approach for robust tuning of the cascade: l develop a model of the actual cascade l specify expected behavior l tune network by searching for valid parameter sets l verify robustness of tuned network

Transcriptional cascade: modeling v PMA differential equation model v Parameter identification (1 input and 4 state variables)
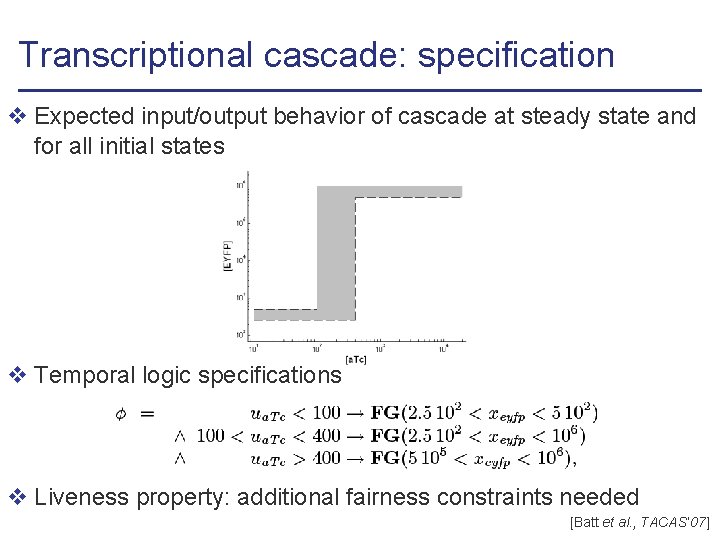
Transcriptional cascade: specification v Expected input/output behavior of cascade at steady state and for all initial states v Temporal logic specifications v Liveness property: additional fairness constraints needed [Batt et al. , TACAS’ 07]

Transcriptional cascade: tuning v Tuning: search for valid parameter sets l Let 3 production rate parameters unconstrained l Answer: 15 sets found (<4 h. , 1500 rectangles, 18 parameter constraints) [Batt et al. , Bioinfo, 07] comparison with numerical simulation results in parameter space and for input/output behavior

Transcriptional cascade: robustness v Robustness: check that tuned network behaves robustly l Let all production and degradation rate parameters range in intervals centered at their reference values (with ± 10% or ± 20% variations) 11 uncertain parameters: l Answer for ± 10% parameter variations: Yes (< 4 hrs) proves that specification holds despite ± 10% parameter variations l Answer for ± 20% parameter variations: No (< 4 hrs) suggests that specification does not hold for some parameters in the ± 20% set (confirmed by manual analysis of counter-example)

Overview I. Introduction II. Problem definition III. Analysis for fixed parameters IV. Analysis for sets of parameters V. Tuning of a synthetic transcriptional cascade VI. Discussion and conclusions

Summary v Gene networks modeled as uncertain PMA systems l piecewise-multiaffine differential equations models l unknown parameters, initial conditions and inputs given by intervals l dynamical properties expressed in temporal logic v Use of tailored combination of parameter constraint synthesis, discrete abstractions, and model checking v Method implemented in publicly-available tool Ro. Ver. Ge. Ne v Approach can answer non-trivial questions on networks of biological interest

Discussion v First computational approach for tuning synthetic gene networks v Related work: l qualitative/discrete approaches (reachability or model checking) [de Jong et al. , Bull. Math. Biol. 04; Ghosh and Tomlin, Syst. Biol. 04; Batt et al. , Bioinfo. 05] [Bernot et al. , J. Theor. Biol. 04; Gonzalez et al. , Biosystems 06, Calzone et al. , Trans. Comput. Syst. Biol 06] l quantitative approaches with fixed parameter values (reachability or MC) [Belta et al. , CDC’ 02; Berman et al. , HSCC’ 07; Fages and Rizk, CMSB’ 07] l quantitative approaches with uncertain parameters (optimisation-based) [Kuepfer et al. , BMC Bioinfo. 07] v Further work: l verification of properties involving timing constraints (post doc, Verimag) l deal with uncertain threshold parameters too l use of compositional verification for design of large modular networks

Acknowledgements Thanks to Calin Belta, Boyan Yordanov, Ron Weiss… Center for Bio. Dynamics Center for Information and Systems Engineering Boston University … and to Ramzi Ben Salah and Oded Maler Verimag Lab Grenoble Polytechnic Institute References: • G. Batt, B. Yordanov, C. Belta and R. Weiss (2007) Robustness analysis and tuning of synthetic gene networks. In Bioinformatics, 23(18): 2415 -1422 • G. Batt, C. Belta and R. Weiss (2007) Temporal logic analysis of gene networks under parameter uncertainty. Accepted to Joint Special Issue on Systems Biology of IEEE Trans. Circuits and Systems and IEEE Trans. Automatic Control
- Slides: 42