RNA interference Forward genetics Reverse genetics phenotype Ricombinazione
RNA interference
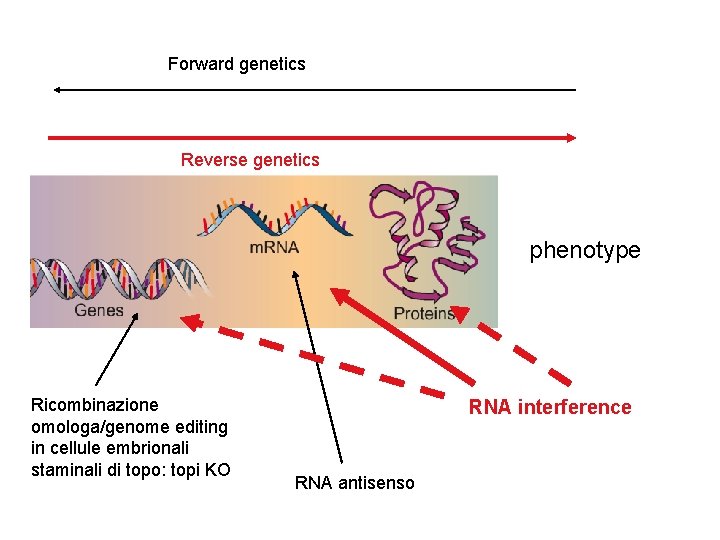
Forward genetics Reverse genetics phenotype Ricombinazione omologa/genome editing in cellule embrionali staminali di topo: topi KO RNA interference RNA antisenso

Discovery of RNA interference: Silencing of target gene fungi plants flies worms mammals Post-Transcriptional Gene Silencing (PTGS): Piante: transgeni ad alto numero di copie e altamente trascritti inducono il silenziamento del gene endogeno Può essere considerato come un primitivo sistema di autodifesa contro RNA esogeno (virus) o RNA endogeno trascritto in modo aberrante (trasposoni)

La Scoperta dell’ RNAi Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans Fire et al, 1998 In C. elegans: introduzione di ds. RNA molto più potente nell’indurre il silenziamento genico rispetto a s. RNA e as. RNA ds. RNA corrispondenti a regioni introniche o al promotore non funzionano ds. RNA microiniettato nelle gonadi induce silenziamento genico in tessuti: amplification/spreading Targeted m. RNA degradation by double stranded RNA in vitro Tuschl et al, 1999 In Drosophila: In vitro: un estratto citoplasmatico di Drosophila + luciferasi m. RNA esogeno + ds. RNA luciferasi: silenziamento genico ds. RNA guida la formazione di un complesso citoplasmatico che determina la degradazione specifica di m. RNA omologo RNA interfernce is mediated by 21 and 22 -nucleotide RNAs Elbashir et al, 2001 In Drosophila: ds. RNA è processato a 21 -22 nt ds. RNA con 5’P, 2 nt protruding 21 -22 -ds. RNA sono sufficienti a indurre RNAi
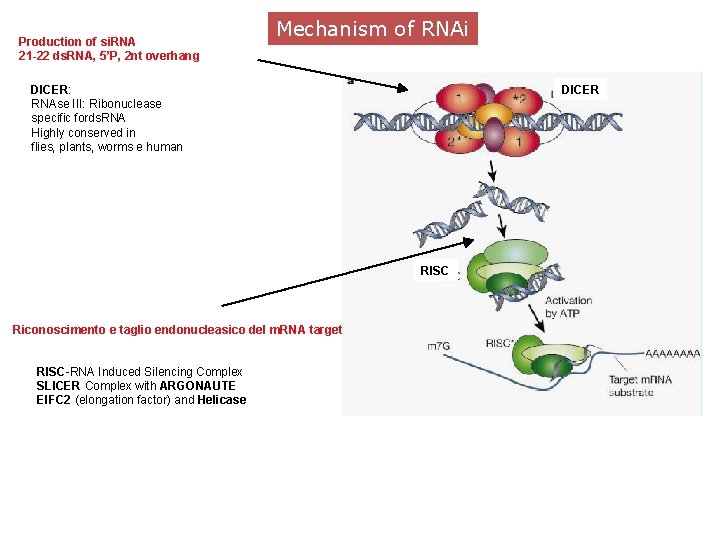
Production of si. RNA 21 -22 ds. RNA, 5’P, 2 nt overhang Mechanism of RNAi DICER: RNAse III: Ribonuclease specific fords. RNA Highly conserved in flies, plants, worms e human DICER RISC Riconoscimento e taglio endonucleasico del m. RNA target RISC-RNA Induced Silencing Complex SLICER Complex with ARGONAUTE EIFC 2 (elongation factor) and Helicase
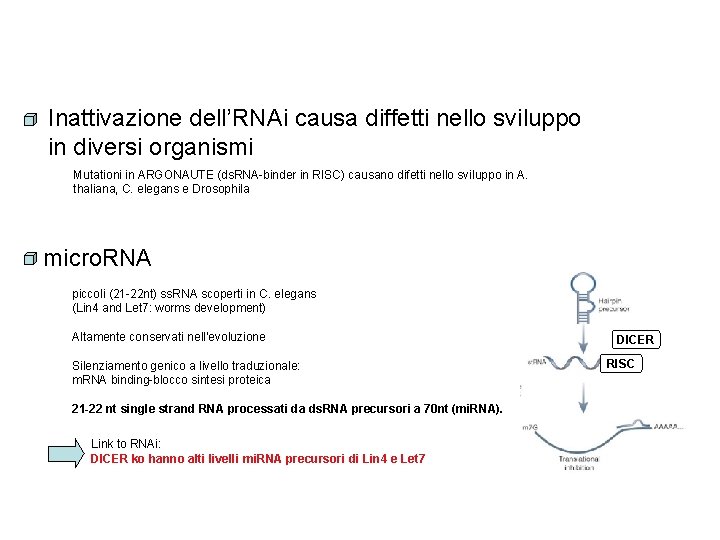
Inattivazione dell’RNAi causa diffetti nello sviluppo in diversi organismi Mutationi in ARGONAUTE (ds. RNA-binder in RISC) causano difetti nello sviluppo in A. thaliana, C. elegans e Drosophila micro. RNA piccoli (21 -22 nt) ss. RNA scoperti in C. elegans (Lin 4 and Let 7: worms development) Altamente conservati nell’evoluzione Silenziamento genico a livello traduzionale: m. RNA binding-blocco sintesi proteica 21 -22 nt single strand RNA processati da ds. RNA precursori a 70 nt (mi. RNA). Link to RNAi: DICER ko hanno alti livelli mi. RNA precursori di Lin 4 e Let 7 DICER RISC

mi. RNA in human

Antisense RNAs in humans
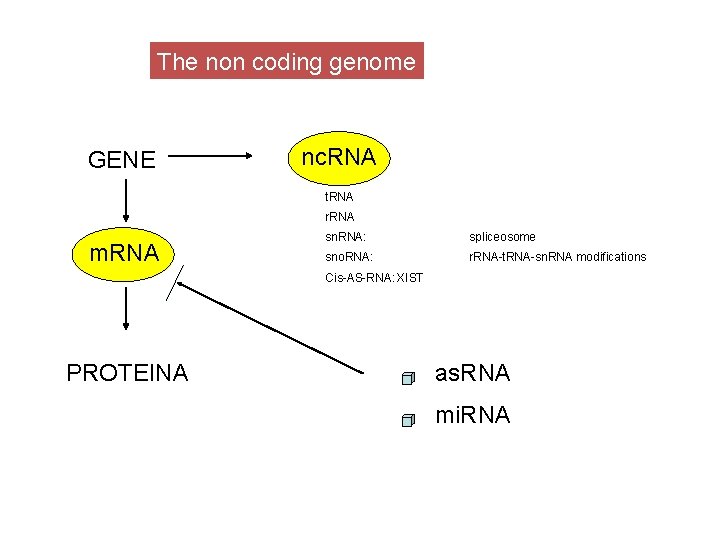
The non coding genome GENE nc. RNA t. RNA r. RNA m. RNA sn. RNA: spliceosome sno. RNA: r. RNA-t. RNA-sn. RNA modifications Cis-AS-RNA: XIST PROTEINA as. RNA mi. RNA

RNAi un nuovo strumento per lo studio della funzione genica nei mammiferi Limitation long ds. RNA >30 nt is effective in mammals, but induce protective pathways Long ds. RNA trigger PKR Phosp of El. F 2 a Block of protein synthesis IFN apoptosis RNAse. L ds. RNA degradation
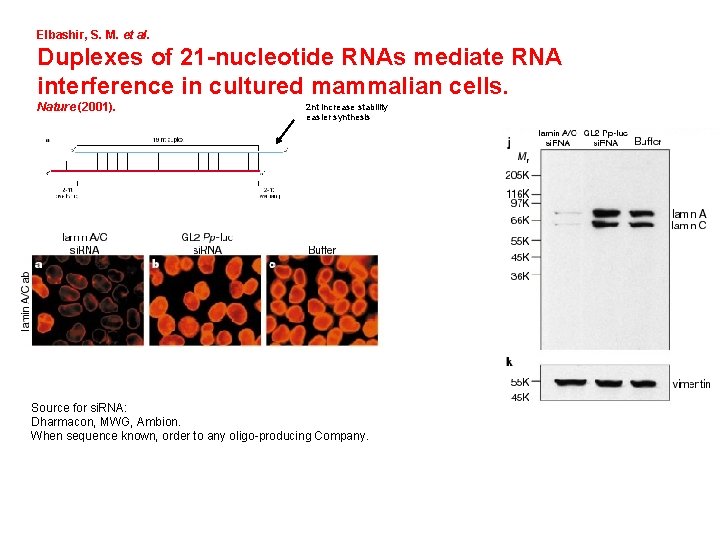
Elbashir, S. M. et al. Duplexes of 21 -nucleotide RNAs mediate RNA interference in cultured mammalian cells. Nature (2001). 2 nt increase stability easier synthesis Source for si. RNA: Dharmacon, MWG, Ambion. When sequence known, order to any oligo-producing Company.
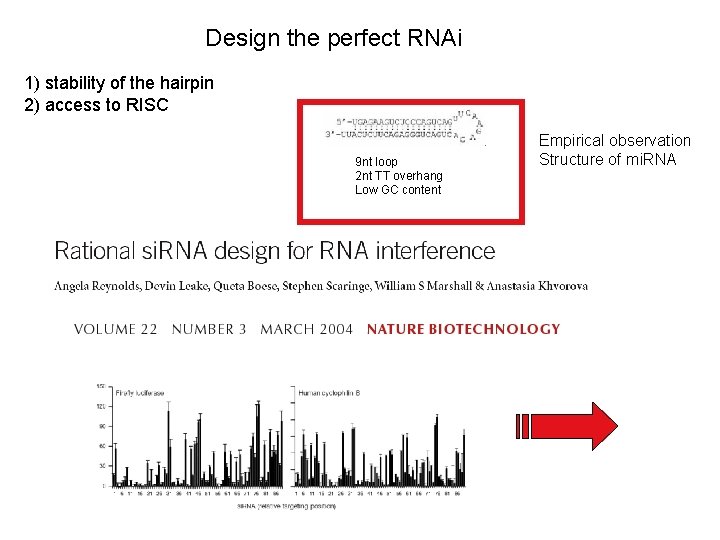
Design the perfect RNAi 1) stability of the hairpin 2) access to RISC 9 nt loop 2 nt TT overhang Low GC content Empirical observation Structure of mi. RNA
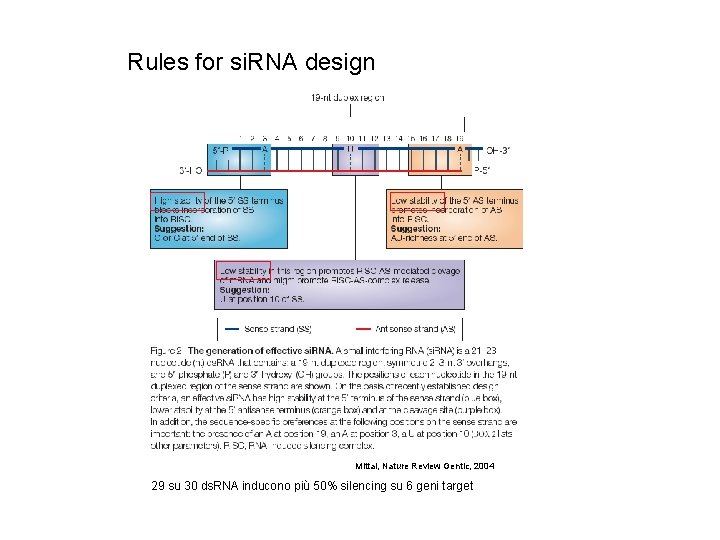
Rules for si. RNA design Mittal, Nature Review Gentic, 2004 29 su 30 ds. RNA inducono più 50% silencing su 6 geni target

1) si. RNA CONTRO PRO si. RNA transfection is very efficient in most cell lines si. RNA silencing is immediate TRANSIENT EFFECT Rate of cell growth? Dilution effect: short time inactivation Ineffectual with Proteis with long half-life EFFICIENCY OF TRANSFECTION Some cells are refractory to transfection Electroporation leads to cell death COST si. RNA are chemically sinthetized, therefore expensive NOT RENEWABLE Use in vivo? ? ? Vettori a DNA per RNAi ?
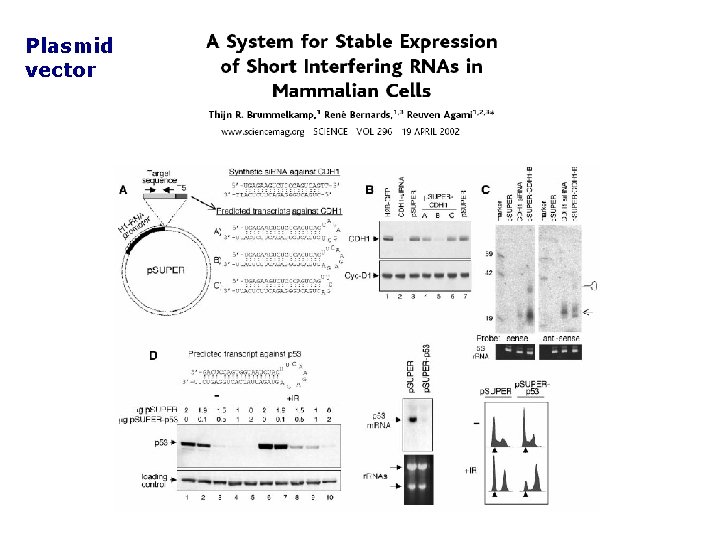
Plasmid vector
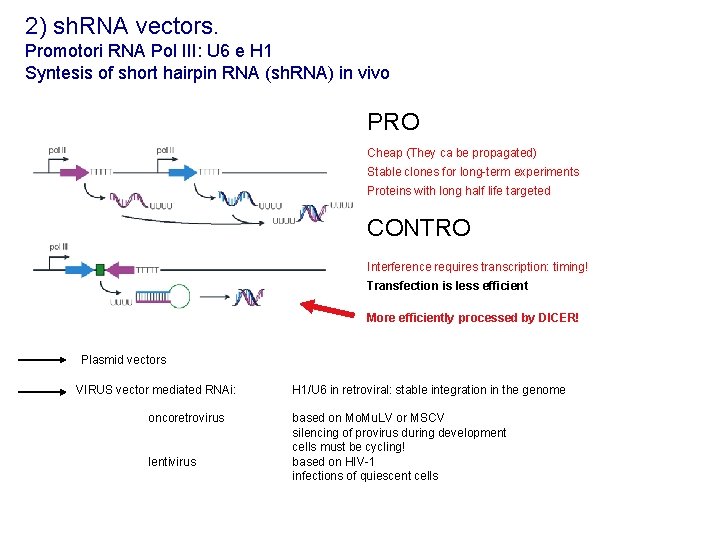
2) sh. RNA vectors. Promotori RNA Pol III: U 6 e H 1 Syntesis of short hairpin RNA (sh. RNA) in vivo PRO Cheap (They ca be propagated) Stable clones for long-term experiments Proteins with long half life targeted CONTRO Interference requires transcription: timing! Transfection is less efficient More efficiently processed by DICER! Plasmid vectors VIRUS vector mediated RNAi: oncoretrovirus lentivirus H 1/U 6 in retroviral: stable integration in the genome based on Mo. Mu. LV or MSCV silencing of provirus during development cells must be cycling! based on HIV-1 infections of quiescent cells
Check Specificity! YES si. RNA against endogenous GFP expression didn’t affecct any other gene NO si. RNA against endogenous gene can affect other genes with weak sequence similarity Use indipendent si. RNA against same target Rescue the observed Loss of Funtion phenotype by expressing the target gene with a silent mutation in the target region

2) Limitations of stable RNAi : impossible to study genes essential for cell survival (housekeeping) or development Vectors for conditional-inducible expression of sh. RNA Tet-OFF/ON H 1 and U 6 promoter system toxicity background
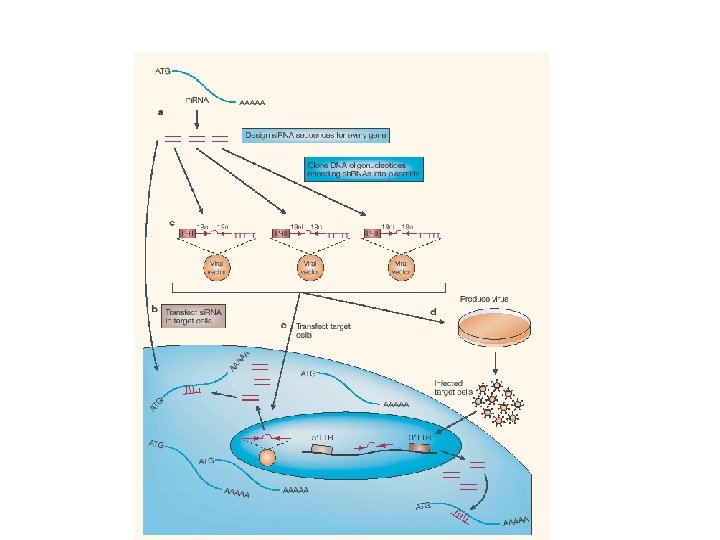
Loss of funtion screenings by RNAi
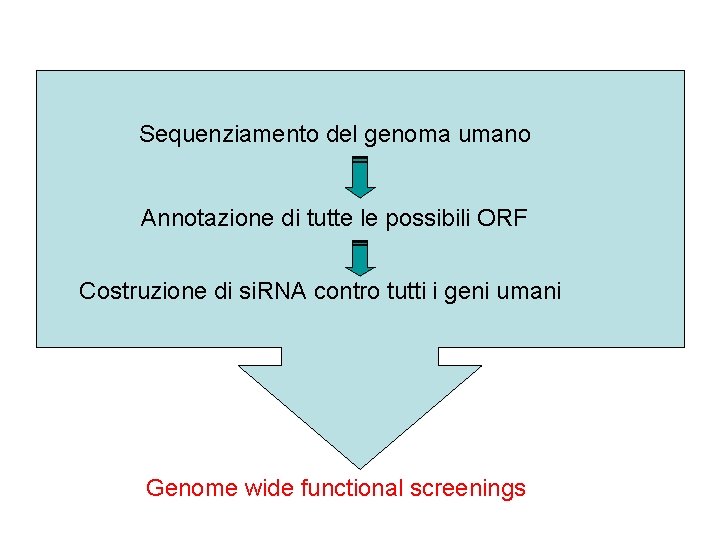
Sequenziamento del genoma umano ? Annotazione di tutte le possibili ORF Costruzione di si. RNA contro tutti i geni umani Genome wide functional screenings
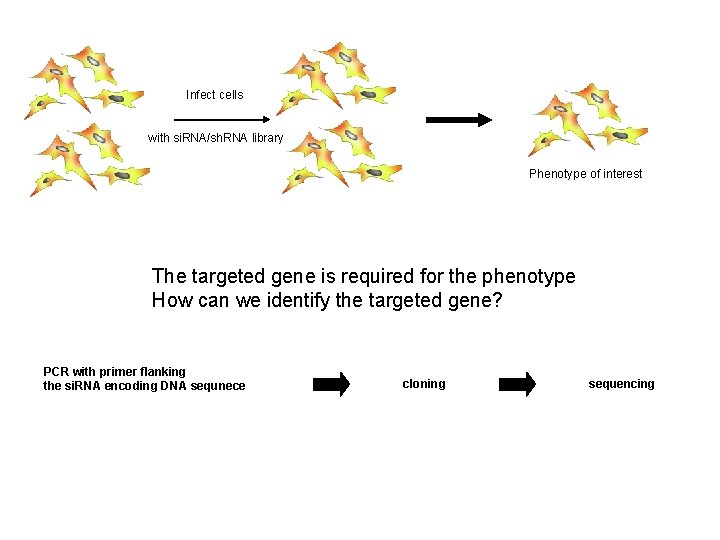
Infect cells with si. RNA/sh. RNA library Phenotype of interest The targeted gene is required for the phenotype How can we identify the targeted gene? PCR with primer flanking the si. RNA encoding DNA sequnece cloning sequencing

High-throughput functional screenings

Limitations: single clone isolation re-cloning and retesting Infect cells select phenotype Single cell derived clones Re-Cloning and sequencing Test again si. RNA BAR-CODE screens Stably integraetd vectors introduce a molecular fingerprint in the infected cell: The 19 hairpin sequence is gene specific and behave as a MOLECULAR BARCODE Relative abundance of BARCODE in a cell population reflect the effect of the knockdown on cell viability Infect cells select population with phenotype PCR of hairpin sequence Fluorescent labeling Hybridize to DNA Microarray containing hairpin sequences from library Identification of highly represented sh. RNA


Silenziamento genico specifico, efficiente e, potenzialmente, stabile nel tempo. Genetica inversa economica e veloce. Rivoluzione nella comprensione dei meccanismi di regolazione dell’espressione genica. Genome-wide functional screenings.
- Slides: 25