RNA editing Liver Intestine By Nesma mohamad What
RNA editing Liver Intestine By Nesma mohamad
What is RNA editing? Discovery of RNA editing sites RNA editing and diseases Significance of RNA editing Mechanism Therapeutic intervention RNA editing and Research References

What IS RNA editing?

RNA editing � Definition: Any process, that results in a change in the sequence of an RNA transcript after it has been generated by RNA polymerase such that it differs from the sequence of the DNA template. so it is any process other than splicing, 5'-capping and polyadenylation Think of splicing as ‘cut and paste, while RNA editing as ‘Spellchecker’ K. Stuart L. Simpson Su, AA; Randau, L (August 2011). "A-to-I and C-to-U editing within transfer RNAs. ". Biochemistry. Biokhimiia. 76 (8): 932– 7 4

RNA editing It is a post transcriptional process with an important role in gene modification In some RNAs, as much as 55% of the nucleotide sequence is not encoded in the (primary) gene, but is added after transcription. Su, AA; Randau, L (August 2011). "A-to-I and C-to-U editing within transfer RNAs. ". Biochemistry. Biokhimiia. 76 (8): 932– 7 5

Discovery of RNA editing

Discovery of RNA Editing in Trypanosome Mitochondria trypanosomes, the kinetoplast exists as a dense granule of DNA within the large mitochondrion � In � The kinetoplast contains circular DNA in two forms, maxicircles and minicircles. ◦ Maxicircles (20 and 40 kb in size), contains most of the genes ◦ Minicircles (1 -3 kb). producing guide RNA (g. RNA) to decode this encrypted maxicircle information, through the insertion or deletion of uridine residues. Gluenz E, et al. (2011). "The kinetoplast replication cycle in Trypanosoma brucei is orchestrated by cytoskeleton-mediated cell morphogenesis". Molecular Cell Biology. 31 (5): 1012– 1021
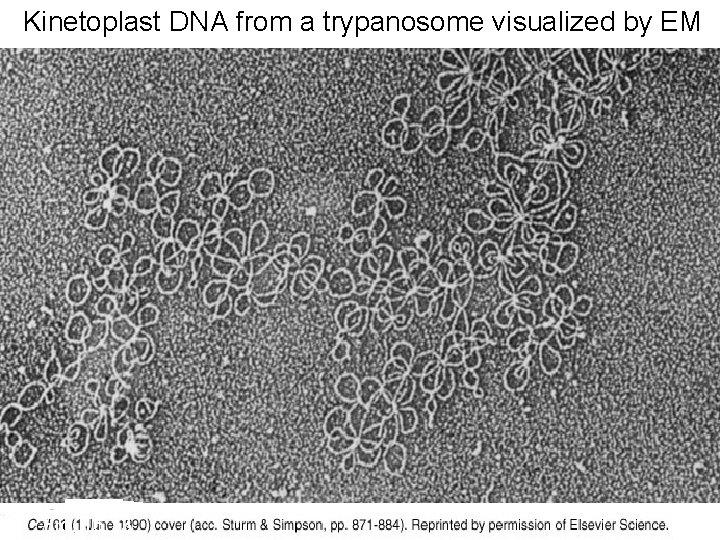
Kinetoplast DNA from a trypanosome visualized by EM Fig. 16. 13
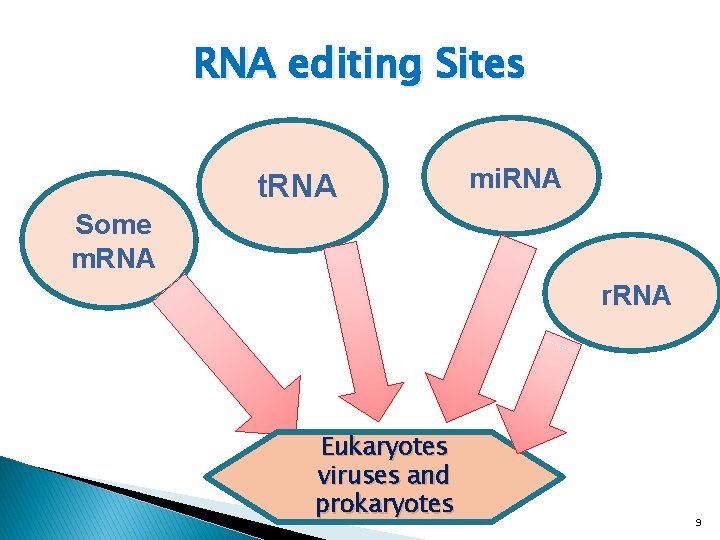
RNA editing Sites t. RNA mi. RNA Some m. RNA r. RNA Eukaryotes viruses and prokaryotes 9

It is also occur in Cell nucleus Cytosol Mitochondria and Plastids Few chloroplast genes of plants. non coding RNAs Kazuko Nishikura (2016). A‑to‑I editing of coding and non‑coding RNAs by ADARs. NATURE REVIEWS | MOLECULAR CELL BIOLOGY. VOLUME 17 | FEBRUARY 2016 | 83 Su, AA; Randau, L (August 2011). "A-to-I and C-to-U editing within transfer RNAs. ". Biochemistry. Biokhimiia. 76 (8): 932– 7. 10

What are the RNA editing mechanisms?

Two mechanisms mediate editing (Conversion) RNA editing by deamination (Insertion / deletion) RNA editing • Li et al. Science doi: 10. 1126/science. 1207018 ; 2011 • Gott & Emeson. Annu Rev Genet. 2000 12

(Conversion / substitution) RNA editing Cytidine to Uridine Editing • Enzyme APOBECs (apolipoprotein B m. RNA editing complex). Adenosie to Inosine Editing • (A-I or A-G) most prevalent in human. • Enzyme ADAR (adenosine deaminases that act on RNA). Alternative editing • Alternative Uridine -to-Cytidine Editing (U-C). • non-classic G-A m. RNA editing.

Now, the question is : What is the difference bet Apo B 100 and Apo B 48 ?
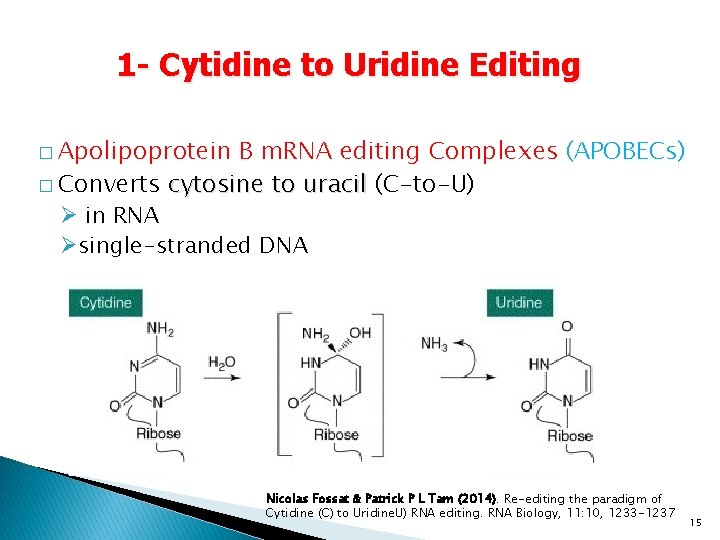
1 - Cytidine to Uridine Editing � Apolipoprotein B m. RNA editing Complexes (APOBECs) � Converts cytosine to uracil (C-to-U) Ø in RNA Øsingle-stranded DNA Nicolas Fossat & Patrick P L Tam (2014). Re-editing the paradigm of Cytidine (C) to Uridine. U) RNA editing. RNA Biology, 11: 10, 1233 -1237 15
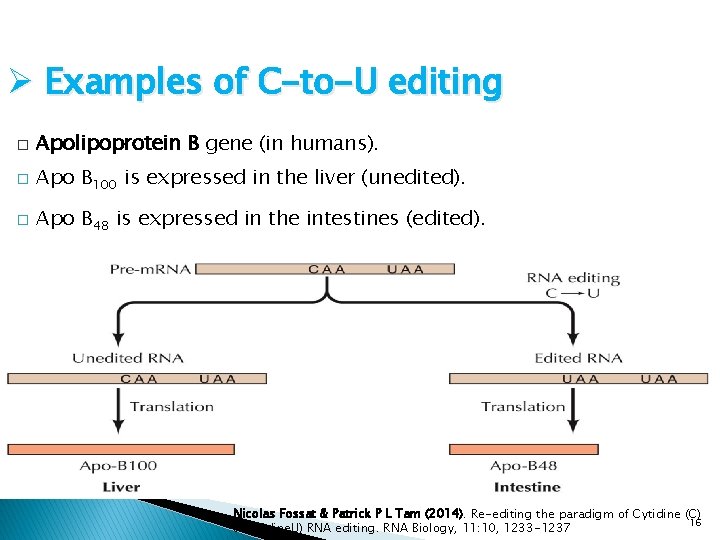
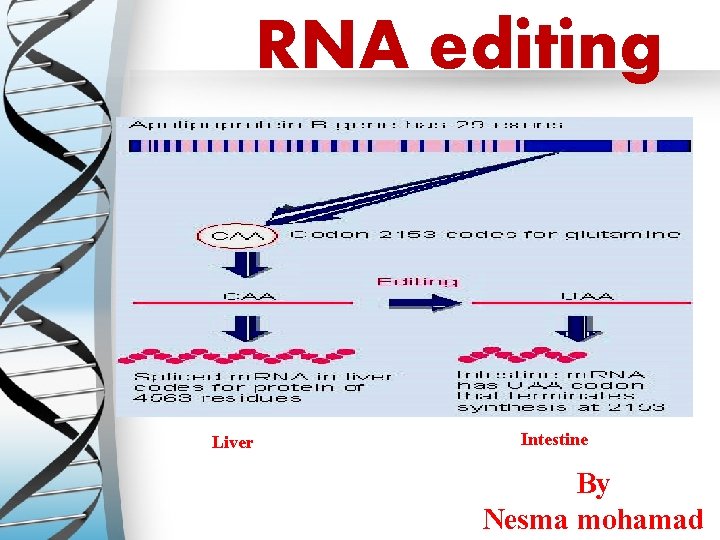
Ø Examples of C-to-U editing � Apolipoprotein B gene (in humans). � Apo B 100 is expressed in the liver (unedited). � Apo B 48 is expressed in the intestines (edited). Nicolas Fossat & Patrick P L Tam (2014). Re-editing the paradigm of Cytidine (C) 16 to Uridine. U) RNA editing. RNA Biology, 11: 10, 1233 -1237
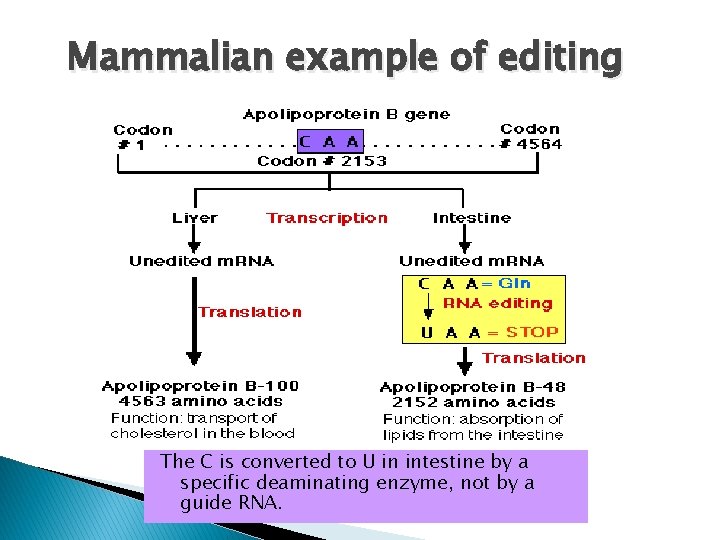
Mammalian example of editing The C is converted to U in intestine by a specific deaminating enzyme, not by a guide RNA.
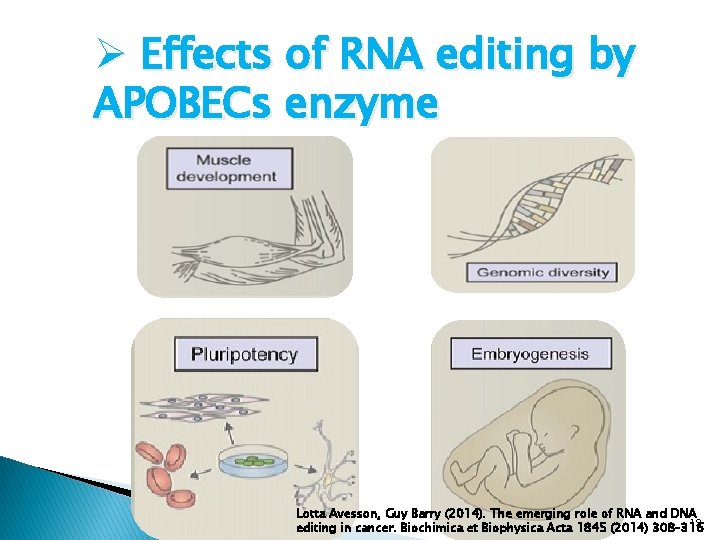
Ø Effects of RNA editing by APOBECs enzyme Lotta Avesson, Guy Barry (2014). The emerging role of RNA and DNA 19 editing in cancer. Biochimica et Biophysica Acta 1845 (2014) 308– 316
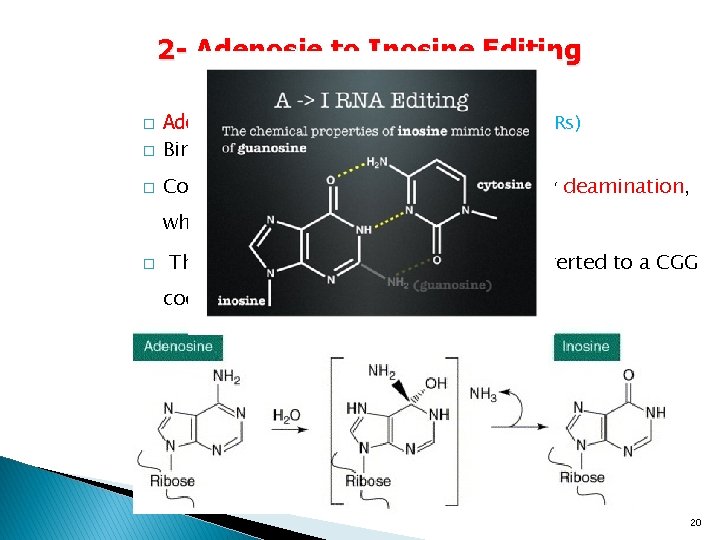
2 - Adenosie to Inosine Editing � � � Adenosine Deaminases Acting on RNA (ADARs) Binds to double stranded RNA Convert adenosine to inosine (A-to-I) by deamination, which the ribosome translates as a G. � Thus a CAG codon (for Gln) can be converted to a CGG codon (for Arg). 20

A-to-I RNA editing: two forms - Promiscuous and Specific ü The promiscuous Occurs within longer duplexes of hundreds of nucleotides (e. g. , Alu elements). (e. g. , pre- or pri-mi. RNAs, ü duplexes arising from transgene or viral expression, duplexes arising from paired repetitive elements). Up to 60% of adenosines could be edited. ü The specific Occurs in short duplexes RNA regions (a single adenosine is edited within the stretch of ds. RNA) Up to 10% of their adenosines selectively could be edited. eg. A-to-I RNA editing in mi. RNA. A-to-I RNA editing sites also abundantly occur in Intronic regions as well as in 3′-UTRs. Fig: Substrates of ADARs Giovanni Nigita, 1, † Salvatore Alaimo, 2, † Alfredo Ferro (2015). Knowledge in the Investigation of A-to-I RNA Editing Signals. Front Bioeng Biotechnol. 2015; 3: 18.
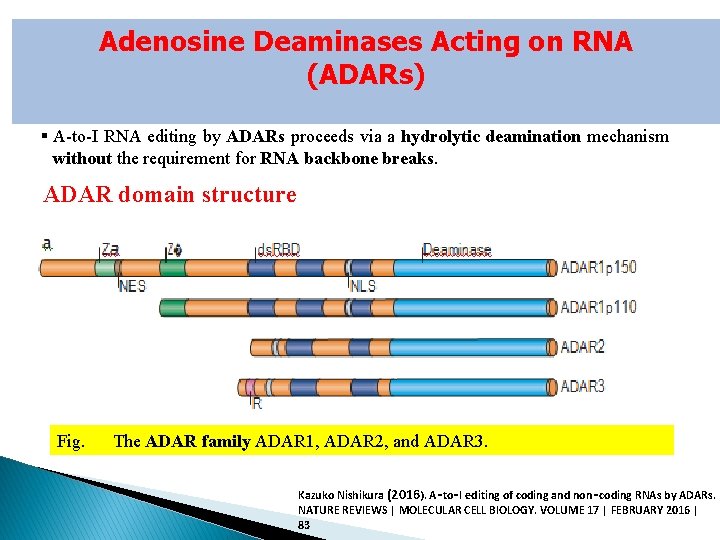
Adenosine Deaminases Acting on RNA (ADARs) § A-to-I RNA editing by ADARs proceeds via a hydrolytic deamination mechanism without the requirement for RNA backbone breaks. ADAR domain structure Fig. The ADAR family ADAR 1, ADAR 2, and ADAR 3. Kazuko Nishikura (2016). A‑to‑I editing of coding and non‑coding RNAs by ADARs. NATURE REVIEWS | MOLECULAR CELL BIOLOGY. VOLUME 17 | FEBRUARY 2016 | 83

SO, Site : selective deamination of A into I enabling functionally different proteins to arise from a single gene. Barbon A. and Barlati S. ( 2011). Glutamate Receptor RNA Editing in Health and Disease. ISSN 00062979, Biochemistry (Moscow), Vol. 76, No. 8, pp. 882889

Alteration of coding capacity Altered splicing. Altered mi. RNA or si. RNA target populations Effects of A-to-I editing Inhibition of mi. RNA and si. RNA processing Heterochromatin formation Nuclear sequestration Cytoplasmic sequestration Garrett, S. ; Rosenthal, J. J. C. (5 January 2012). "RNA Editing Underlies Temperature Adaptation in K+ Channels from Polar Octopuses". Science. 335: 848– 851.

Effects of RNA editing by ADARs enzyme Lotta Avesson, Guy Barry (2014). The emerging role of RNA and DNA editing in cancer. Biochimica et Biophysica Acta 1845 (2014) 308– 316

Ø Examples of (A-to-I) editing: 26

Glutamate receptor L-glutamate is the predominant excitatory neurotransmitter in vertebrate nervous systems. RNA editing affects protein function. In mammals, particular attention is given to the Glu. A 2, an ionotropic glutamate receptor subunit, Channels open and close, or gate, in response to an extrinsic factor, such as concentration of a specific ligand (eg. ionotropic glutamate receptors). Four basic groups of ionotropic glutamate receptors: α-amino-3 -hydroxy-5 -methyl-4 -isoxazolepropionic acid receptor (AMPA) receptors, kinate receptors, Delta receptors and N-methyl-d-aspartate (NMDA) receptors. Four AMPA subunits: Glu. A 1– 4 Five Kainate receptor subunits: Glu. K 1– 5 Barbon A. and Barlati S. (2011). Glutamate Receptor RNA Editing in Health and Disease. Biochemistry (Moscow), Vol. 76, No. 8, pp. 882 -889. 27

Regulation of Ion Channel and Transporter Function Through RNA Editing Glu. A 1 (AMPA receptor) m. RNAs in humans – no R/G editing Fig: Schematic glutamate receptor subunit structure. Svenja Pachernegg, Yvonne Münster. Elke Muth-Köhne, Gloria Fuhrmann and Michael Hollmann 1(2015): Glu. A 2 is rapidly edited at the Q/R site during neural differentiation in vitro. Front. Cell. Neurosci. , 05 March 2015
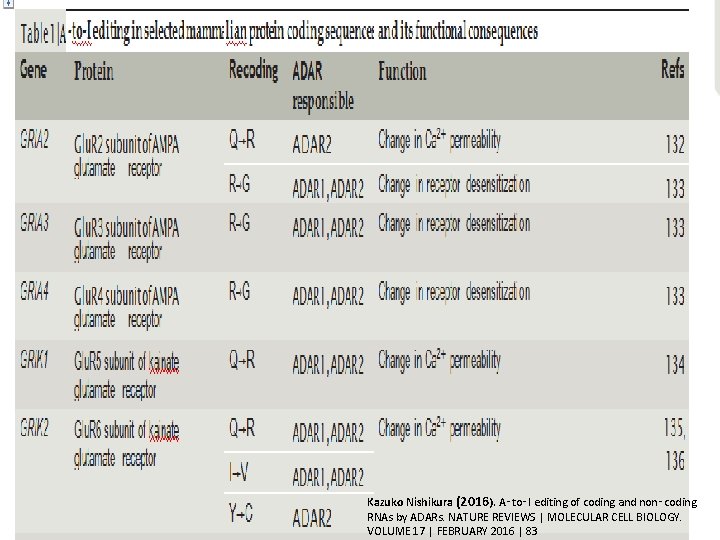
Kazuko Nishikura (2016). A‑to‑I editing of coding and non‑coding RNAs by ADARs. NATURE REVIEWS | MOLECULAR CELL BIOLOGY. 29 VOLUME 17 | FEBRUARY 2016 | 83

VOLTAGE-ACTIVATED POTASSIUM CHANNELS Voltage-gated potassium channels (VGKCs) are transmembrane channels specific for potassium and sensitive to voltage changes in the cell's membrane potential. They open during depolarization followed by fast inactivation, This due to pore blockage by the inactivating particle, which decreases or completely abolishes the K conductance The human Kv 1. 1 potassium channel undergoes RNA editing in its pore domain. Editing replaces an isoleucine with a valine that results in rapid recovery from fast inactivation. Barbon A. and Barlati S. (2011). Glutamate Receptor RNA Editing in Health and Disease. Biochemistry (Moscow), Vol. 76, No. 8, pp. 882 -889. 30
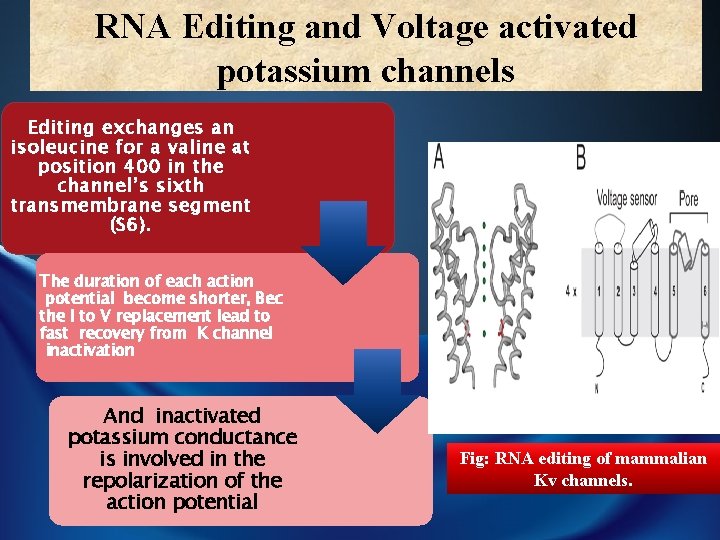
RNA Editing and Voltage activated potassium channels Editing exchanges an isoleucine for a valine at position 400 in the channel’s sixth transmembrane segment (S 6). The duration of each action potential become shorter, Bec the I to V replacement lead to fast recovery from K channel inactivation And inactivated potassium conductance is involved in the repolarization of the action potential Fig: RNA editing of mammalian Kv channels.
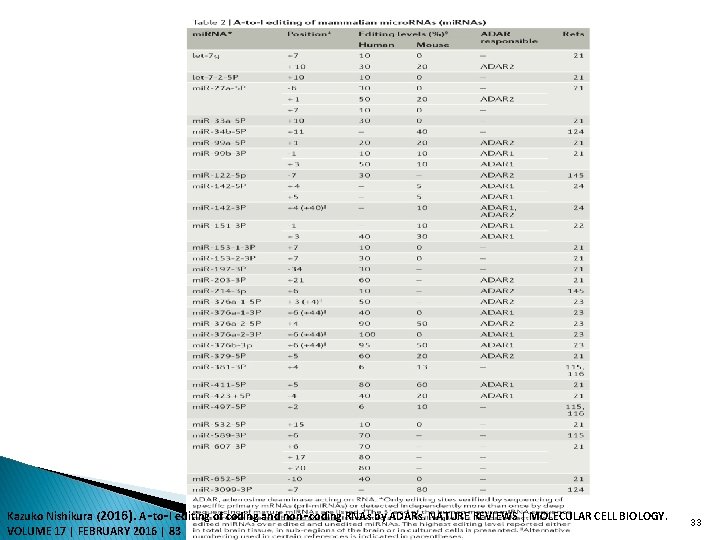
Kazuko Nishikura (2016). A‑to‑I editing of coding and non‑coding RNAs by ADARs. NATURE REVIEWS | MOLECULAR CELL BIOLOGY. VOLUME 17 | FEBRUARY 2016 | 83 33

3 - Alternative m. RNA editing Alternative U -to-C m. RNA editing non-classic GA m. RNA editing • It was first reported in • (Wilms Tumor-1) transcripts • It also seen alongside (U-to. C) alterations in brain cell TPH 2(tryptophan hydroxylase 2) transcri pts. Grohmann M. ; Hammer P. ; Walther M. ; Paulmann N. ; Buttner A. ; Eisenmenger W. ; et al. (2010). "Alternative splicing and extensive RNA editing of human TPH 2 transcripts". PLOS ONE. 5: e 8956. 34

Two mechanisms mediate editing (Conversion) RNA editing by deamination (Insertion / deletion) RNA editing • Li et al. Science doi: 10. 1126/science. 1207018 ; 2011 • Gott & Emeson. Annu Rev Genet. 2000 35

B. Insertion/Deletion Editing Ø It is called "pan-editing“ because This may involve a large fraction of the sites in a gene. Ø Requires a special class of RNA called guide RNA (g. RNA). Ø Multiple U’s are inserted into specific region of m. RNAs after transcription (or U’s may be deleted). 36

Insertion/Deletion Editing 37
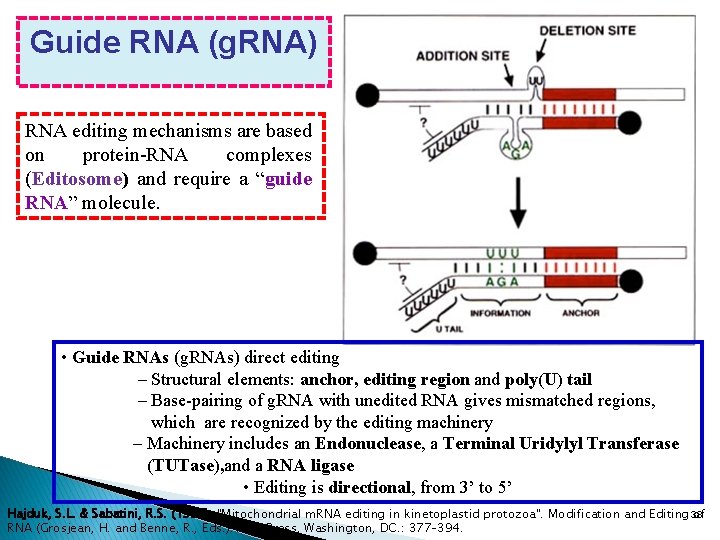
Guide RNA (g. RNA) RNA editing mechanisms are based on protein-RNA complexes (Editosome) and require a “guide RNA” molecule. • Guide RNAs (g. RNAs) direct editing – Structural elements: anchor, editing region and poly(U) tail – Base-pairing of g. RNA with unedited RNA gives mismatched regions, which are recognized by the editing machinery – Machinery includes an Endonuclease, a Terminal Uridylyl Transferase (TUTase), and a RNA ligase • Editing is directional, from 3’ to 5’ Hajduk, S. L. & Sabatini, R. S. (1998). "Mitochondrial m. RNA editing in kinetoplastid protozoa". Modification and Editing 38 of RNA (Grosjean, H. and Benne, R. , Eds. ). ASM Press, Washington, DC. : 377– 394.
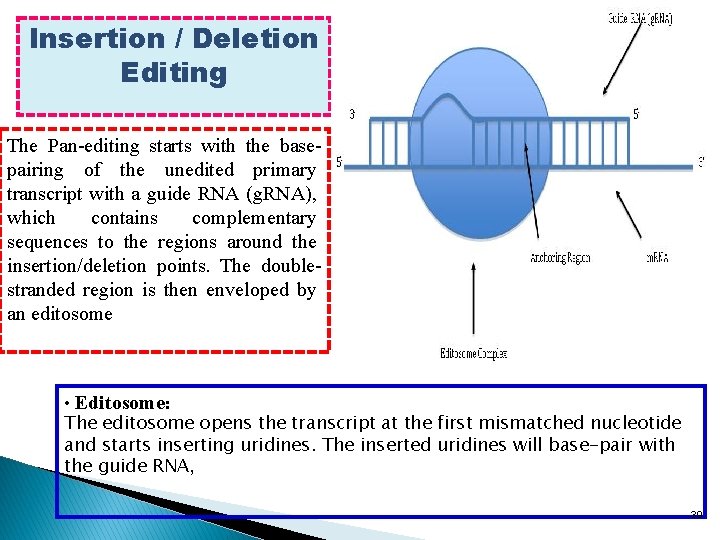
Insertion / Deletion Editing The Pan-editing starts with the basepairing of the unedited primary transcript with a guide RNA (g. RNA), which contains complementary sequences to the regions around the insertion/deletion points. The doublestranded region is then enveloped by an editosome • Editosome: The editosome opens the transcript at the first mismatched nucleotide and starts inserting uridines. The inserted uridines will base-pair with the guide RNA, 39
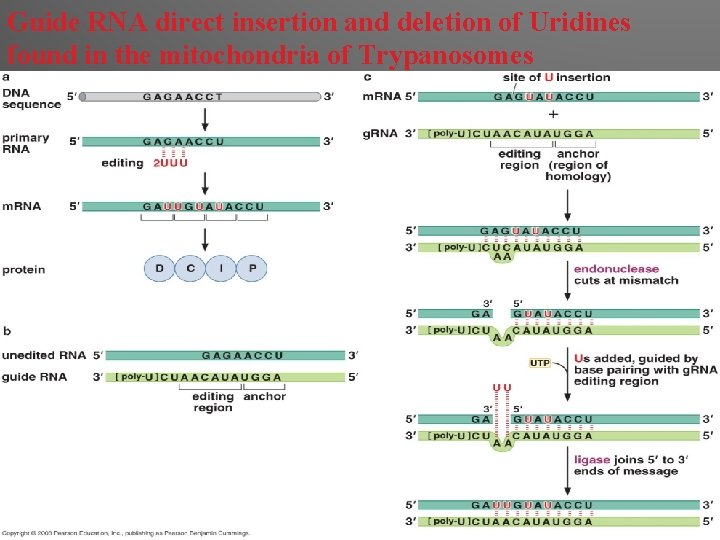
Guide RNA direct insertion and deletion of Uridines found in the mitochondria of Trypanosomes

Significans of RNA editing

Significance of RNA editing • RNA editing mutant strong defects in organelle development. • Deficiency cancers) diseases(mostly • Increase the number of different proteins without the need to increase the number of genes in the genome. 42
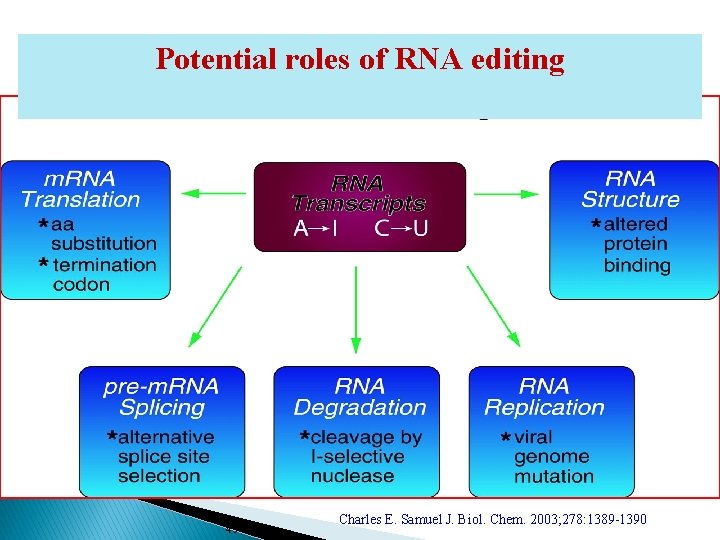
Potential roles of RNA editing 43 Charles E. Samuel J. Biol. Chem. 2003; 278: 1389 -1390

RNA editing and Diseases
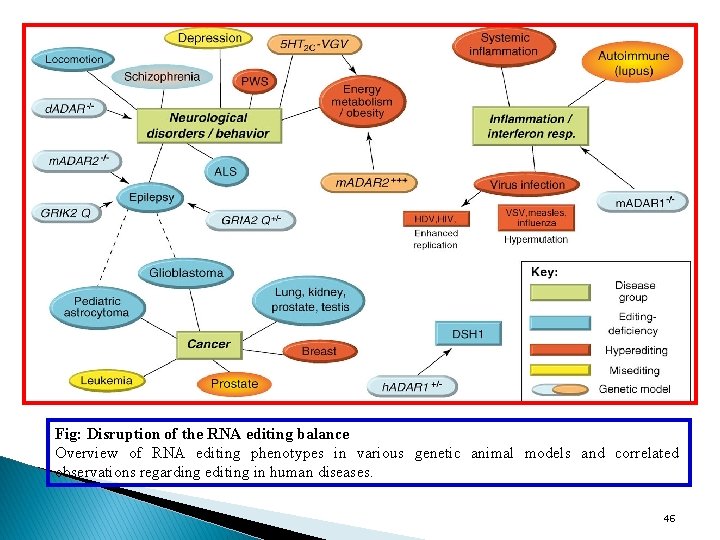
Fig: Disruption of the RNA editing balance Overview of RNA editing phenotypes in various genetic animal models and correlated observations regarding editing in human diseases. 46
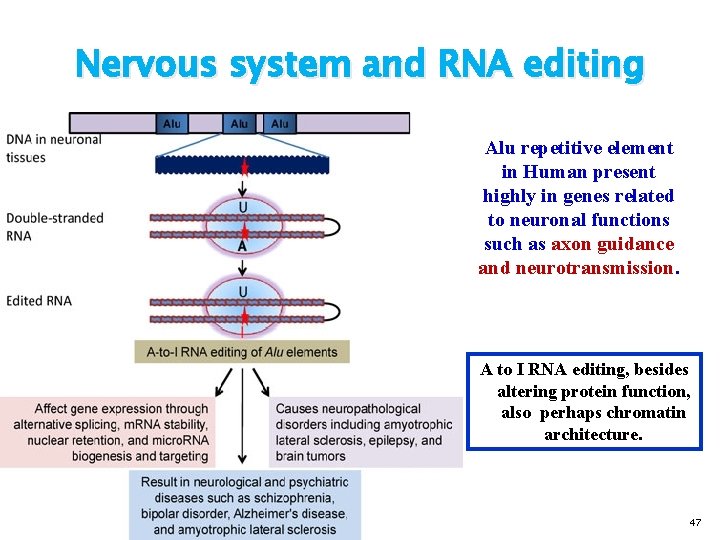
Nervous system and RNA editing Alu repetitive element in Human present highly in genes related to neuronal functions such as axon guidance and neurotransmission. A to I RNA editing, besides altering protein function, also perhaps chromatin architecture. 47

Immune system and RNA editing ADAR 1 -S and ADAR 1 -L involved in HDV editing APOBEC 3 G ØPlay a vital role in innate defense against viruses ØHas antiviral properties (inactivate the HIV) Qingde Wang *, Xiaoni Li, Ruofan Qi and Timothy Billiar (2017): RNA Editing, ADAR 1, and the Innate. Immune Response. Genes, 8, 41 48

RNA editing and cancers A-to-I A class of phospholipids found in mammals. editing is involved in They represent 80– 90% of the myelin membrane phospholipids human embryogenesis. RNA regulation can modulate the expression of oncogenes or tumor suppressor genes So A-to-I editing may contribute to cancer development and progression 49
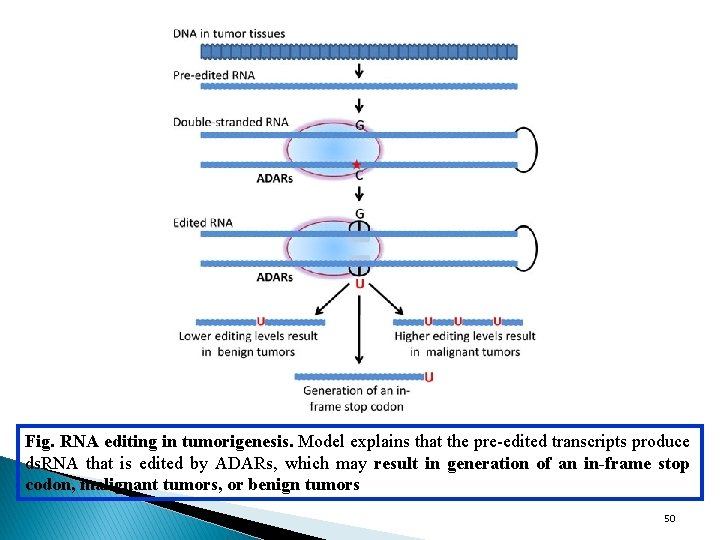
Fig. RNA editing in tumorigenesis. Model explains that the pre-edited transcripts produce ds. RNA that is edited by ADARs, which may result in generation of an in-frame stop codon, malignant tumors, or benign tumors 50

RNA editing and cancers: more examples ADARs APOBECs Brain Reduced editing of Glu. R in malignant gliomas Bladder Reduced editing levels in tumors Leukemia Disease-associated alternative splicing of PTPN 6 caused by RNA hyper-editing Liver APOBEC 1 -mediated editing of NAT 1 m. RNA in liver tumors Breast APOBEC 3 B upregulated in breast cancer cell lines General APOBEC signatures are observed in multiple cancer types Lotta Avesson, Guy Barry (2014). The emerging role of RNA and DNA editing 51 in cancer. Biochimica et Biophysica Acta 1845 (2014) 308– 316

52

RNA editing And therapeutic intervension

Therapeutic intervention of RNA editing • Therapeutic target for CNS disorders • Drugs for inhibiting RNA editing enzymes • Editing enzyme levels could be used as prognostic markers of cancer or determine post-therapeutic outcome. 54
55

56

RNA editing And Research

58
59

60

61
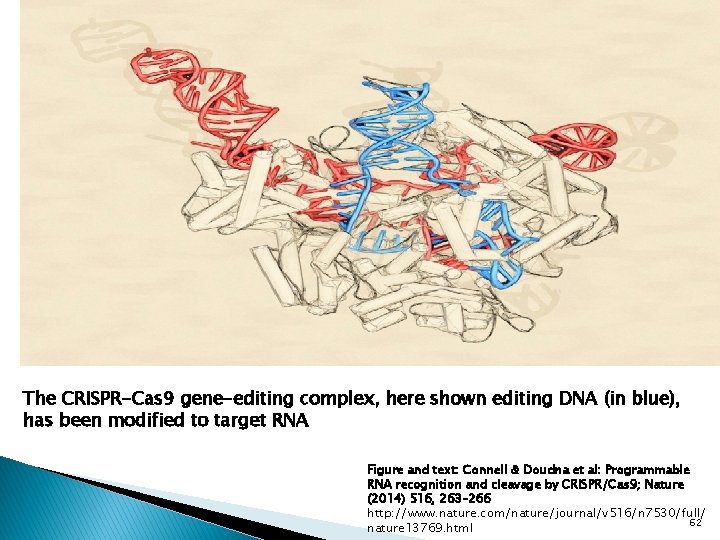
The CRISPR-Cas 9 gene-editing complex, here shown editing DNA (in blue), has been modified to target RNA Figure and text: Connell & Doudna et al: Programmable RNA recognition and cleavage by CRISPR/Cas 9; Nature (2014) 516, 263– 266 http: //www. nature. com/nature/journal/v 516/n 7530/full/ 62 nature 13769. html

References 1. Burri, PH; Hlushchuk, R; Djonov, V (2004). "Intussusceptive angiogenesis: its emergence, its characteristics, and its significance". Dev Dyn. 231 (3): 474– 88. 2. Klimek-Tomczak K. ; Mikula M. ; Dzwonek A. ; Paziewska A. ; Karczmarski J. ; Hennig E. ; et al. (2006). "Editing of hn. RNP K protein m. RNA in colorectal adenocarcinoma and surrounding mucosa". Br. J. Cancer. 94 (4): 586– 92. 3. Grohmann M. ; Hammer P. ; Walther M. ; Paulmann N. ; Buttner A. ; Eisenmenger W. ; et al. (2010 "Alternative splicing and extensive RNA editing of human TPH 2 transcripts". PLOS ONE. 5: e 8956. 4. Lotta Avesson, Guy Barry : The emerging role of RNA and DNA editing in cancer. Review Article. Biochimica et Biophysica Acta (BBA) – Review on Cancer, Volume 1845, Issue 2, April 2014, Pages 308 -316 5. Huang WH, Tseng CN, Tang JY, Yang CH, Liang SS, Chang HW. RNA editing and drug discovery for cancer therapy. Scientific World Journal. 2013 Apr 24; 2013: 804505. doi: 10. 1155/2013/804505. Print 2013. Review. 6. Su, AA; Randau, L (August 2011). "A-to-I and C-to-U editing within transfer RNAs. ". Biochemistry. Biokhimiia. 76 (8): 932– 7 7. Gluenz E, et al. (2011). "The kinetoplast replication cycle in Trypanosoma brucei is orchestrated by cytoskeleton-mediated cell morphogenesis". Molecular Cell Biology. 31 (5): 1012– 1021

References 8. Connell & Doudna et al (2014 ): Programmable RNA recognition and cleavage by CRISPR/Cas 9; Nature) 516, 263– 266 9. Qingde Wang *, Xiaoni Li, Ruofan Qi and Timothy Billiar (2017): RNA Editing, ADAR 1, and the Innate. Immune Response. Genes, 8, 41 10. Charles E. Samuel J. Biol. Chem. 2003; 278: 1389 -1390 11. Hajduk, S. L. & Sabatini, R. S. (1998). "Mitochondrial m. RNA editing in kinetoplastid protozoa". Modification and Editing of RNA (Grosjean, H. and Benne, R. , Eds. ). ASM Press, Washington, DC. : 377– 394. 12. Grohmann M. ; Hammer P. ; Walther M. ; Paulmann N. ; Buttner A. ; Eisenmenger W. ; et al. (2010). "Alternative splicing and extensive RNA editing of human TPH 2 transcripts". PLOS ONE. 5: e 8956 13. Kazuko Nishikura (2016). A‑to‑I editing of coding and non‑coding RNAs by ADARs. NATURE REVIEWS | MOLECULAR CELL BIOLOGY. VOLUME 17 | FEBRUARY 2016 | 83 14. Nicolas Fossat & Patrick P L Tam (2014). Re-editing the paradigm of Cytidine (C) to Uridine. U) RN editing. RNA Biology, 11: 10, 1233 -1237

- Slides: 62