RNA catalysis Understand the basics of RNADNA catalysts

RNA catalysis Understand the basics of RNA/DNA catalysts - what functional groups used for catalysis? structures formed? Know about transesterification & cleavage reactions Know four types of natural catalytic RNAs (group I introns, group II introns, RNase P, small self-cleaving), what reactions they perform, know basics of their secondary and tertiary structure, requirements for cofactors/metals/proteins/ATP Know details of glm. S ribozyme self-cleavage Understand use of ribozymes as therapeutics In vitro selection - understand the process Know some of the ribozymes and deoxyribozymes that have been discovered using in vitro selection

Outline • RNA transesterification • Naturally occurring catalysts • Catalytic functions • Catalytic mechanisms
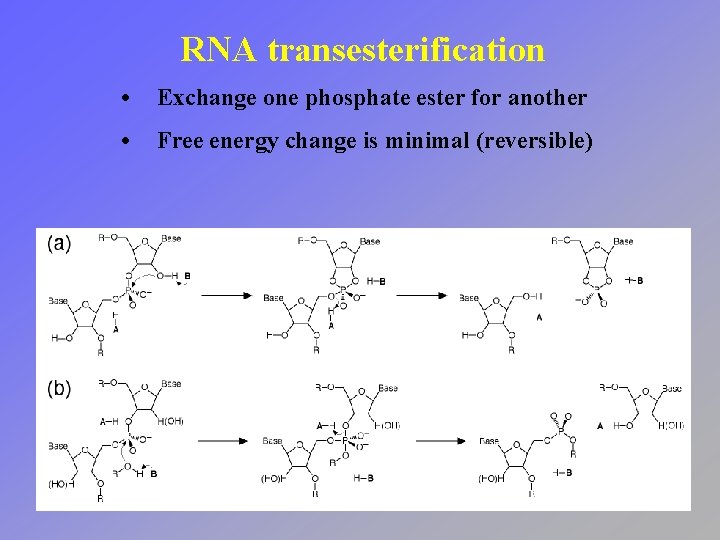
RNA transesterification • Exchange one phosphate ester for another • Free energy change is minimal (reversible)
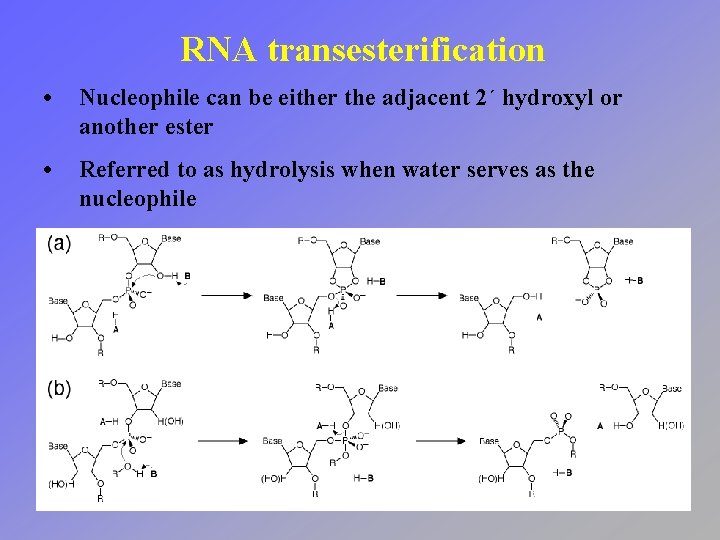
RNA transesterification • Nucleophile can be either the adjacent 2´ hydroxyl or another ester • Referred to as hydrolysis when water serves as the nucleophile
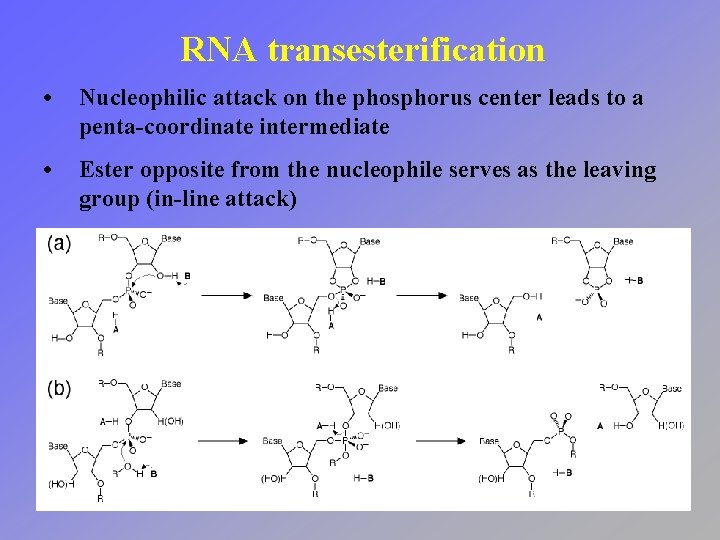
RNA transesterification • Nucleophilic attack on the phosphorus center leads to a penta-coordinate intermediate • Ester opposite from the nucleophile serves as the leaving group (in-line attack)
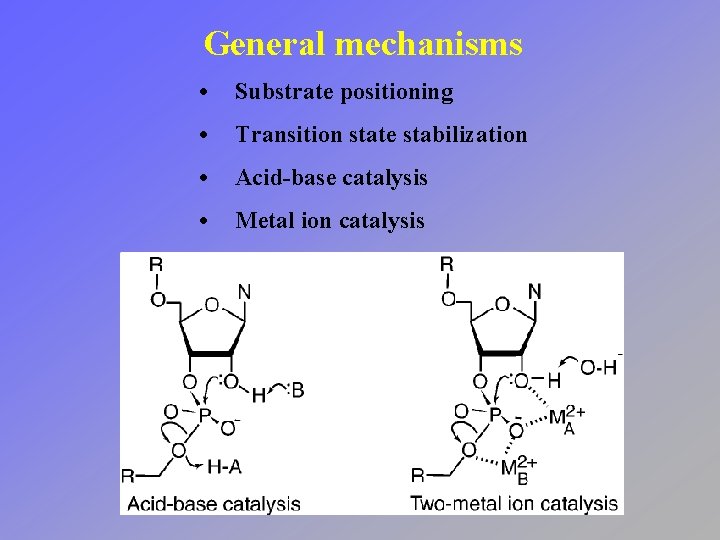
General mechanisms • Substrate positioning • Transition state stabilization • Acid-base catalysis • Metal ion catalysis

RNA Catalysts

Naturally occurring catalysts • RNA cleavage glm. S ribozyme (crystal structure) hammerhead ribozyme (crystal structure) hairpin ribozyme (crystal structure) Varkud satellite (VS) ribozyme (partial NMR structure) hepatitis delta virus (HDV) ribozyme (crystal structure) M 1 RNA (RNase P) (partial crystal structure) • RNA splicing group I introns (crystal structure) group II introns (crystal structure) *** U 2 -U 6 sn. RNA (spliceosome) (partial NMR structure) *** • Peptide bond formation ribosome (crystal structure)
Small self-cleaving ribozymes • Hammerhead, hairpin, VS, HDV ribozymes • Derivative of viral, viroid, or satellite RNAs • Involved in RNA processing during rolling circle replication • RNA transesterification via 2´ hydroxyl • Reversible: cleavage and ligation (excepting HDV)

Hammerhead ribozyme • Three-stem junction with conserved loop regions • Coaxial stacking of stems II and III through extended stem II structure containing canonical Watson-Crick and non-canonical base pairs • Metal-ion catalysis
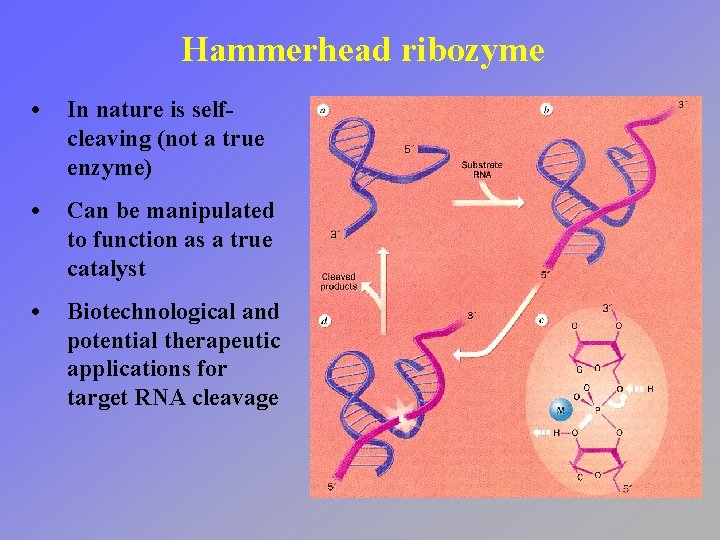
Hammerhead ribozyme • In nature is selfcleaving (not a true enzyme) • Can be manipulated to function as a true catalyst • Biotechnological and potential therapeutic applications for target RNA cleavage

Hammerhead ribozyme • Separation of catalytic and substrate strands • Strand with hairpin is the enzyme • Single strand is substrate • KM = 40 n. M; kcat = ~1 min-1; kcat/KM = ~107 M -1 min -1 (catalytic efficiency) • Compare to protein enzymes?
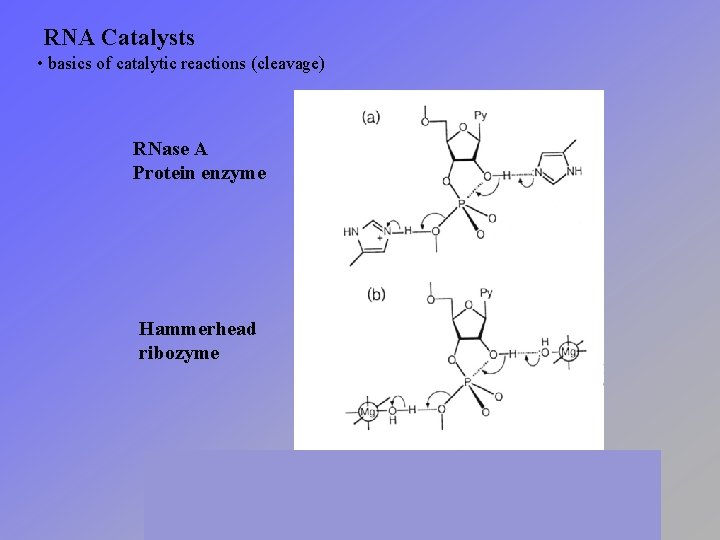
RNA Catalysts • basics of catalytic reactions (cleavage) RNase A Protein enzyme Hammerhead ribozyme
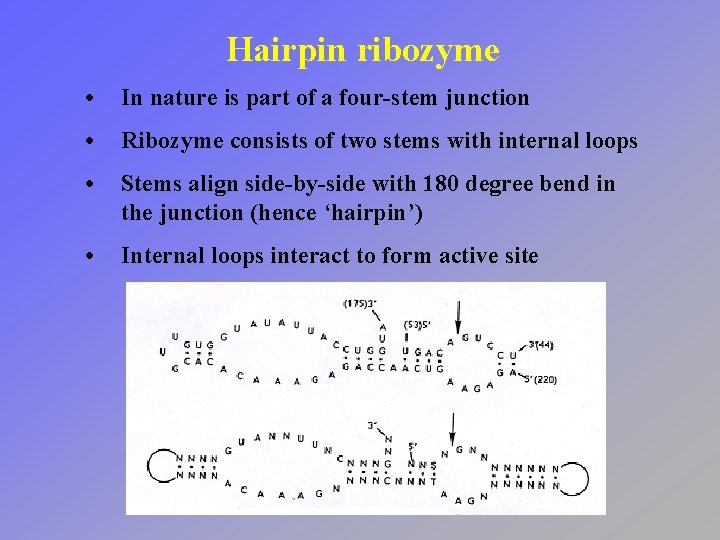
Hairpin ribozyme • In nature is part of a four-stem junction • Ribozyme consists of two stems with internal loops • Stems align side-by-side with 180 degree bend in the junction (hence ‘hairpin’) • Internal loops interact to form active site
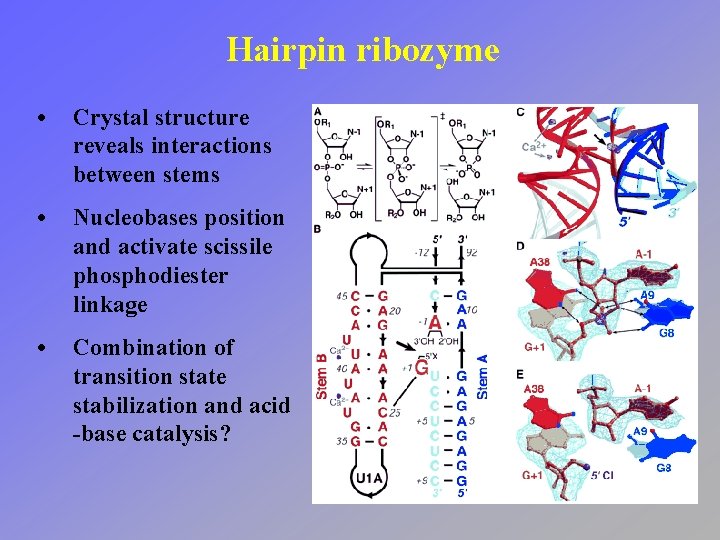
Hairpin ribozyme • Crystal structure reveals interactions between stems • Nucleobases position and activate scissile phosphodiester linkage • Combination of transition state stabilization and acid -base catalysis?
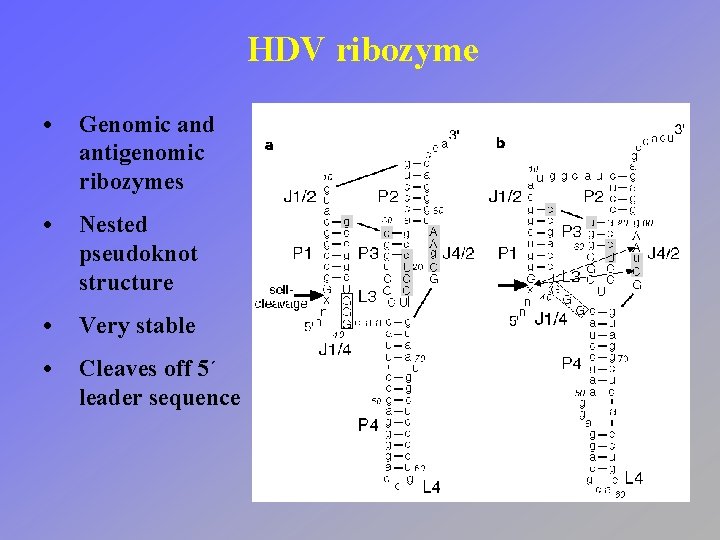
HDV ribozyme • Genomic and antigenomic ribozymes • Nested pseudoknot structure • Very stable • Cleaves off 5´ leader sequence
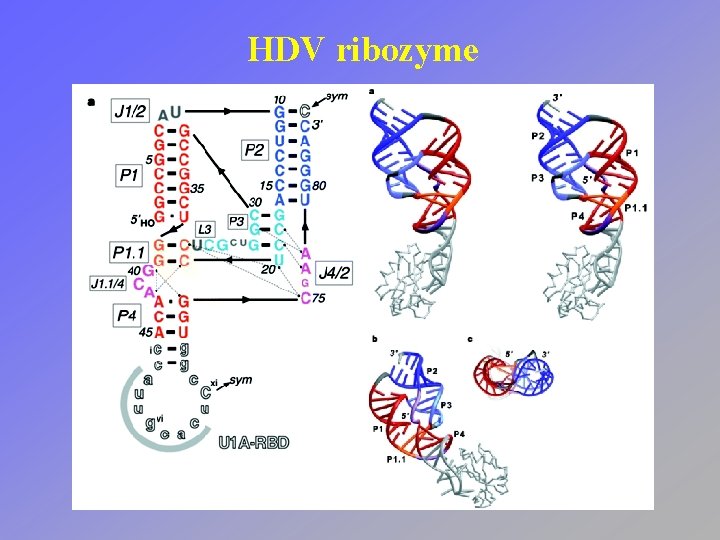
HDV ribozyme

HDV ribozyme • Active site positions an important cytidine near the scissile phosphodiester bond
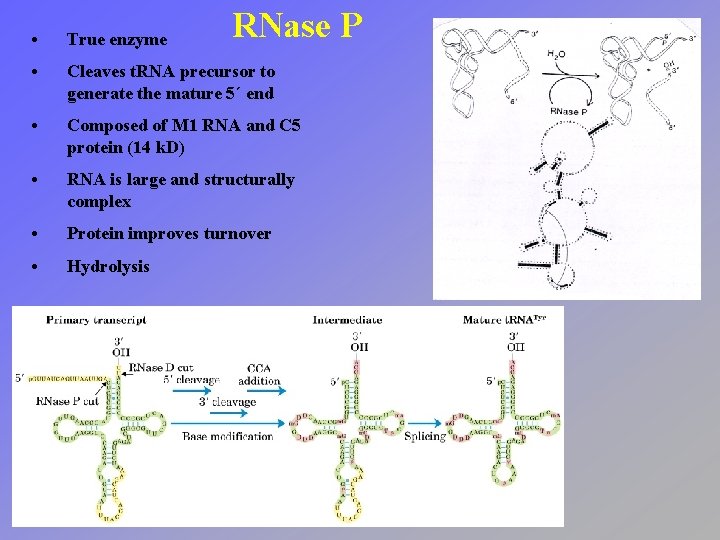
RNase P • True enzyme • Cleaves t. RNA precursor to generate the mature 5´ end • Composed of M 1 RNA and C 5 protein (14 k. D) • RNA is large and structurally complex • Protein improves turnover • Hydrolysis
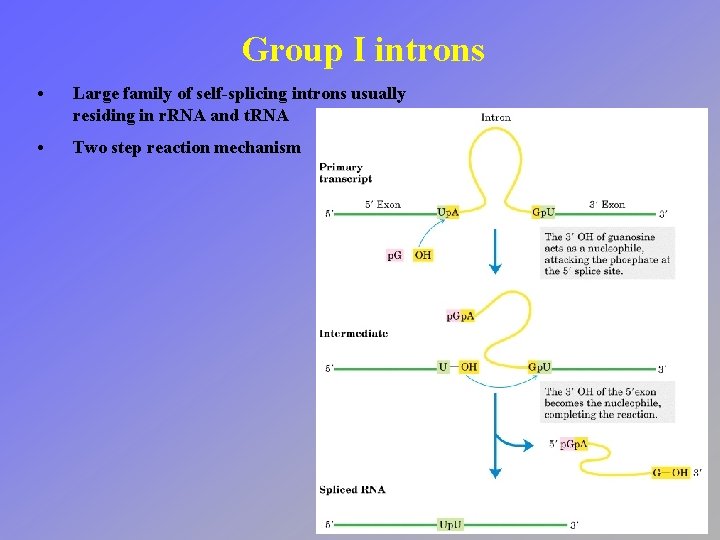
Group I introns • Large family of self-splicing introns usually residing in r. RNA and t. RNA • Two step reaction mechanism
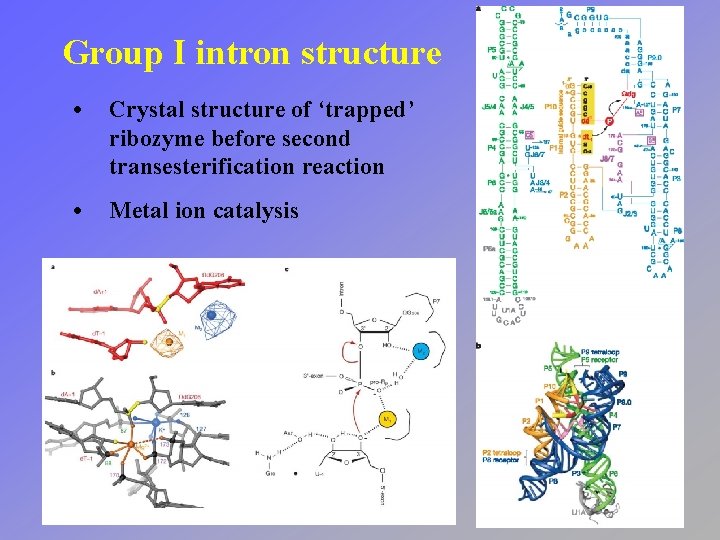
Group I intron structure • Crystal structure of ‘trapped’ ribozyme before second transesterification reaction • Metal ion catalysis
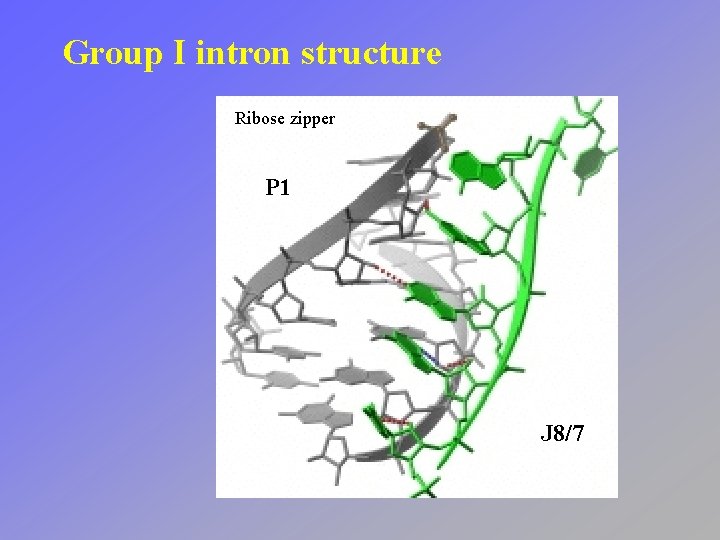
Group I intron structure Ribose zipper P 1 J 8/7
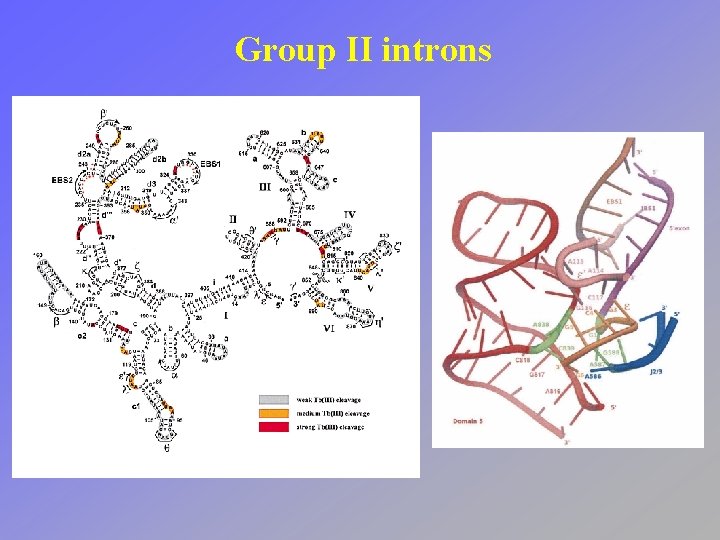
Group II introns
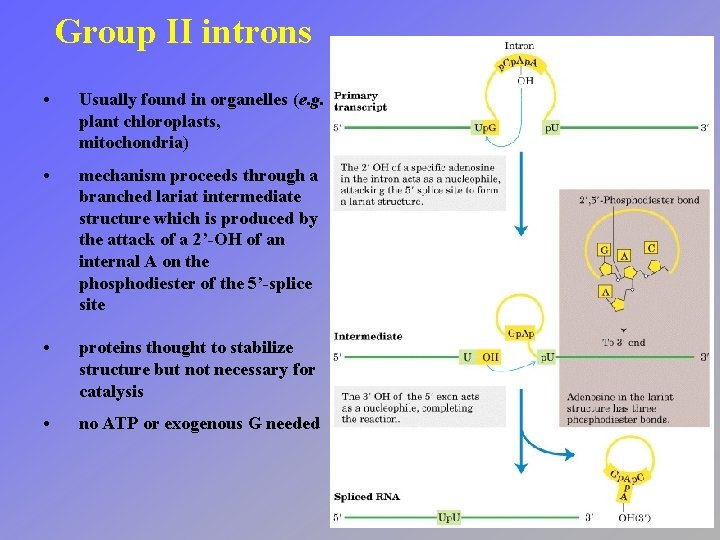
Group II introns • Usually found in organelles (e. g. plant chloroplasts, mitochondria) • mechanism proceeds through a branched lariat intermediate structure which is produced by the attack of a 2’-OH of an internal A on the phosphodiester of the 5’-splice site • proteins thought to stabilize structure but not necessary for catalysis • no ATP or exogenous G needed

Summary of splicing reactions
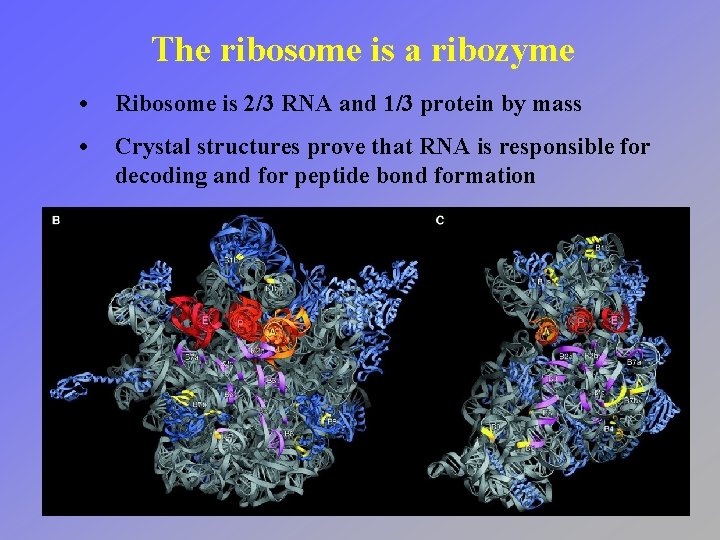
The ribosome is a ribozyme • Ribosome is 2/3 RNA and 1/3 protein by mass • Crystal structures prove that RNA is responsible for decoding and for peptide bond formation

Peptidyl transferase • Crystal structure of 50 S subunit shows no protein within 20 Å of peptidyl transferase center • Closest component to aa-t. RNA is adenosine 2451 in 23 S r. RNA • Proposed acid-base mechanism for peptide bond formation • Recent evidence shows substrate positioning accounts for catalysis
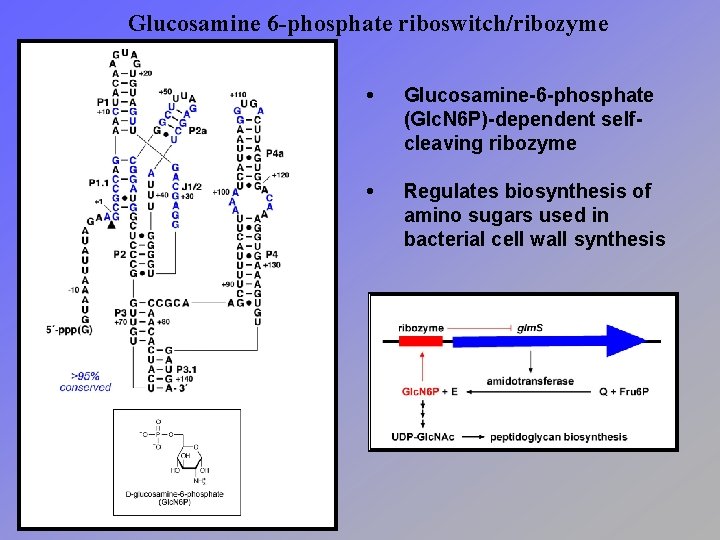
Glucosamine 6 -phosphate riboswitch/ribozyme • Glucosamine-6 -phosphate (Glc. N 6 P)-dependent selfcleaving ribozyme • Regulates biosynthesis of amino sugars used in bacterial cell wall synthesis
![glm. S is a metabolite-responsive ribozyme Effects of [glc. N 6 P] on the glm. S is a metabolite-responsive ribozyme Effects of [glc. N 6 P] on the](http://slidetodoc.com/presentation_image/42f2f8bfa959a53be8ddbb6205874e42/image-29.jpg)
glm. S is a metabolite-responsive ribozyme Effects of [glc. N 6 P] on the rate constant. M
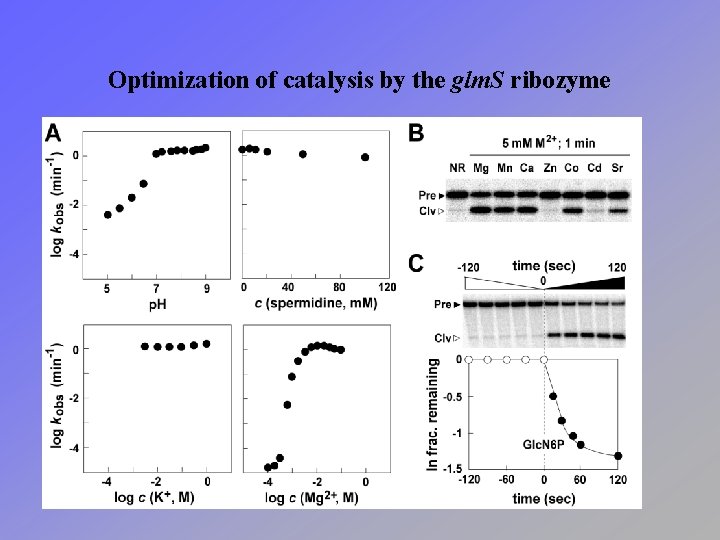
Optimization of catalysis by the glm. S ribozyme
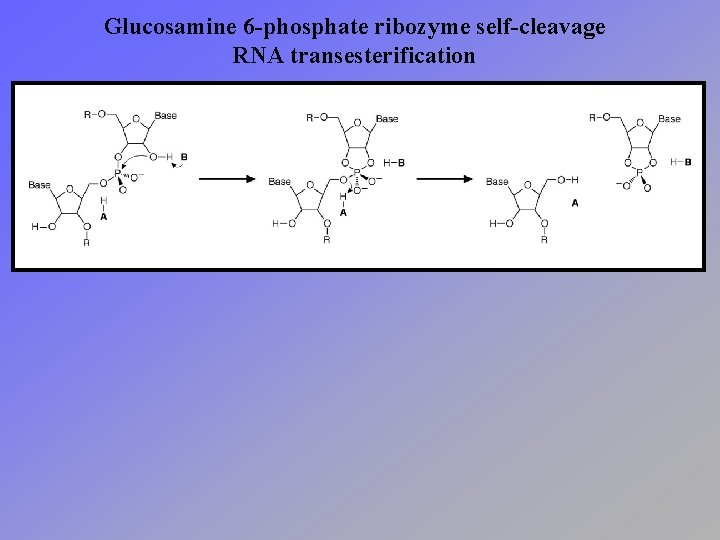
Glucosamine 6 -phosphate ribozyme self-cleavage RNA transesterification
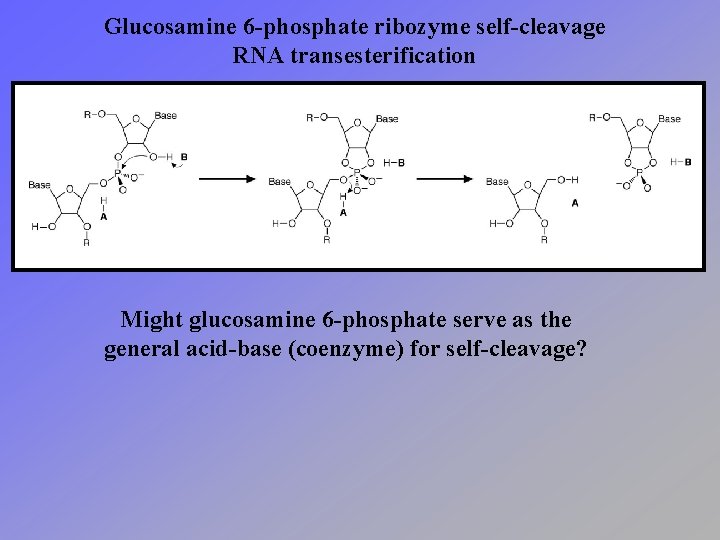
Glucosamine 6 -phosphate ribozyme self-cleavage RNA transesterification Might glucosamine 6 -phosphate serve as the general acid-base (coenzyme) for self-cleavage?
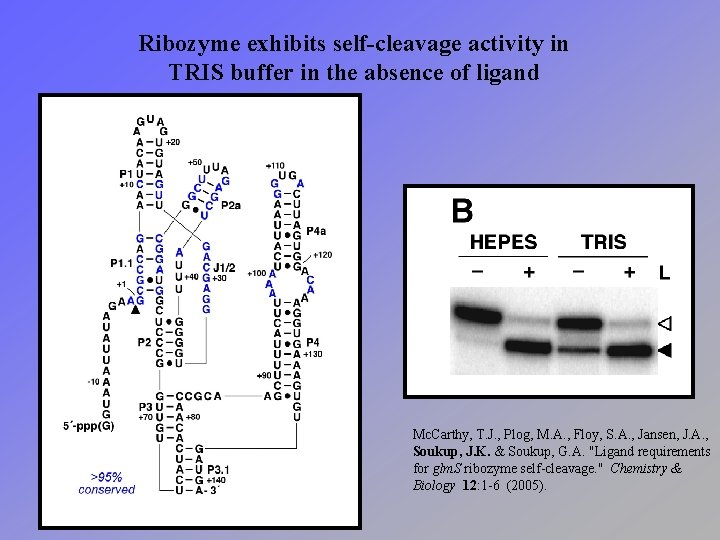
Ribozyme exhibits self-cleavage activity in TRIS buffer in the absence of ligand Mc. Carthy, T. J. , Plog, M. A. , Floy, S. A. , Jansen, J. A. , Soukup, J. K. & Soukup, G. A. "Ligand requirements for glm. S ribozyme self-cleavage. " Chemistry & Biology 12: 1 -6 (2005).
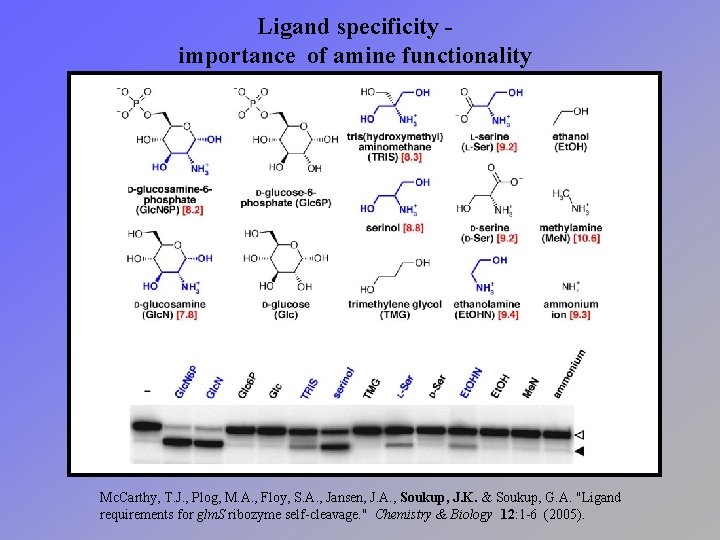
Ligand specificity importance of amine functionality Mc. Carthy, T. J. , Plog, M. A. , Floy, S. A. , Jansen, J. A. , Soukup, J. K. & Soukup, G. A. "Ligand requirements for glm. S ribozyme self-cleavage. " Chemistry & Biology 12: 1 -6 (2005).

Observed rate constants and apparent binding of ligand analogs Mc. Carthy, T. J. , Plog, M. A. , Floy, S. A. , Jansen, J. A. , Soukup, J. K. & Soukup, G. A. "Ligand requirements for glm. S ribozyme self-cleavage. " Chemistry & Biology 12: 1 -6 (2005).
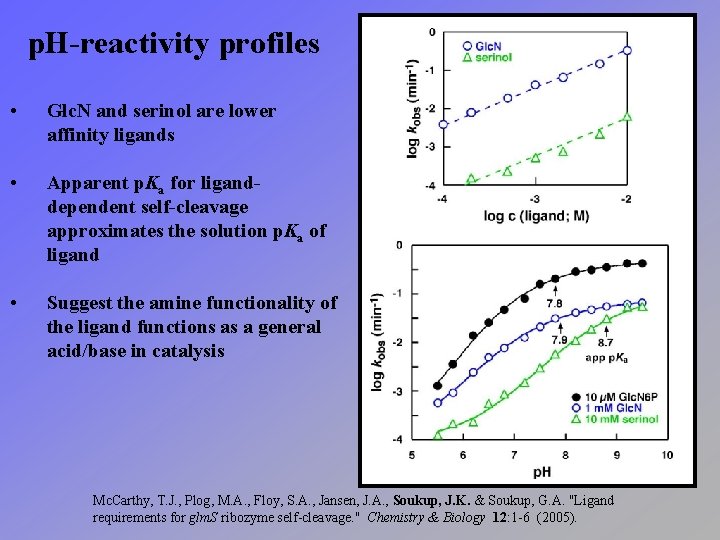
p. H-reactivity profiles • Glc. N and serinol are lower affinity ligands • Apparent p. Ka for liganddependent self-cleavage approximates the solution p. Ka of ligand • Suggest the amine functionality of the ligand functions as a general acid/base in catalysis Mc. Carthy, T. J. , Plog, M. A. , Floy, S. A. , Jansen, J. A. , Soukup, J. K. & Soukup, G. A. "Ligand requirements for glm. S ribozyme self-cleavage. " Chemistry & Biology 12: 1 -6 (2005).
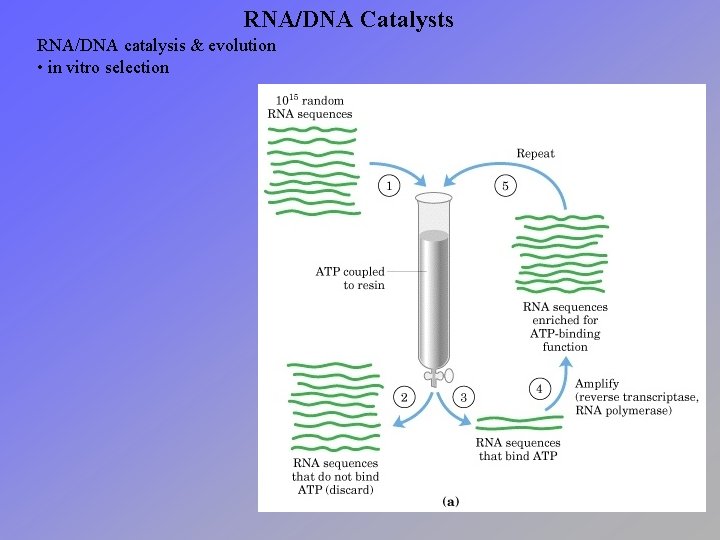
RNA/DNA Catalysts RNA/DNA catalysis & evolution • in vitro selection

RNA/DNA Catalysts RNA/DNA catalysis & evolution • increasing numbers of examples of reactions catalyzed by nucleic acids
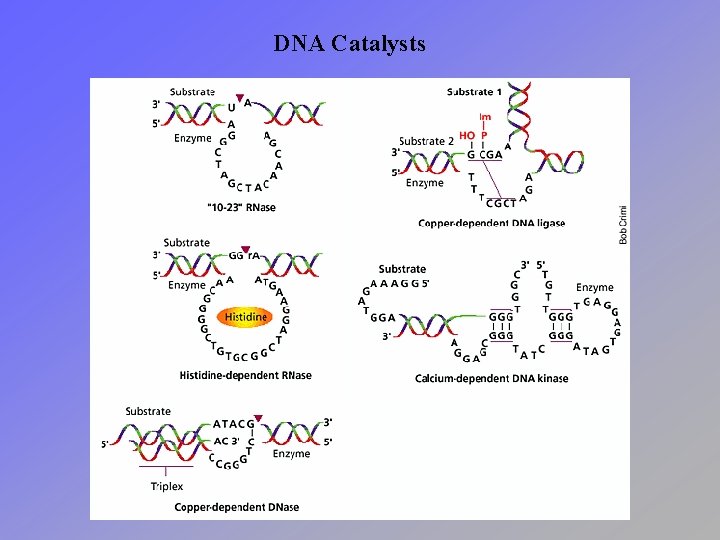
DNA Catalysts
- Slides: 39