RIP TRANSCRIPT EXPRESSION LEVELS OUTLINE RNA ImmunoPrecipitation RIP















- Slides: 20

RIP – TRANSCRIPT EXPRESSION LEVELS

OUTLINE RNA Immuno-Precipitation (RIP) NGS on RIP & its alternatives Alternate splicing Transcription as a graph Distribution of tags in exons Pipeline on RIP-seq dataset

RNA IMMUNO-PRECIPITATION (RIP) Global identification of multiple RNA targets of RNABinding Proteins (RBPs) Identify proteins associated with RNAs in RNP complexes Identify subsets of RNAs that are functionally-related and potentially co-regulated

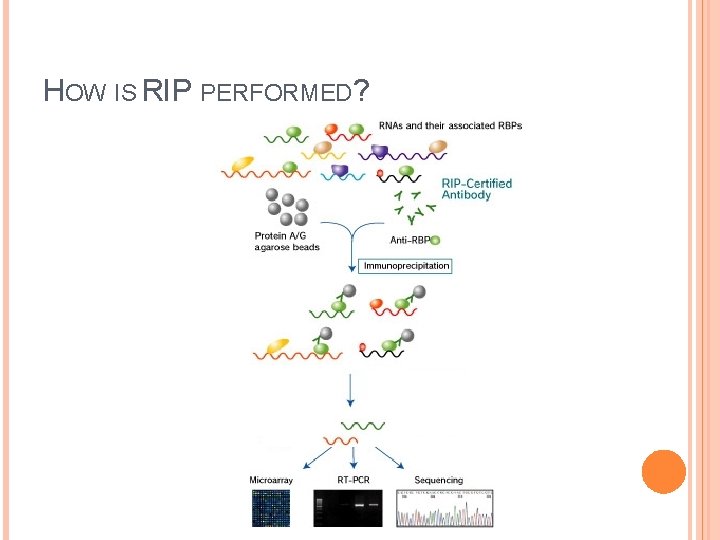
HOW IS RIP PERFORMED?

SEQUENCING ON RIP-Chip � Noisy � May miss out rare transcripts RIP-RT-PCR � PCR introduces mutations RIP tilting-arrays � Very expensive � Too sensitive to ‘transcriptional noise’

NGS ON RIP-Seq �A more complete and unbiased assessment of the global population of RNAs associated with a RNP complex � Minimize sequencing bias and high backgrounds known to the previously-mentioned methods

ALTERNATE SPLICING • A simple example Regions with the numbers of reads – Exon 1: chr 1: 13113087 -13113138(5, 1); – Exon 2: chr 1: 13113270 -13113299(2, 0); – Exon 3: chr 1: 13113312 -13113343(3, 0); • Splice reads – chr 1, 13113107, 13113138, chr 1, 13113312, 13113343, 3. 0; – chr 1, 13113087, 13113116, chr 1, 13113270, 13113299, 2. 0; Exon 1(5) Exon 2(2) Exon 3(3) Exon_Num(Tags)

ALTERNATE SPLICING • A less ideal example Regions with the numbers of reads – Exon 1: chr 4: 145149018 -145149181(29, 0); – Exon 2: chr 4: 145149265 -145149402(8, 0); – Exon 3: chr 4: 146893298 -146895275(116, 1); • Splice reads – chr 4, 145149059, 145149088, chr 4, 146894246, 146894276, 3. 0; – chr 4, 145149374, 145149402, chr 4, 146894470, 146894498, 2. 0; Exon 1(29) Exon 2(8) Exon 3(116)


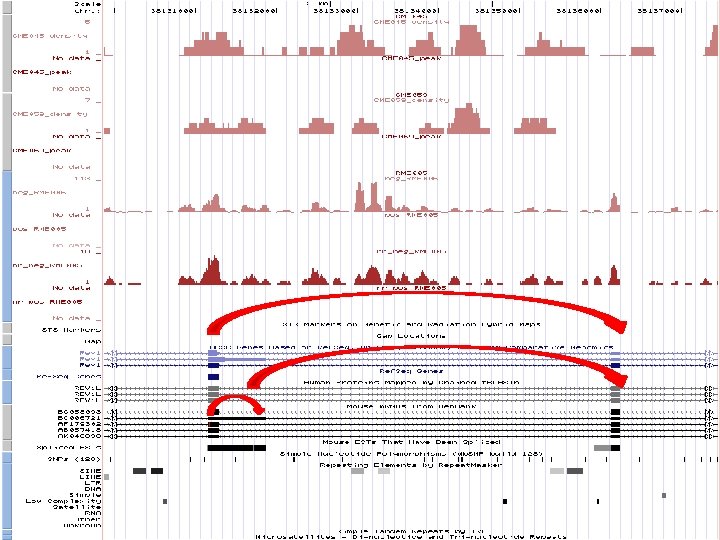
TRANSCRIPTION AS A GRAPH From RNA-seq data, check the overlap of the tags If a region has more than one tag, we call it an enriched region � Nodes Using the splice reads, we will connect the enriched regions � Edges

TRANSCRIPTION AS A GRAPH Represent transcriptome in a topologically sorted acyclic graph Some Observed Errors (RME 005) � Out-of-range edges in graphs � Self-looping nodes Default action: Ignore them

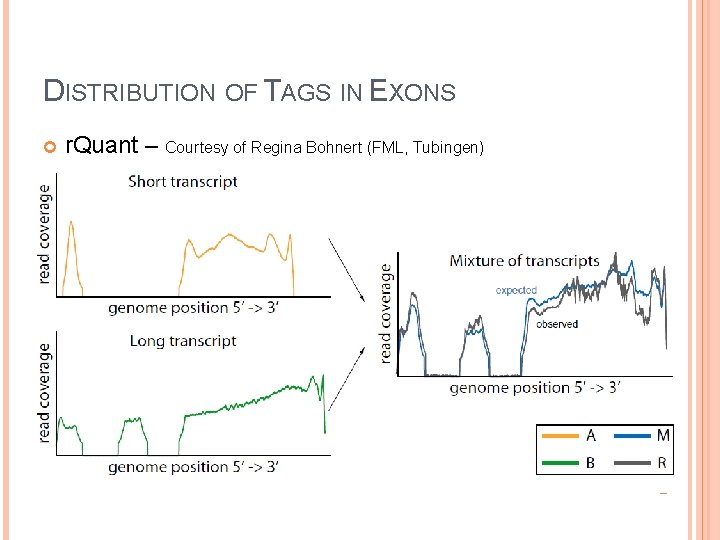
DISTRIBUTION OF TAGS IN EXONS r. Quant – Courtesy of Regina Bohnert (FML, Tubingen)

RNA-SEQ RIP-SEQ The previous results are from RNA-seq � Will � And we have similar observations on RIP-seq datasets? possibly link the observations to transcription expression levels in transcriptome

PIPELINE ON RIP-SEQ DATASET 1. 2. 3. 4. 5. Dataset RME 005 is used Use Top. Hat / Eland to map RNA back to genome Generate transcription-graphs for each transcript with alternate splicing Express the paths of all transcriptions in the graph using a set of linear equations Use R to solve the linear equations

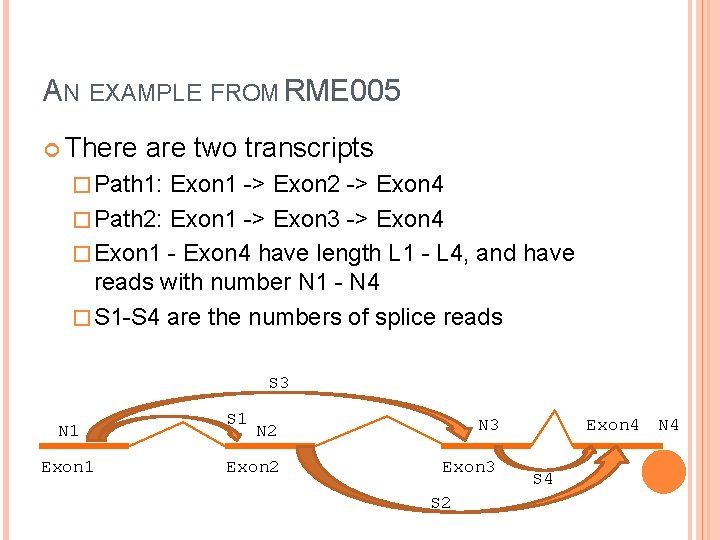
AN EXAMPLE FROM RME 005 There are two transcripts � Path 1: Exon 1 -> Exon 2 -> Exon 4 � Path 2: Exon 1 -> Exon 3 -> Exon 4 � Exon 1 - Exon 4 have length L 1 - L 4, and have reads with number N 1 - N 4 � S 1 -S 4 are the numbers of splice reads S 3 N 1 Exon 1 S 1 N 2 N 3 Exon 2 Exon 3 S 2 Exon 4 N 4 S 4

ASSUMPTIONS The transcript expression levels are: � Path 1: x 1 � Path 2: x 2 The read length = constant The reads are uniformly sampled from the transcripts Use density of reads instead of read_coverage � Differentiate reads on both long & short exons

EQUATIONS FOR LINEAR PROGRAMMING Objective function: minimize the sum of d_i Constraints � � � � � N 1/L 1 = x 1 + x 2 + d 1 - d 2 S 1/R = x 1 + d 3 - d 4 N 2/L 2 = x 1 + d 5 - d 6 S 2/R = x 1 + d 7 - d 8 N 1 S 3/R = x 2 + d 9 - d 10 N 3/L 3 = x 2 + d 11 - d 12 S 4/R = x 2 + d 13 - d 14 N 4/L 4 = x 1 + x 2 + d 15 - d 16 x 1 , x 2 >= 0 d_i >= 0 S 3 S 1 N 2 The solution should be the values of x 1, x 2 and all d_i N 3 N 4 S 2

ANOTHER PROBLEM An implicit assumption on enriched regions in RME 005 � RIP is known to be ~10% efficient � Noise will overwhelm true RNP-targets � Should use total-RNA as control dataset True-positive regions from RIP should be relatively enriched with tags than

HANDLING THE ASSUMPTION Obtain RNA-seq from the same source of transcriptome � Directly compare both RNA-seq and RIP-seq data � RIP-chip discriminate enriched region with >4 -fold than RNA-chip data Maybe 4 -fold is the magic number ? Current tag distribution observed by Dr Li Guoliang � Non-uniform on RNA-seq as opposed to what r. Quant has observed

Q&A