Rhythmic growth explained by coincidence between internal and

Rhythmic growth explained by coincidence between internal and external cues; what gene networks are underlying? Kazunari (Kazu) Nozue College of Biological Sciences, University of California Davis May 16, 2007

Acknowledgements Stacey Hamer Maloof lab Julin Maloof Andreah Wallace (Andii) Mike Covington
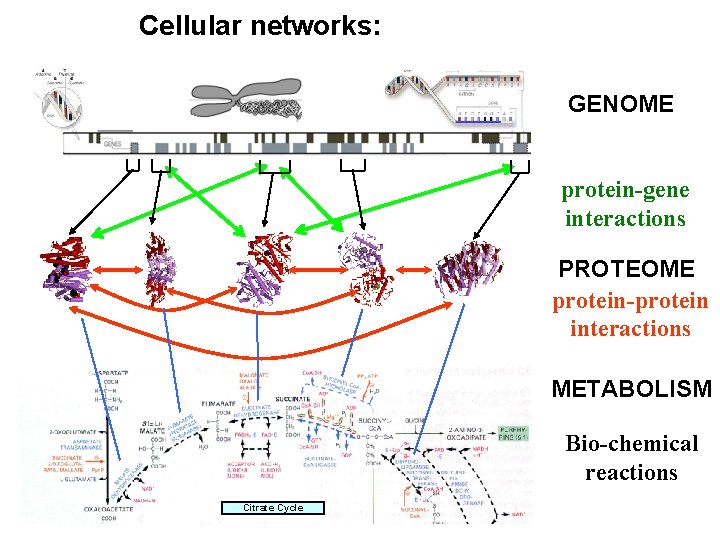
Cellular networks: GENOME protein-gene interactions PROTEOME protein-protein interactions METABOLISM Bio-chemical reactions Citrate Cycle
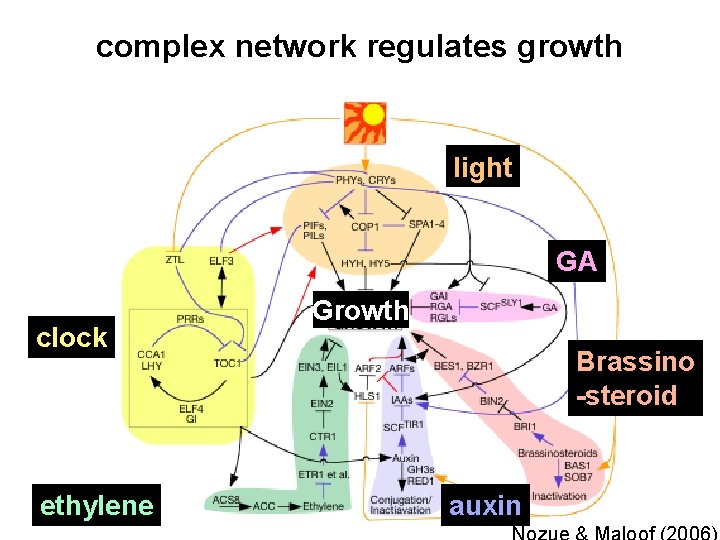
complex network regulates growth light GA clock ethylene Growth Brassino -steroid auxin


Molecular mechanisms of circadian clock negative feedback loop Salome (2005)

Hypocotyl elongation has circadian rhythm Time in continuous light (hrs) Dowson-Day (1999) Plant J. 17: 63 -71 2 d 12 L/12 D entrainment image recording under continuous dim light
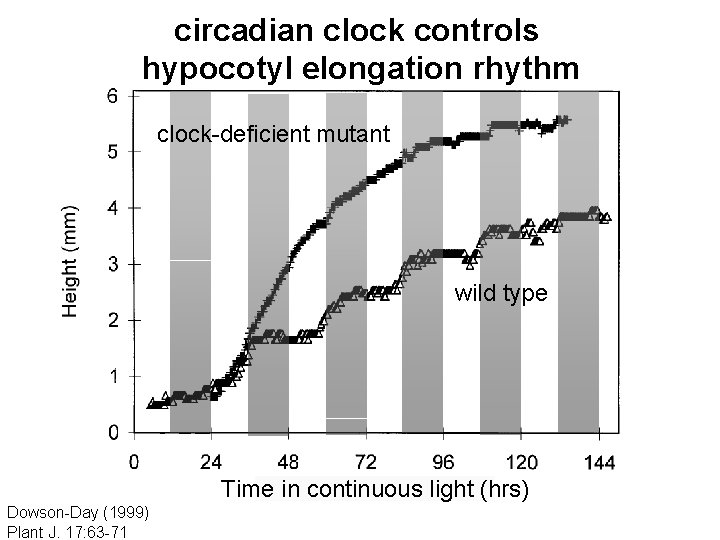
circadian clock controls hypocotyl elongation rhythm clock-deficient mutant wild type Time in continuous light (hrs) Dowson-Day (1999) Plant J. 17: 63 -71

Real world is not continuous light …day-night cycles!

time-lapse photography (Col; short day (SD))

what molecular mechanisms are underlying? • Hypothesis: transcriptional regulation is involved in gating of dark-induced elongation. • Method: whole genome microarray analysis • Purpose: to find genes which expression patterns are correlated with growth pattern

what is microarray?

TOP 10 of up-regulated genes in growing phase (Dark) vs non-growing phase (Dark) rank-product non-parmetric method(Breitling 2004, FEBS 573: 83) pfp; percentage of false-positives, FC; fold-change Nozue (2007) Nature
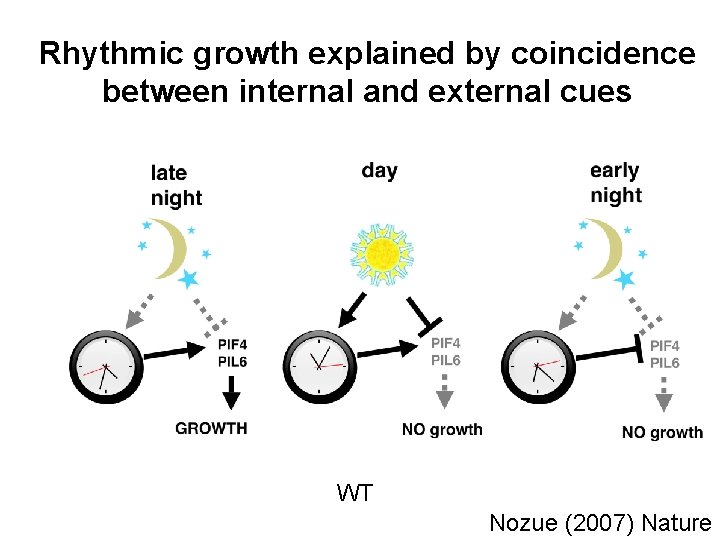
Rhythmic growth explained by coincidence between internal and external cues WT Nozue (2007) Nature

coincidence between internal & external cues another example: flowering time Imaizumi (2006)

questions • what are genes in my lists? • Do the genes control growth? If so, • how they control growth?

what gene networks control plant growth? • • genotype interaction (cf. Jose’s talk) protein-protein interaction • co-expression
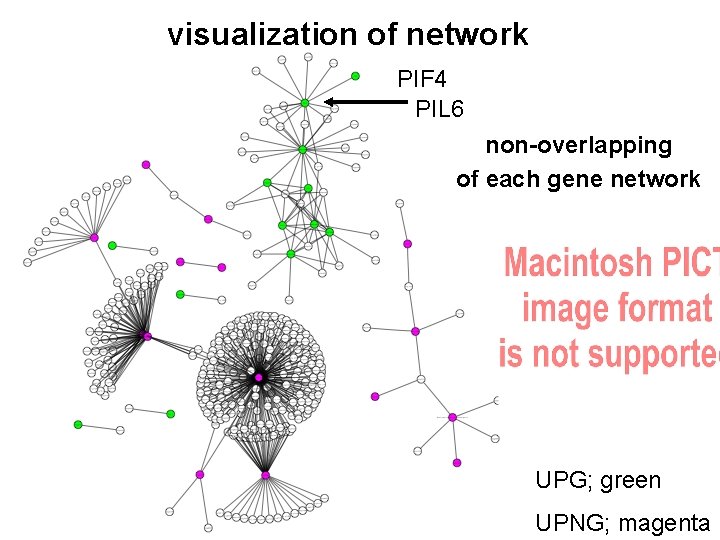
visualization of network PIF 4 PIL 6 non-overlapping of each gene network UPG; green UPNG; magenta
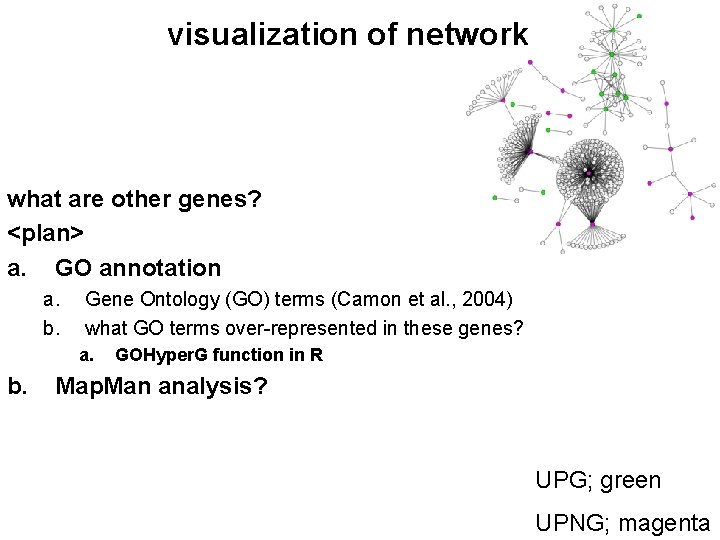
visualization of network what are other genes? <plan> a. GO annotation a. b. Gene Ontology (GO) terms (Camon et al. , 2004) what GO terms over-represented in these genes? a. b. GOHyper. G function in R Map. Man analysis? UPG; green UPNG; magenta
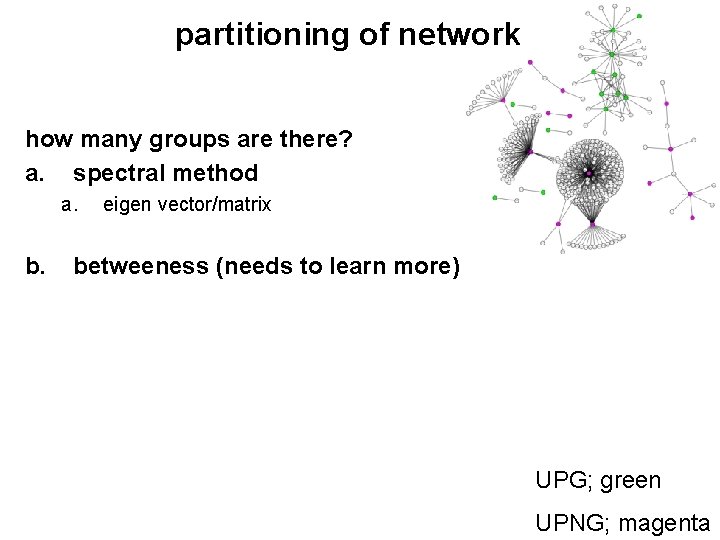
partitioning of network how many groups are there? a. spectral method a. b. eigen vector/matrix betweeness (needs to learn more) UPG; green UPNG; magenta
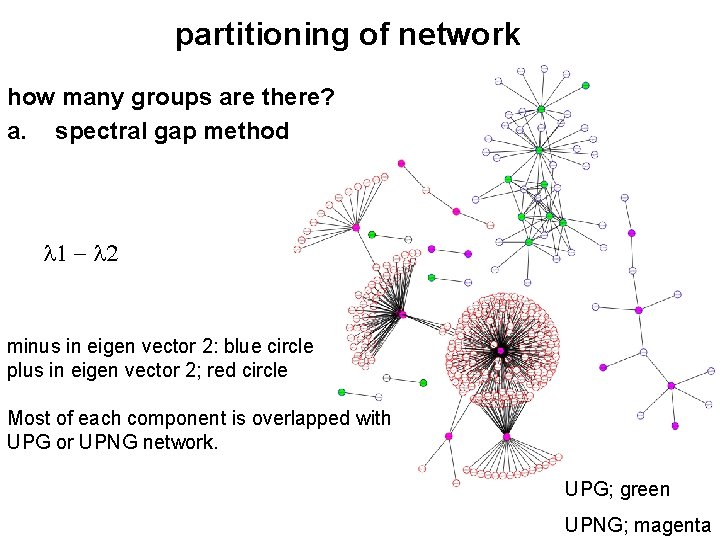
partitioning of network how many groups are there? a. spectral gap method minus in eigen vector 2: blue circle plus in eigen vector 2; red circle Most of each component is overlapped with UPG or UPNG network. UPG; green UPNG; magenta
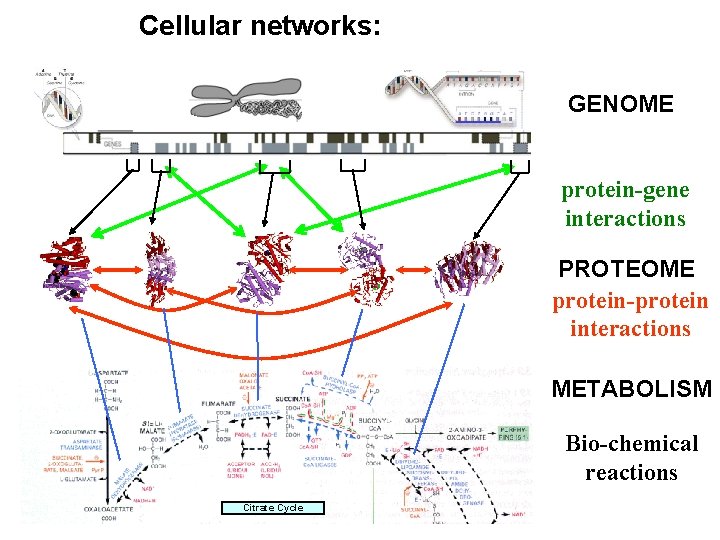
Cellular networks: GENOME protein-gene interactions PROTEOME protein-protein interactions METABOLISM Bio-chemical reactions Citrate Cycle

biological network layered
- Slides: 23