Restriction Enzyme Digest October 8 2015 Restriction Enzymes

Restriction Enzyme Digest October 8, 2015

Restriction Enzymes

Restriction Enzymes • • • Also known as restriction endonucleases Scan the DNA sequence Find a very specific set of nucleotides Make a specific cut Used to construct recombinant DNA plasmids
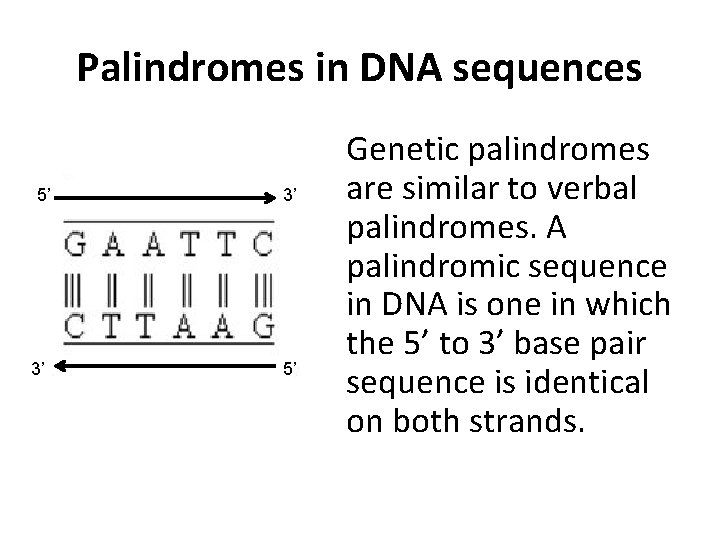
Palindromes in DNA sequences 5’ 3’ 3’ 5’ Genetic palindromes are similar to verbal palindromes. A palindromic sequence in DNA is one in which the 5’ to 3’ base pair sequence is identical on both strands.

Restriction enzymes recognize and make a cut within specific palindromic sequences, known as restriction sites, in the DNA. This is usually a 4 - or 6 base pair sequence.

Restriction Endonuclease Types Type I- multi-subunit, both endonuclease and methylase activities, cleave at random up to 1000 bp from recognition sequence Type II- most single subunit, cleave DNA within recognition sequence Type III- multi-subunit, endonuclease and methylase about 25 bp from recognition sequence

Hae III Hae. III is a restriction enzyme that searches the DNA molecule until it finds this sequence of four nitrogen bases. 5’ TGACGGGTTCGAGGCCAG 3’ 3’ ACTGCCCAAGGTCCGGTC 5’

Once the recognition site is found Hae III will cleave the DNA at that site 5’ TGACGGGTTCGAGGCCAG 3’ 3’ ACTGCCCAAGGTCCGGTC 5’

These cuts produce “blunt ends” 5’ TGACGGGTTCGAGG 3’ ACTGCCCAAGGTCC CCAG 3’ GGTC 5’

The names for restriction enzymes come from: • the type of bacteria in which the enzyme is found • the order in which the restriction enzyme was identified and isolated. Eco. RI for example R strain of E. coli bacteria I as it is was the first E. coli restriction enzyme to be discovered.
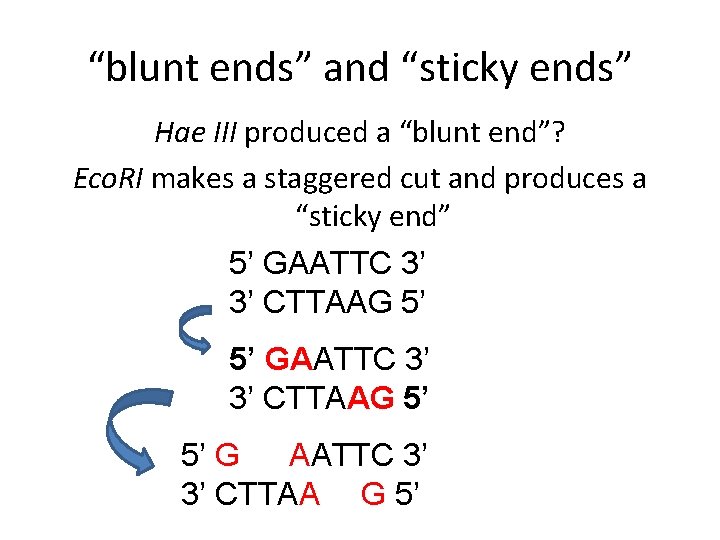
“blunt ends” and “sticky ends” Hae III produced a “blunt end”? Eco. RI makes a staggered cut and produces a “sticky end” 5’ GAATTC 3’ 3’ CTTAAG 5’ 5’ G AATTC 3’ 3’ CTTAA G 5’

More examples of restriction sites of restriction enzymes with their cut sites Hind III: 5’ AAGCTT 3’ 3’ TTCGAA 5’ Bam HI: 5’ GGATCC 3’ 3’ CCTAGG 5’ Alu I: 5’ AGCT 3’ 3’ TCGA 5’
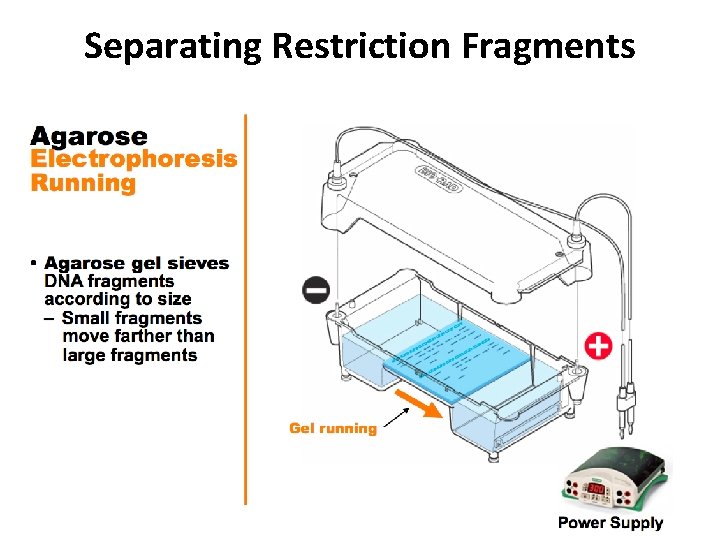
Separating Restriction Fragments
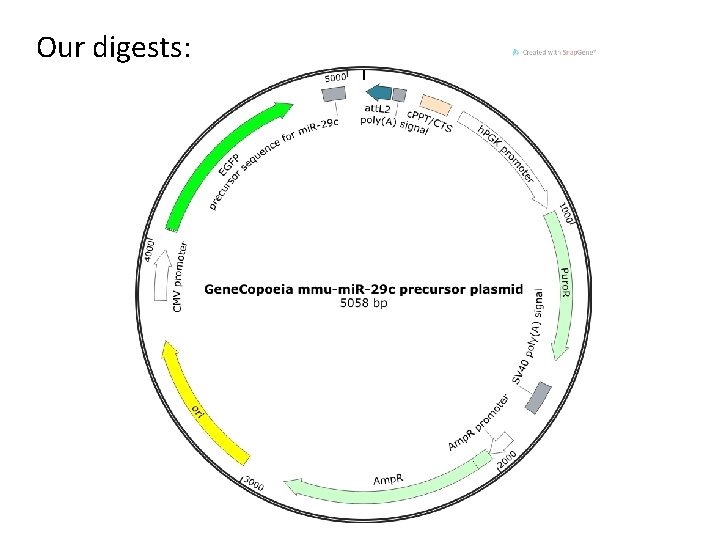
Our digests:
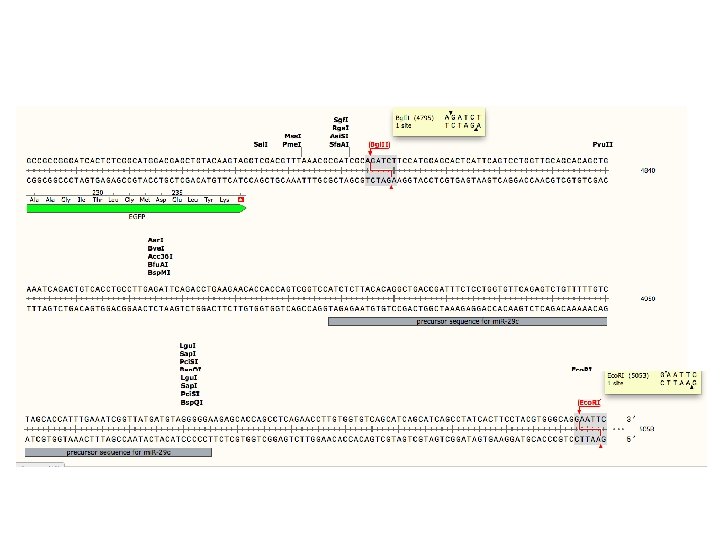
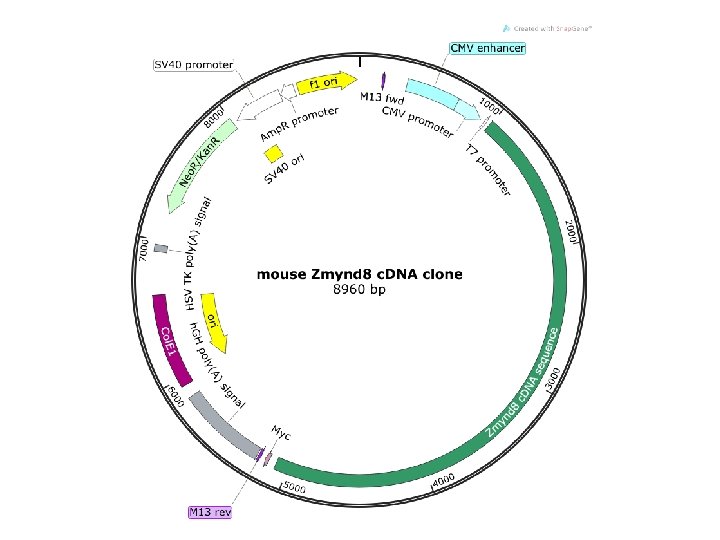
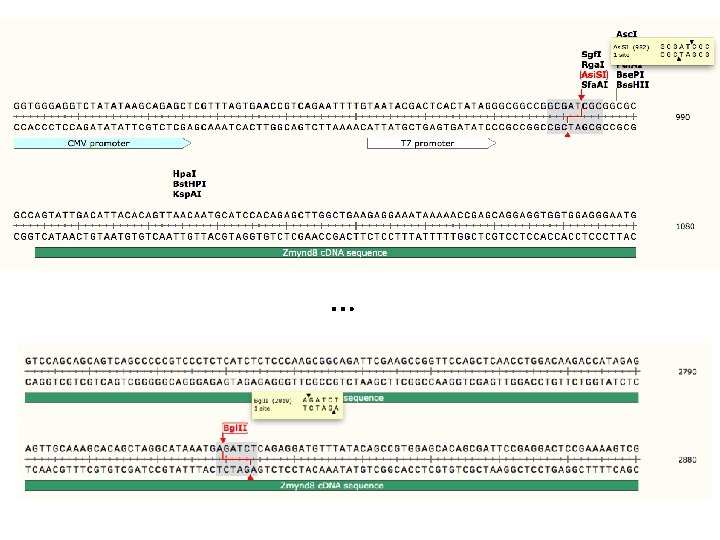
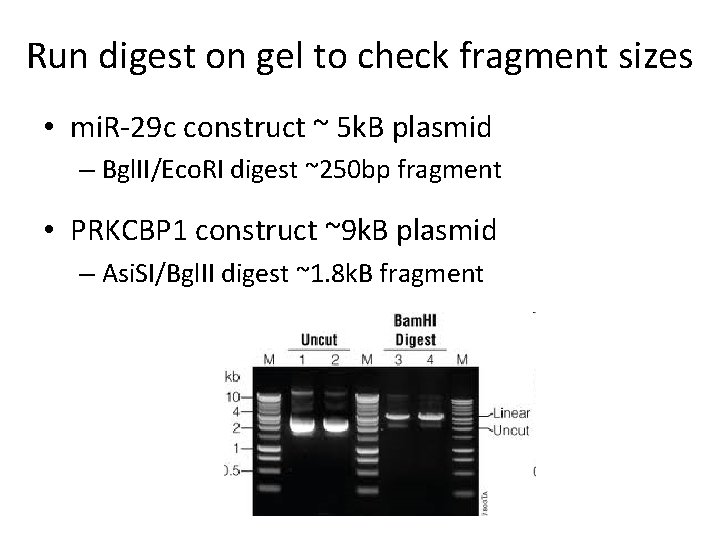
Run digest on gel to check fragment sizes • mi. R-29 c construct ~ 5 k. B plasmid – Bgl. II/Eco. RI digest ~250 bp fragment • PRKCBP 1 construct ~9 k. B plasmid – Asi. SI/Bgl. II digest ~1. 8 k. B fragment
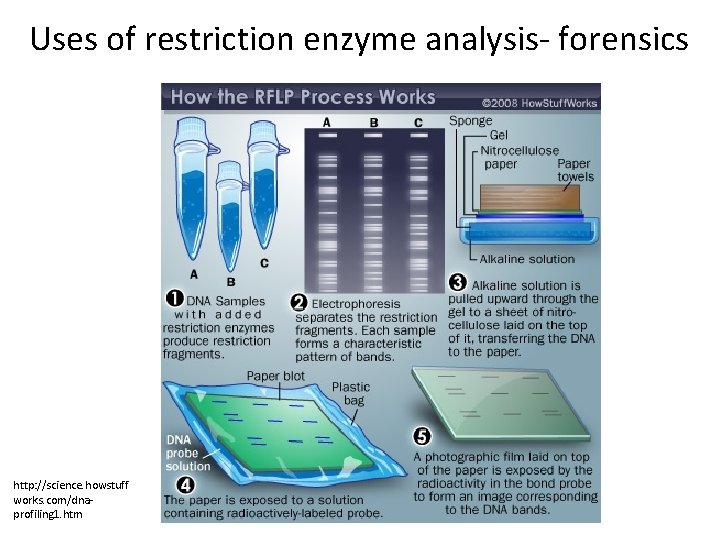
Uses of restriction enzyme analysis- forensics http: //science. howstuff works. com/dnaprofiling 1. htm
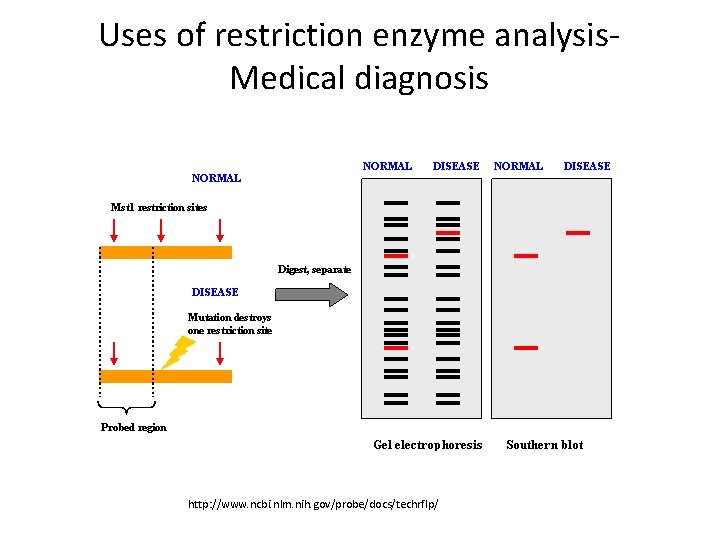
Uses of restriction enzyme analysis. Medical diagnosis http: //www. ncbi. nlm. nih. gov/probe/docs/techrflp/
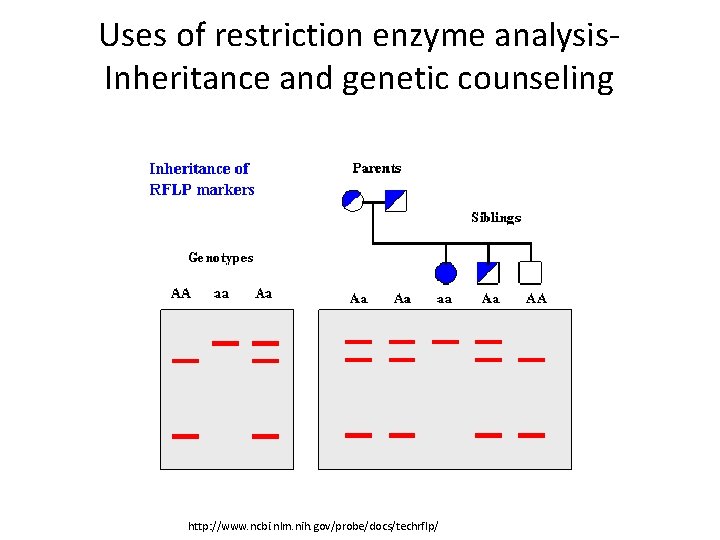
Uses of restriction enzyme analysis. Inheritance and genetic counseling http: //www. ncbi. nlm. nih. gov/probe/docs/techrflp/
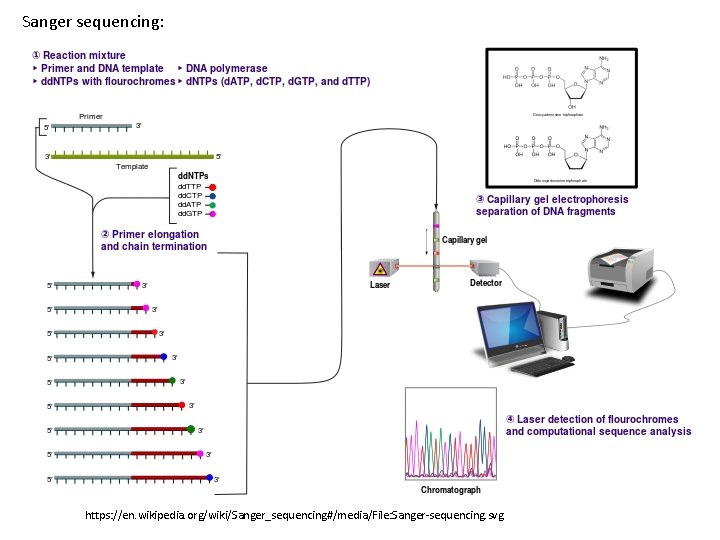
Sanger sequencing: https: //en. wikipedia. org/wiki/Sanger_sequencing#/media/File: Sanger-sequencing. svg
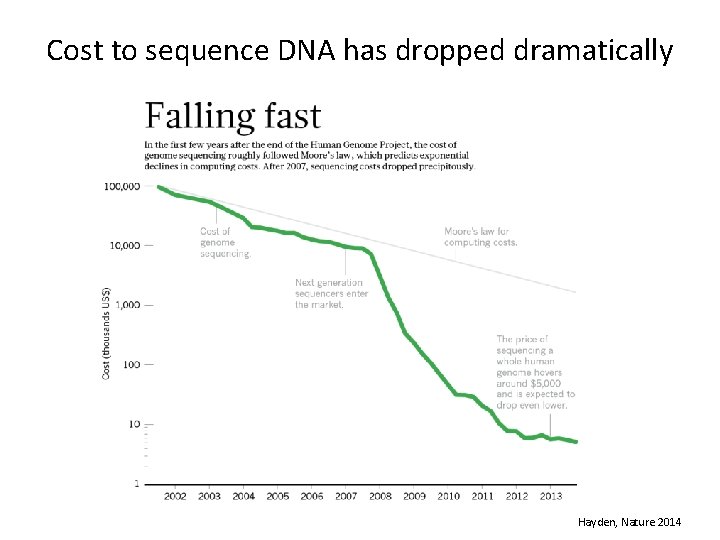
Cost to sequence DNA has dropped dramatically Hayden, Nature 2014
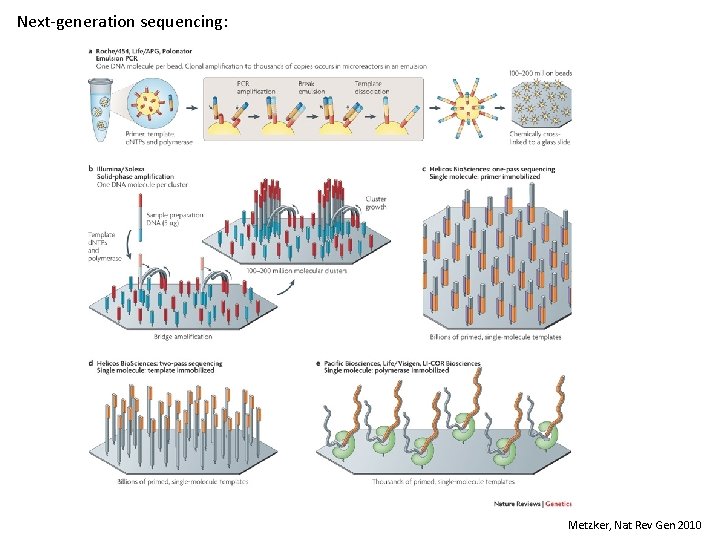
Next-generation sequencing: Metzker, Nat Rev Gen 2010
- Slides: 24