Research Title Researchers names and affiliations Glass Bead
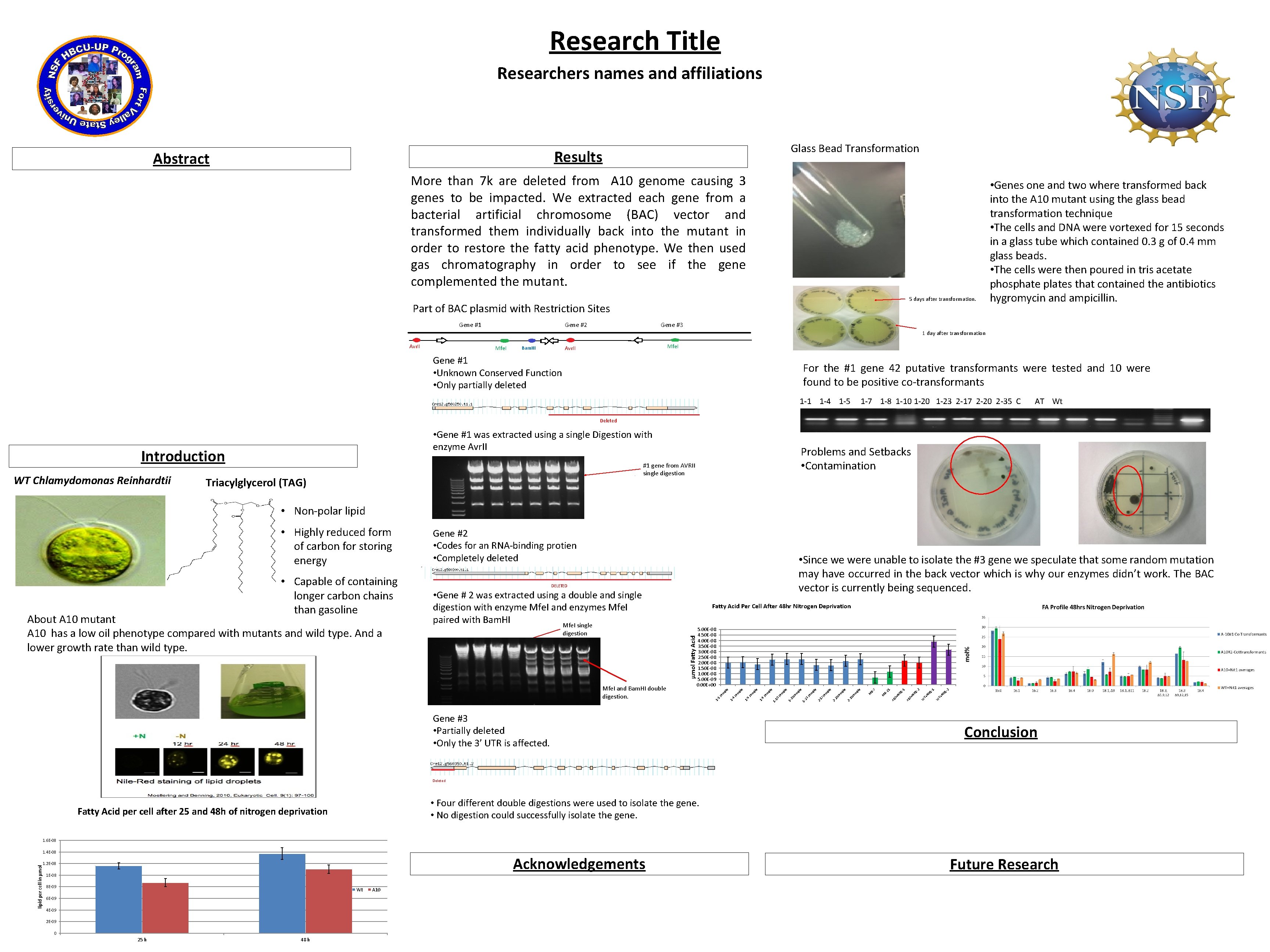
Research Title Researchers names and affiliations Glass Bead Transformation Results Abstract More than 7 k are deleted from A 10 genome causing 3 genes to be impacted. We extracted each gene from a bacterial artificial chromosome (BAC) vector and transformed them individually back into the mutant in order to restore the fatty acid phenotype. We then used gas chromatography in order to see if the gene complemented the mutant. 5 days after transformation. Part of BAC plasmid with Restriction Sites Gene #1 Gene #2 • Genes one and two where transformed back into the A 10 mutant using the glass bead transformation technique • The cells and DNA were vortexed for 15 seconds in a glass tube which contained 0. 3 g of 0. 4 mm glass beads. • The cells were then poured in tris acetate phosphate plates that contained the antibiotics hygromycin and ampicillin. Gene #3 1 day after transformation Avr. II Mfe. I Avr. II Bam. HI Gene #1 • Unknown Conserved Function • Only partially deleted For the #1 gene 42 putative transformants were tested and 10 were found to be positive co-transformants 1 -1 1 -4 1 -5 1 -7 1 -8 1 -10 1 -23 2 -17 2 -20 2 -35 C AT Wt Deleted • Gene #1 was extracted using a single Digestion with enzyme Avr. II Introduction WT Chlamydomonas Reinhardtii Problems and Setbacks • Contamination #1 gene from AVRII single digestion Triacylglycerol (TAG) • Non-polar lipid DELETED • Gene # 2 was extracted using a double and single digestion with enzyme Mfe. I and enzymes Mfe. I paired with Bam. HI Mfe. I single Gene #3 • Partially deleted • Only the 3’ UTR is affected. -2 -1 T+ Ni t 1 W T+ Ni t 1 -2 A 1 0+ Ni t 1 Ni 0+ A 1 W -1 21 BM 7 B- pl sa m 30 M e e 2 - sa m 2 - 20 pl sa m 17 pl e e 2 - 23 sa m pl e 1 - 1 - 20 sa m pl pl sa m 1 - 10 m pl sa 8 e e e 1 - 5 sa m pl e 1 - 4 sa m pl e 1 - m pl Mfe. I and Bam. HI double digestion. 5. 00 E-08 4. 50 E-08 4. 00 E-08 3. 50 E-08 3. 00 E-08 2. 50 E-08 2. 00 E-08 1. 50 E-08 1. 00 E-08 5. 00 E-09 0. 00 E+00 sa digestion Fatty Acid Per Cell After 48 hr Nitrogen Deprivation 1 About A 10 mutant A 10 has a low oil phenotype compared with mutants and wild type. And a lower growth rate than wild type. • Since we were unable to isolate the #3 gene we speculate that some random mutation may have occurred in the back vector which is why our enzymes didn’t work. The BAC vector is currently being sequenced. 1 - • Capable of containing longer carbon chains than gasoline Gene #2 • Codes for an RNA-binding protien • Completely deleted µmol Fatty Acid • Highly reduced form of carbon for storing energy Conclusion Deleted • Four different double digestions were used to isolate the gene. • No digestion could successfully isolate the gene. Fatty Acid per cell after 25 and 48 h of nitrogen deprivation 1. 6 E-08 lipid per cell in µmol 1. 4 E-08 Acknowledgements 1. 2 E-08 1 E-08 8 E-09 Wt 6 E-09 4 E-09 2 E-09 0 25 h 48 h A 10 Future Research
- Slides: 1