Reporting Karyotype FISH Molecular Results Janet BrunnerGrady PAC

Reporting Karyotype, FISH & Molecular Results Janet Brunner-Grady, PA-C February 2019 TRAINING & DEVELOPMENT | .

Hematologic Malignancy Evaluation • Histology/ Morphology (Least sensitive) – What the cells look like • Immunohistochemistry (IHC) – Staining the cells to identify specific markers • Flow cytometry – Looks at individual cells based on staining for specific markers • Cytogenetics – Karyotype - FISH analysis • Molecular studies (Most sensitive) – Identifying abnormal genes and/or gene products TRAINING & DEVELOPMENT | 2.

Types of Cytogenetic Analysis • Karyotype vs. FISH Ø Karyotype- is the characterization of structural and numerical changes in chromosomes while cells are dividing. They are displayed as a systematized arrangement of chromosome pairs in descending order of size. Ø FISH (Fluorescent in situ hybridization) • A molecular cytogenetic technique using fluorescent probes that bind to a specific part of a chromosome • Used to detect the presence or absence of specific DNA sequences on chromosomes TRAINING & DEVELOPMENT | 3.
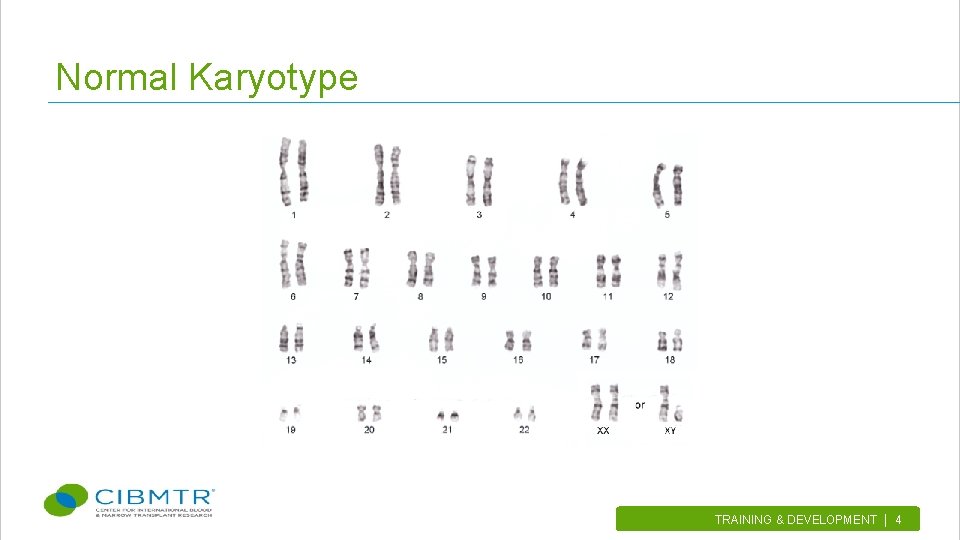
Normal Karyotype TRAINING & DEVELOPMENT | 4.
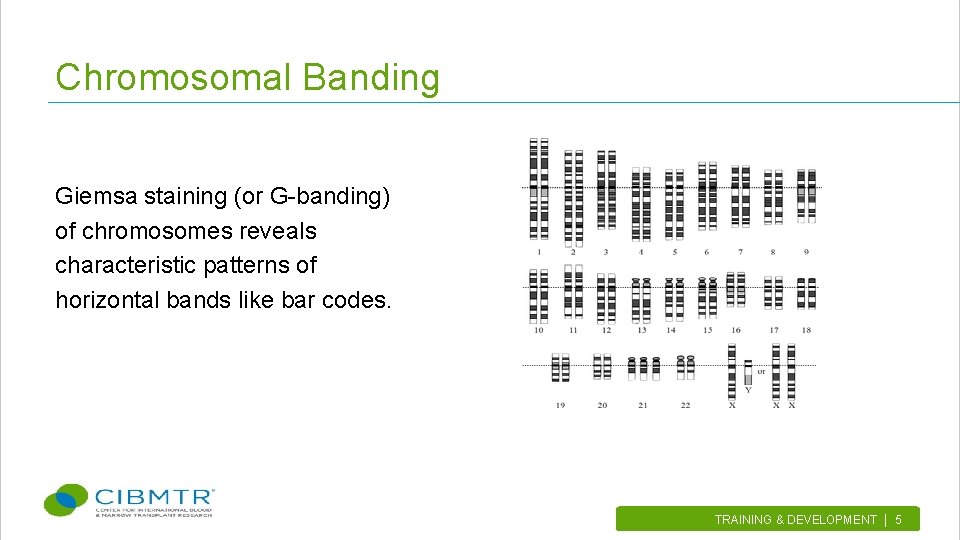
Chromosomal Banding Giemsa staining (or G-banding) of chromosomes reveals characteristic patterns of horizontal bands like bar codes. TRAINING & DEVELOPMENT | 5.

Chromosome Definitions TRAINING & DEVELOPMENT | 6.
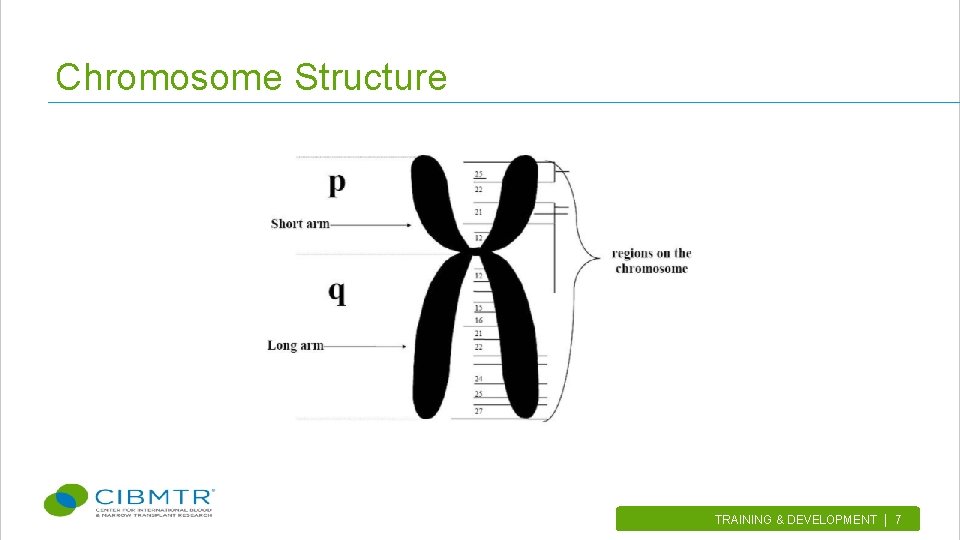
Chromosome Structure TRAINING & DEVELOPMENT | 7.

Definitions of a Clone • Two or more cells with gain of a specific chromosome (e. g. , trisomy 8 or +8) • Two or more cells with the same structural chromosomal abnormality (e. g. , t(9; 22)) • Three or more cells with loss of a specific chromosome (e. g. , monosomy 7 or -7) TRAINING & DEVELOPMENT | 8.

Types of FISH probes • Centromere enumeration probes (CEP) – Monitors number of a specific chromosome in a cell (e. g. , trisomy 8 or +8) • Locus specific probes – Used to rule out deletions, gains or rearrangements of specific loci (e. g. , del(7 q) or del(7)(q 22 q 31)) TRAINING & DEVELOPMENT | 9.

Types of FISH probes (continued) • Dual fusion probes – Used to confirm presence of a translocation (e. g. , t(9; 22)) – Fusion signal on each partner chromosome – Highly specific (very low false positives) • Break-apart probes – Used to confirm rearrangements of genes – 3’ portion of gene or region in one color, 5’ in another – If rearranged, colors are separated TRAINING & DEVELOPMENT | 10.

Examples of karyotype & FISH abnormalities • Trisomy 8 (or +8) using CEP • Translocation of chromosome 9 & 22 (t(9; 22)) using dualfusion probe TRAINING & DEVELOPMENT | 11.
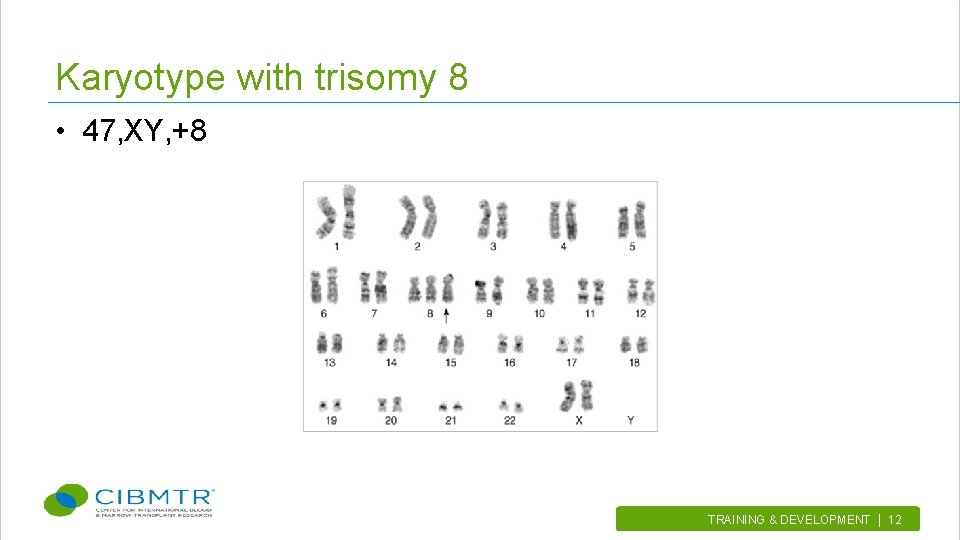
Karyotype with trisomy 8 • 47, XY, +8 TRAINING & DEVELOPMENT | 12.
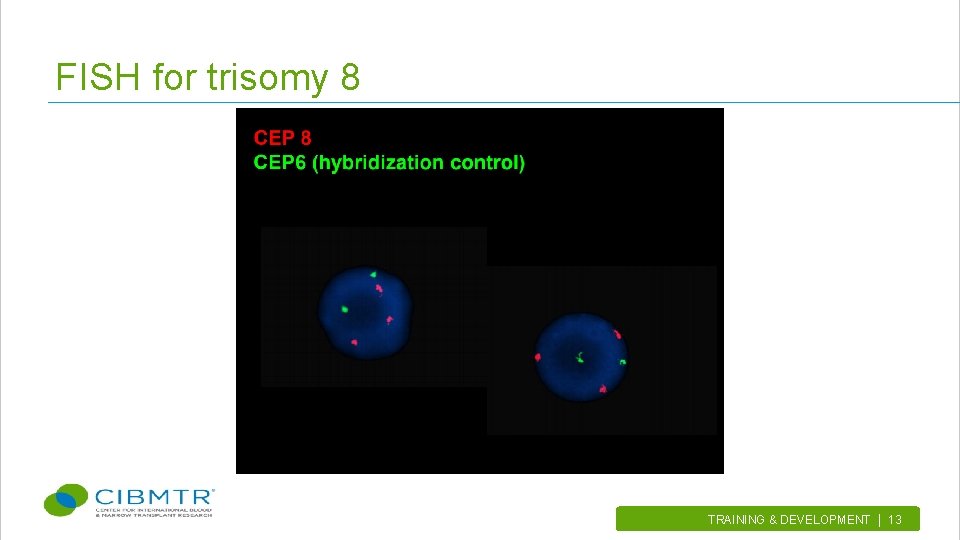
FISH for trisomy 8 TRAINING & DEVELOPMENT | 13.
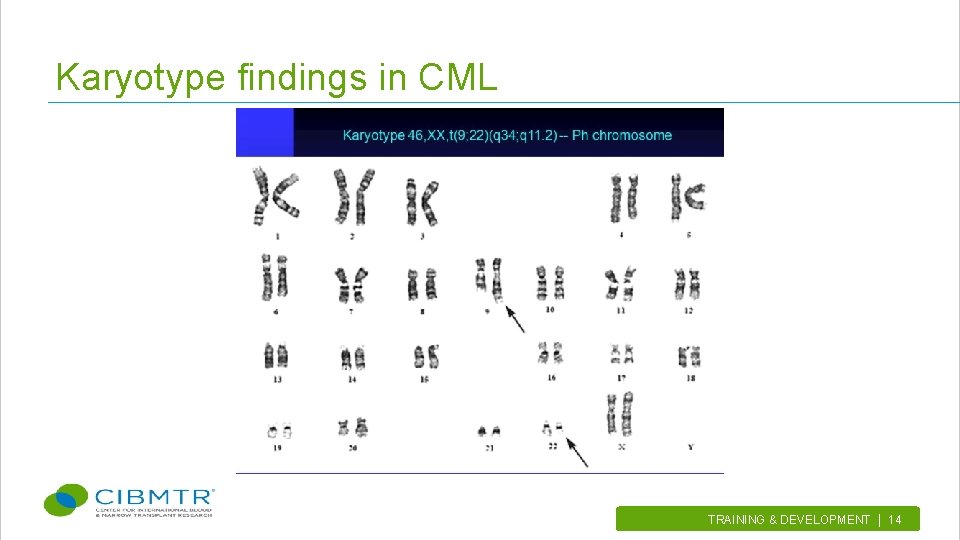
Karyotype findings in CML TRAINING & DEVELOPMENT | 14.

Dual-fusion probes TRAINING & DEVELOPMENT | 15.
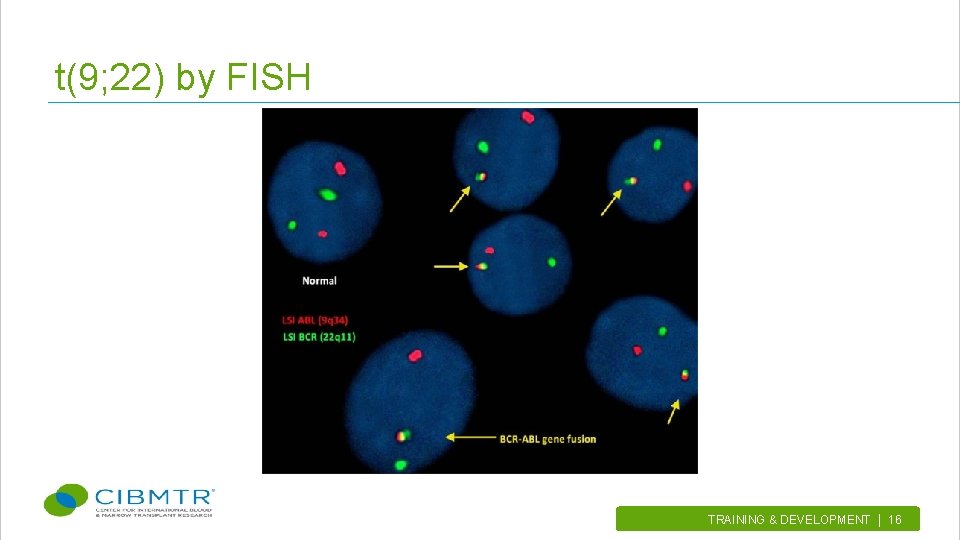
t(9; 22) by FISH TRAINING & DEVELOPMENT | 16.

Case Study #1 • The following karyotype is from a 55 y/o WM with newly diagnosed AML. • Cytogenetic report 20 metaphases analyzed 43 XY, add(4)(q 21), add(5)(q 11. 2), add 7(q 11. 2), -11, add(12)(p 11. 2), add(15)(p 11. 2), -16, -18, add(19)(q 13. 3), add(22)(p 11. 2)[4] 46, XY[16] TRAINING & DEVELOPMENT | 17.

Case Study #1 • What to report on the Pre-TED Disease form (F 2402)…. • Cytogenetic report 43 XY, add(4)(q 21), add(5)(q 11. 2), add(7)(q 11. 2), -11, add(12)(p 11. 2), add(15)(p 11. 2), -16, -18, add(19)(q 13. 3), add(22)(p 11. 2) TRAINING & DEVELOPMENT | 18.

Case Study #1 • F 2402 r 3 AML disease section Q 3 -89 Cytogenetics tested via karyotyping: • There are three monosomy chromosomes listed in the report: -11, -16, 18 • Q 18 option ‘-18’ would be checked, while the -11 & -16 would be reported in Q 19 ‘Specify other abnormality’ TRAINING & DEVELOPMENT | 19.

Case Study #1 • How to report the rest of the abnormalitiesadd(4 q), add(5 q), add(7 q), add(12 p), add(15 p), add(19 q) & add(22 p) Q 18 Specify abnormalities • All the abnormalities listed above would be reported in Q 19 or simply write “See attached cytogenetic report”. TRAINING & DEVELOPMENT | 20.

CIBMTR Request • Please submit a copy of the chromosome reports (karyotype and /or FISH) • It will decrease the number of queries sent related to chromosome abnormalities! TRAINING & DEVELOPMENT | 21.

Case Study #2 • The following karyotype is from a 10 y/o AAF with relapsed ALL. • Cytogenetic report 20 metaphases analyzed 28, X, +8, +10, +18, +21[7]; 46, XX[13] Interpretation: near haploid clone detected • The report could have been written this way- 28, XX, -1, -2, -3, -4, -5, -6, -7, -9, -11, -12, -13, -14, -15, -16, -17, -19, -20, -22[7]; 46, XX[13] TRAINING & DEVELOPMENT | 22.

Case Study #2 • How to report the following karyotype on the Pre-TED Disease form (F 2402 r 3)…. . 28, X, +8, +10, +18, +21[7]; 46, XX[13] • How would you answer Q 104? A) Check options +8 & +21 B) None of the abnormalities should be reported here TRAINING & DEVELOPMENT | 23.

Case Study #2 • Keep in mind this karyotype was a “near haploid clone”. What does that even mean? – It means most of the chromosomes only had one copy instead of the normal two. – In this case, indicating +8, +10, +18 & +21 means there are two copies and not three copies as in the case of a trisomy finding. 28, X, +8, +10, +18, +21[7]; 46, XX[13] TRAINING & DEVELOPMENT | 24.

Case Study #2 • How to report the rest of the abnormalities… -1, -2, -3, -4, -5, -6, -7, -9, -11, -12, -13, -14, -15, -16, -17, -19, -20, -22 TRAINING & DEVELOPMENT | 25.

Case Study #2 • How to report the rest of the abnormalities… -1, -2, -3, -4, -5, -6, -9, -11, -12, -13, -14, -15, -16, -17, -19, -20, -22 Q 104 • ‘Hypodiploid’ and ‘Other abnormality’ would be checked • Report all the remaining abnormalities in Q 105 or write “See attached cytogenetic report”. TRAINING & DEVELOPMENT | 26.

CIBMTR Request • Please submit a copy of the chromosome reports (karyotype and /or FISH) • Again, it will decrease the number of queries sent related to chromosome abnormalities! TRAINING & DEVELOPMENT | 27.

Case Study #3 • Interpreting FISH results can be a challenge…. – The CEP 8 (8 centromere) and D 20 S 108 (20 q 12) dual-color probe set were used for enumeration analysis. Of 200 cells examined, 15 (7. 5%) demonstrated three signals for the 8 centromere and two signals for 20 q 12. – The RUNX 1 T 1 (8 q 21. 3) / RUNX 1 (21 q 22. 1) dual-color, dual fusion probe set was used for detection of RUNX 1 T 1/RUNX 1 fusion associated with t(8; 21). Of 200 cells examined, 14 (7%) demonstrated three signals for RUNX 1 T 1 and two signals for RUNX 1. There was no evidence of fusion. TRAINING & DEVELOPMENT | 28.

Case Study #3 • What do the FISH results tell us? • 15 of 200 cells (7. 5%) demonstrated three signals for the 8 centromere and two signals for 20 q 12. § The three signals indicate the presence of 3 copies of chromosome 8 (i. e. , trisomy 8 or +8) and the two signals for 20 q 12 indicate a normal copy number of chromosome 20. • 14 of 200 cells (7%) demonstrated three signals for RUNX 1 T 1 (associated with chromosome 8 q 21. 3) and 2 signals for RUNX 1 (21 q 22. 1). § The three signals indicate the presence of 3 copies of chromosome 8 (i. e. , trisomy 8 or +8) and the 2 signals for RUNX 1 (21 q 22. 1) indicate a normal copy number of chromosome 21. There’s no evidence of t(8; 21). TRAINING & DEVELOPMENT | 29.

Molecular Marker Testing • Purpose Ø Diagnostic accuracy Ø Prognostic markers to guide therapy options and estimate outcomes Ø Monitor for minimal residual disease TRAINING & DEVELOPMENT | 30.

Types of Molecular Testing • Polymerase Chain Reaction (PCR) – Amplifies DNA fragments • Gene Expression Profiling (microarray technology) – Identifies a molecular signature of a tumor • Proteomics (microarray technology) – Identifies protein expression profiles of tissue/cell type • Single nucleotide polymorphism (SNP) & Comparative genomic hybridization (CGH) • Next Generation Sequencing (NGS) TRAINING & DEVELOPMENT | 31.

Mutations in AML • • CEBPA FLT 3 - D 835 point mutation FLT 3 - ITD mutation IDH 1 IDH 2 KIT NPM 1 TRAINING & DEVELOPMENT | 32.

AML: Example • AML/MDS Gene Mutation Panel TRAINING & DEVELOPMENT | 33.

AML: F 2402 r 3 Q 26 -27 • FLT 3 - ITD allelic ratio – Refers to the # of ITD-mutant alleles compared to the number of wild -type alleles (normal alleles) – In the example, the mutant allele burden is 70%, therefore the wildtype alleles (normal) would be 30%. The ratio = 2. 3 (70: 30) TRAINING & DEVELOPMENT | 34.
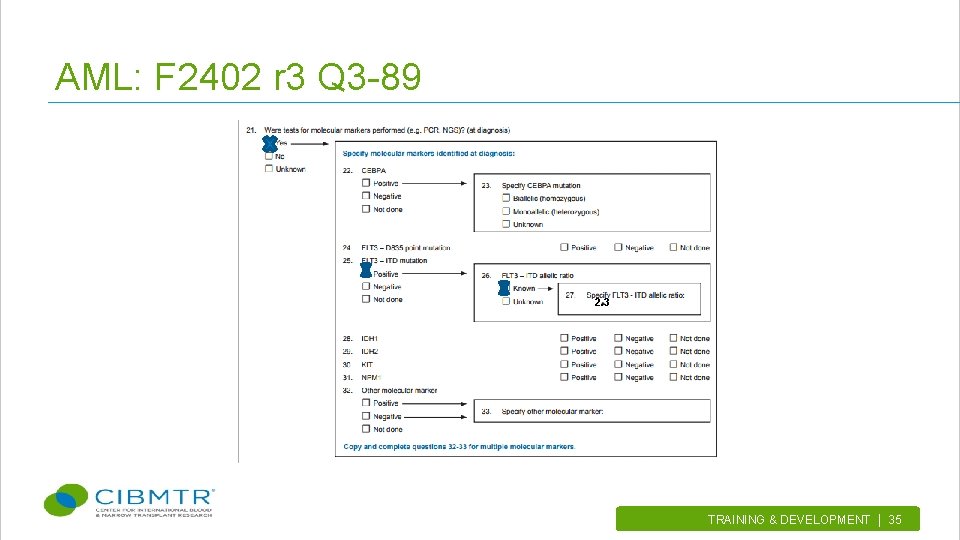
AML: F 2402 r 3 Q 3 -89 2. 3 TRAINING & DEVELOPMENT | 35.

FLT 3 -ITD allelic ratio • 2017 European Leukemia Net (ELN) guidelines defined 0. 5 as the cutoff between low & high allelic ratios – Low allelic ratio < 0. 5 (FLT 3 -ITDlow) – High allelic ratio > 0. 5 (FLT 3 -ITDhigh) • A low FLT 3 -ITD allelic ratio is a favorable prognostic factor TRAINING & DEVELOPMENT | 36.

Mutations in MDS/MPN • • • ASXL 1 JAK 2 (MPN only) ETV 6 EZH 2 P 53 RUNX 1 TRAINING & DEVELOPMENT | 37.

Case Report. Patient with Primary Myelofibrosis Panel reports data on 21 genes for myeloid malignancies TRAINING & DEVELOPMENT | 38.

Case Report Continued
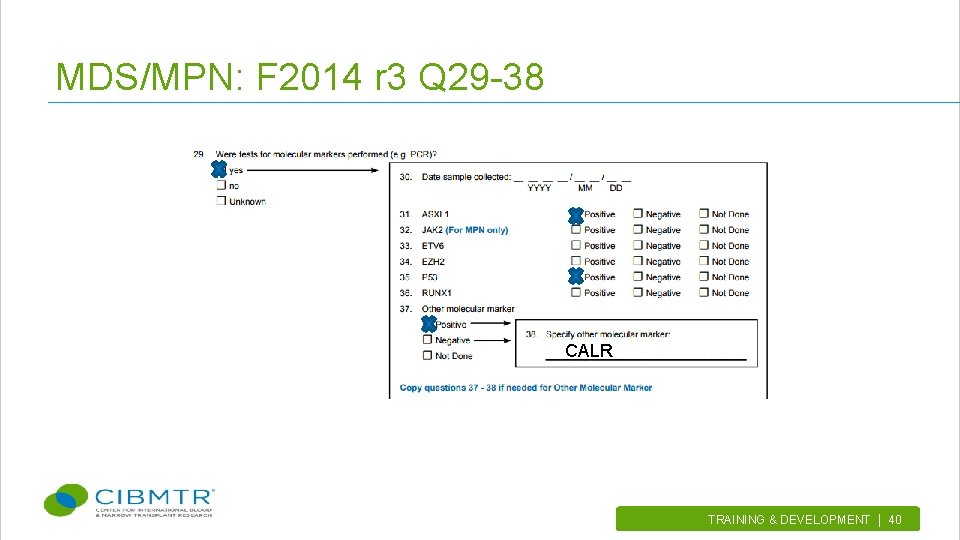
MDS/MPN: F 2014 r 3 Q 29 -38 CALR TRAINING & DEVELOPMENT | 40.
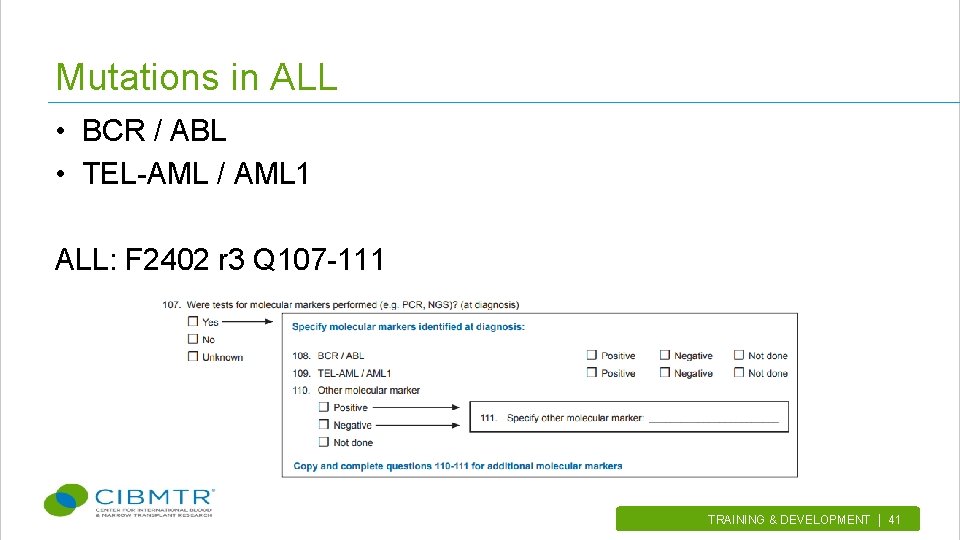
Mutations in ALL • BCR / ABL • TEL-AML / AML 1 ALL: F 2402 r 3 Q 107 -111 TRAINING & DEVELOPMENT | 41.
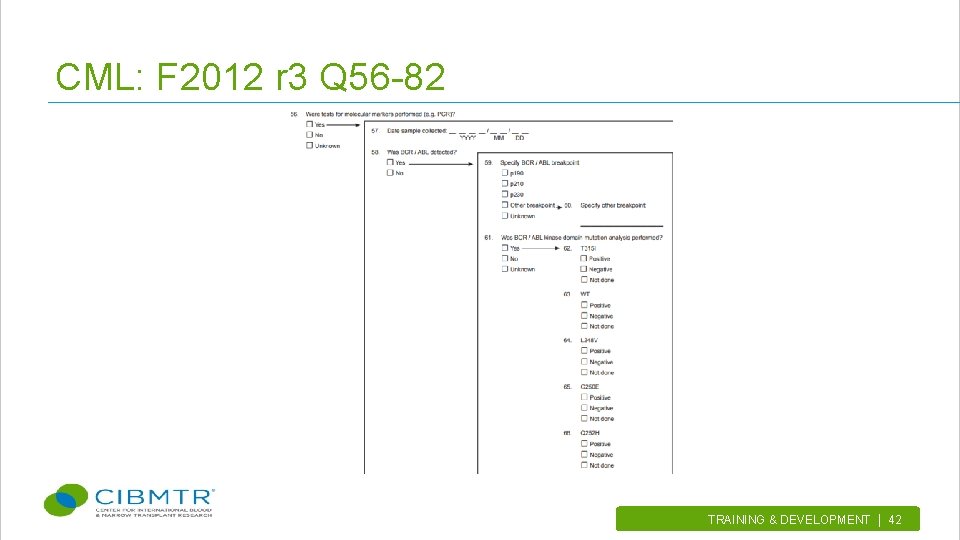
CML: F 2012 r 3 Q 56 -82 TRAINING & DEVELOPMENT | 42.
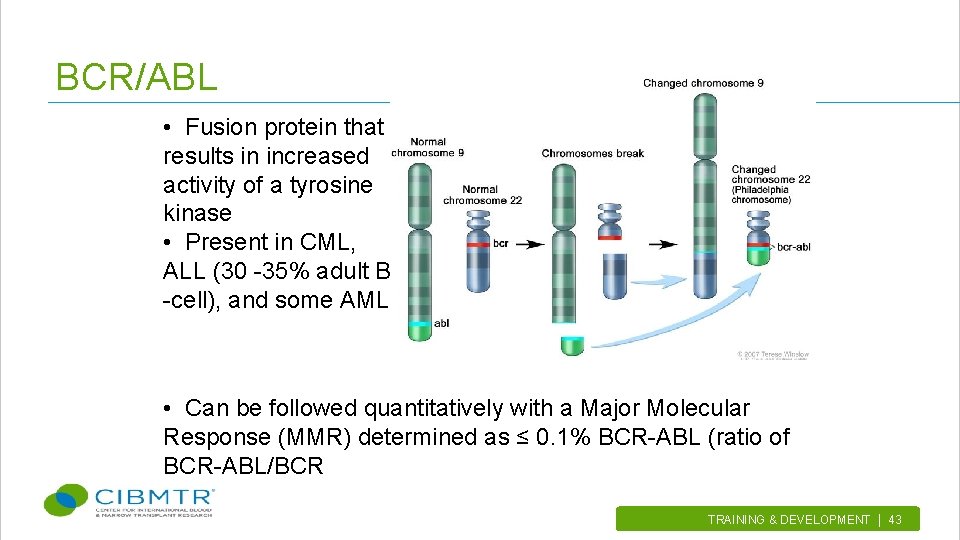
BCR/ABL • Fusion protein that results in increased activity of a tyrosine kinase • Present in CML, ALL (30 -35% adult B -cell), and some AML • Can be followed quantitatively with a Major Molecular Response (MMR) determined as ≤ 0. 1% BCR-ABL (ratio of BCR-ABL/BCR TRAINING & DEVELOPMENT | 43.

Mutations in CLL • • Immunoglobulin heavy chain variable (IGHV) mutation NOTCH 1 TP 53 (or P 53) SF 3 B 1 TRAINING & DEVELOPMENT | 44.
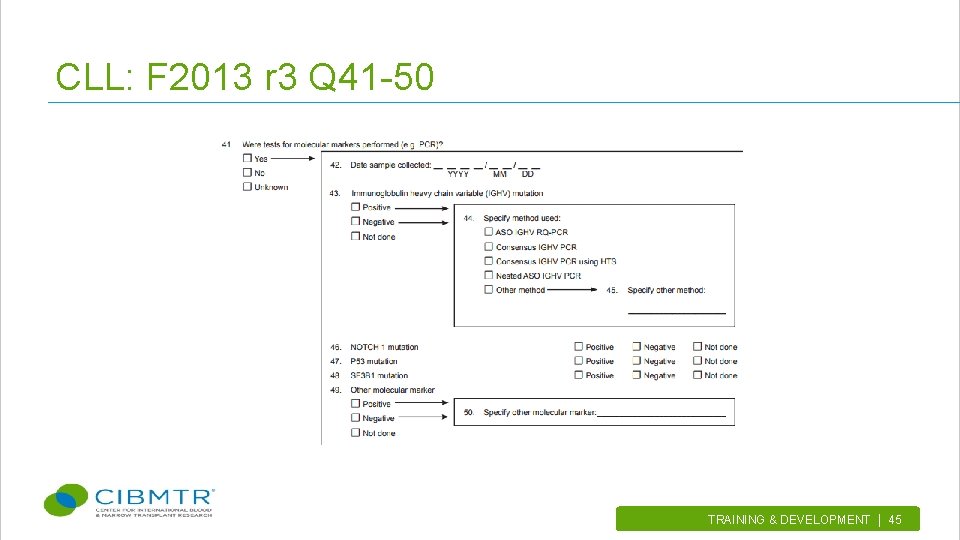
CLL: F 2013 r 3 Q 41 -50 TRAINING & DEVELOPMENT | 45.

Mutations in Lymphomas • • • BCL-1/Cyclin D 1; t(11; 14) BCL 2; t(14; 18) BCL-6 B-cell, Immunoglobulin Heavy (Ig. H) chain rearrangement T-cell receptor (TCR) gene rearrangement TRAINING & DEVELOPMENT | 46.

Questions TRAINING & DEVELOPMENT | 47.
- Slides: 47