Remediation of Phenolic Resin Using E coli Containing





























- Slides: 31

Remediation of Phenolic Resin Using E. coli Containing Li. P Gene from P. chrysosporium Team Fun Guys: Lorena Christensen, Michelle Fretheim, Gabriel Martin

Our Goal Insert fungus Phanerochaete chrysosporium’s Lignin Peroxidase (Li. PD) gene into E. coli to see if we could enable the bacteria to break down phenolic resin (plastic). Enzymes coded for by Li. PD are able to break carbon bonds in phenolic resins.

Phanerochaete chrysosporium’s Lignin Peroxidase (Li. PD) gene

Original Plan Build smaller exon segments using oligonucleotides. Use primers to extract larger exons from DNA. Run PCR for both to amplify gene’s exons. Ligate exons. Transform E. coli with Li. PD gene from fungus P. chrysosporium. Plate on phenolic resin to observe remediation if successful.

Modified Plan Due to difficulty of constructing primer sets and removing introns, we decided to attempt RNA extraction instead. Discovered there are no existing biobricks for Lip. D gene. We would have to build and integrate these ourselves.

What Actually Went Down… (summary… details on the following slides) Cultured P. chrysosporium DNA extraction from P. chrysosporium Built and ordered primers - one set with biobrick extension, one without. PCR with primers Attempted RNA extraction from P. chrysosporium (FAIL) Ligation of Li. PD gene into p. GEM-T Easy Vector Transformation of E. coli with vector containing Li. PD gene Miniprep of plasmid, digestion using Eco. R 1 Sequenced select samples from digestion to see if Li. PD gene was present.

What Actually Went Down… Cultured P. chrysosporium Liquid cultures of mycelium using slant culture obtained from Dr. Bumpus. Incubated at room temperature for five days in an agar suspension.

What Actually Went Down… DNA Extraction from P. chrysosporium Filtered mycelial mats from liquid agar, let dry. Divided into two samples as a precautionary measure: S 1 and S 2. Protocol used: “Rapid procedure for the extraction of DNA from fungal spores and mycelia, ” by John J. Weiland. http: //www. fgsc. net/fgn 44/weiland. html Deviation from protocol: During extraction initial mixture was too viscous, so increased volume of extraction buffer added.

What Actually Went Down… DNA Extraction from P. chrysosporium Ran agarose gel electrophoresis to verify if DNA was present DNA!!!! Sample #1 Ladder Sample #2 18 µl 6 µl 18 µl

What Actually Went Down… Built and Ordered Primers Sneezy (forward + biobrick extension) GTTTCTTC GAATTCGCGG CCGCTTCTAG CCTTCGTATG TAAGTCGCTG Grumpy (forward without biobrick extension) CCTTCGTATG TAAGTCGCTG Cowgirl (reverse + biobrick extension) TACTAGTAG CGGCCGCTGC AGGAAGAAAC CGCCGTGCGC GAGTCGCGCG Cowboy (reverse without biobrick extension) CGCCGTGCGC GAGTCGCGCG

What Actually Went Down… PCR With Primers Obtained primers in DNA form, made a working stock (100 u. M). Ran PCR to test primer combinations: 1) Sneezy + Cowgirl 2) Grumpy + Cowboy 3) Sneezy + Cowboy 4) Grumpy + Cowgirl Ran each set at three different temperatures: 1. 58. 0 o C 2. 51. 7 o C 3. 48. 0 o C

What Actually Went Down… PCR With Primers Ran gel electrophoresis to verify primers worked Lane 1: Sneezy Cowgirl 58. 0°C 10 µl Lane 2: empty Lane 3: Sneezy Cowgirl 51. 7°C 5 µl Lane 4: Sneezy Cowgirl 51. 7°C 15 µl Lane 5: Sneezy Cowgril 48. 0°C 5 µl Lane 6: Sneezy Cowgirl 48. 0°C 15 µl Lane 7: Grumpy Cowboy 58. 0°C 10 µl Lane 8: empty Lane 9: Grumpy Cowboy 51. 7°C 5 µl Lane 10: Grumpy Cowboy 51. 7°C 15 µl Lane 11: Grumpy Cowboy 48. 0°C 5 µl Lane 12: Grumpy Cowboy 48. 0°C 15 µl Lane 13: Ladder (Fast. Ruler Low Range) 15 µl Lane 14: Control 20 µl Lane 15: Sneezy Cowboy 58. 0°C 10 µl Lane 16: empty Lane 17: Sneezy Cowboy 51. 7°C 5 µl Lane 18: Sneezy Cowboy 51. 7°C 15 µl Lane 19: Sneezy Cowboy 48. 0°C 5 µl Lane 20: Sneezy Cowboy 48. 0°C 15 µl Lane 21: Grumpy Cowgirl 58. 0°C 10 µl Lane 22: empty Lane 23: Grumpy Cowgirl 51. 7°C 5 µl Lane 24: Grumpy Cowgirl 51. 7°C 15 µl Lane 25: Grumpy Cowgirl 48. 0°C 5 µl Lane 26: Grumpy Cowgirl 48. 0°C 15 µl

What Actually Went Down… Attempted RNA Extraction from P. chrysosporium (failtastic) Used “Rneasy Plant Mini. Kit” protocol. Ground mycelium with liquid nitrogen. Purified sample for Reverse Transcriptase – PCR. Ran RT-PCR following protocol from “QIAGEN One Step RT PCR Kit” Ran gel electrophoresis.

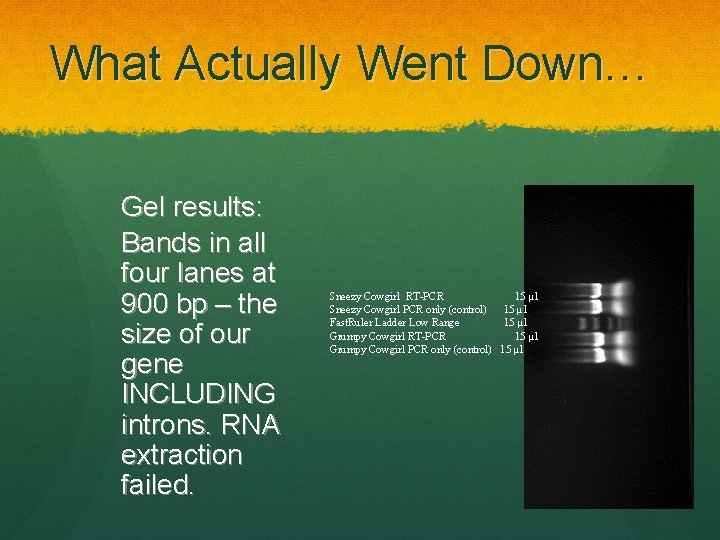
What Actually Went Down… Gel results: Bands in all four lanes at 900 bp – the size of our gene INCLUDING introns. RNA extraction failed. Sneezy Cowgirl RT-PCR 15 µl Sneezy Cowgirl PCR only (control) 15 µl Fast. Ruler Ladder Low Range 15 µl Grumpy Cowgirl RT-PCR 15 µl Grumpy Cowgirl PCR only (control) 15 µl

What Actually Went Down… RNA Extraction round two Followed same protocol with the following adjustments: Added DNase to bind up excess DNA Used RNase free materials to not destroy any RNA present Heated extraction at 56 o. C to lower viscosity. However, it had the opposite effect. Ran RT-PCR, gel electrophoresis, and still got same results as first attempt.

What Actually Went Down… Sneezy Cowgirl PCR (control) 15 µl Sneezy Cowgirl RT-PCR 15 µl Fast. Ruler Ladder Low Range 15 µl Grumpy Cowgirl PCR (control) 15 µl Grumpy Cowgirl RT-PCR 15 µl

What Actually Went Down… Ligation of Li. PD gene into p. GEM-T Easy Vector Following protocol from “QIAquick PCR Purification Kit 50”, we purified DNA obtained from failed RT-PCR. Purified two samples Grumpy+Cowgirl Sneezy+Cowgirl Ran gel electrophoresis

What Actually Went Down… Gel used to determine concentration of DNA in our samples (needed for ligation). Compared to ladder containing 60 ng DNA/ 15µl, samples are estimated to contain 30 ng DNA/ 15µl Grumpy Cowgirl Fast. Ruler Ladder Low Range Sneezy Cowgirl 15µl

What Actually Went Down… Ligation of Li. PD gene into p. GEM-T Easy Vector Did five ligation combinations: 1) Grumpy+Cowgirl 1: 1 ratio 3. 5 u. L sample + 0. 5 u. L plasmid 2) Grumpy+Cowgirl 3. 0 u. L sample + 1. 0 u. L plasmid 3) Sneezy+Cowgirl 1: 1 ratio 3. 5 u. L sample + 0. 5 u. L plasmid 4) Sneezy+Cowgirl 3. 0 u. L sample + 1. 0 u. L plasmid 5) Control 3. 0 u. L H 2 O + 1. 0 u. L plasmid

What Actually Went Down… Transformation of E. coli with vector containing Li. PD gene Followed protocol from “PGem from Promega” with adjustments. Plated ligations and controls: Control 1) H 2 O + 40 u. L competent cells Control 2) H 2 O + 40 u. L competent cells Control 3) SOC + 50 u. L Competent cells 1) 3. 5 u. L GC to 0. 5 u. L plasmid 2) 3. 0 u. L GC to 1. 0 u. L plasmid 3) 3. 5 u. L SC to 0. 5 u. L plasmid 4) 3. 0 u. L SC to 1. 0 u. L plasmid 5) 3. 0 u. L H 2 O + 1. 0 u. L plasmid (ligation control) on LB+Xgal+Amp on LB+Xgal+Amp

What Actually Went Down… Transformation of E. coli with vector containing Li. PD gene Picked four white colonies each from plates #1 -4. Added to liquid culture for miniprep of plasmid. Shook for one hour at 125 rpm

What Actually Went Down… Miniprep of plasmid, digestion using Eco. R 1 Used protocol “Gene. Jet Plasmid Mini. Prep Kit”. Cut plasmid with Eco. R 1 using protocol “Fast Digestion DNA” from class packet. Ran a gel electrophoresis to verify sizes of fragments.

What Actually Went Down… Miniprep of plasmid, digestion using Eco. R 1 gel results Lane 1: 3. 0 Grumpy Cowgirl Lane 2: 3. 5 Grumpy Cowgirl Lane 3: 3. 5 Grumpy Cowgirl Lane 4: 3. 5 Grumpy Cowgirl Lane 5: Fast. Ruler Ladder Low Range 19µl Lane 6: 3. 0 Sneezy Cowgirl Lane 7: 3. 0 Sneezy Cowgirl Lane 8: 3. 0 Sneezy Cowgirl Lane 9: 3. 5 Sneezy Cowgirl Lane 10: 3. 5 Sneezy Cowgirl Lane 11: 3. 5 Sneezy Cowgirl 19 µl 19µl 19µl 19µl

What Actually Went Down… Sent four samples (lanes 3, 4, 6, 7) from digestion to be sequenced. Ran the sequences with NCBI BLAST to see if our gene was present.

What Actually Went Down… Grumpy Cowgirl B 8 BLAST hits Red indicates an excellent alignment score for 5 of the 8 hits. Pink indicates a very good score. Descriptions of sequences with significant alignment. The top hit IS our gene of interest. (Note: Evalues indicate the likelihood that this hit is unrelated to our query. )

What Actually Went Down… Grumpy Cowgirl C Same results Again, our Lip. D gene is the top match.

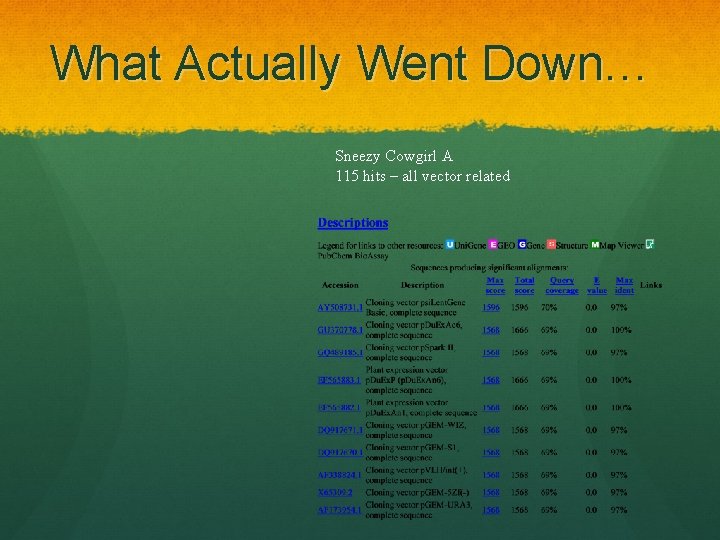
What Actually Went Down… Sneezy Cowgirl A 115 hits – all vector related

What Actually Went Down… Sneezy Cowgirl B Same results as the first two samples. Lip. D gene is top match.

SUCCESS and FAIIIIIIIL Successes: Primers worked well in multiple combinations Extracted DNA from P. chrysosporium Transformation Ligated correct gene

SUCCESS and FAIIIIIIIL Failures: RNA extractions Steps in various protocols Not limiting media, therefore suppressing Li. PD gene expression!!!!

Suggestions for further experiments RNA extraction using nutrient deprived P. chrysosporium sample Clone and assemble exon fragments, as we originally planned.