Regulatory Motif Finding Clustering of Genes Find binding

(Regulatory-) Motif Finding

Clustering of Genes Find binding sites responsible for common expression patterns
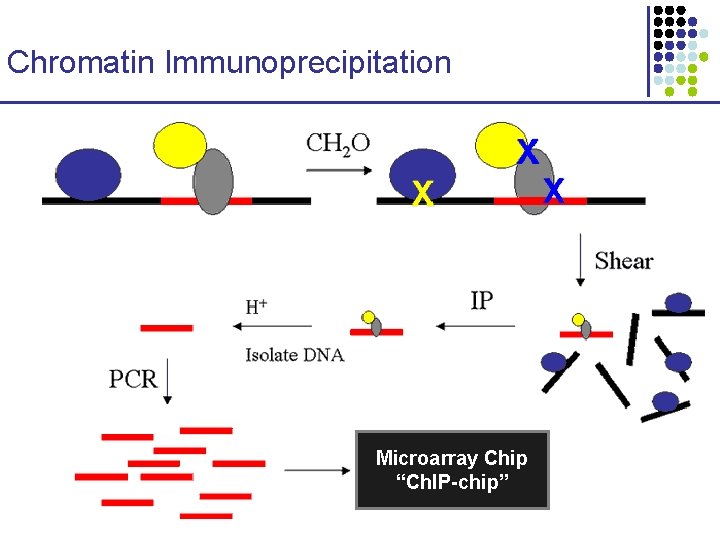
Chromatin Immunoprecipitation Microarray Chip “Ch. IP-chip”

Finding Regulatory Motifs . . . Given a collection of genes with common expression, Find the TF-binding motif in common

Characteristics of Regulatory Motifs • Tiny • Highly Variable • ~Constant Size § Because a constant-size transcription factor binds • Often repeated • Low-complexity-ish
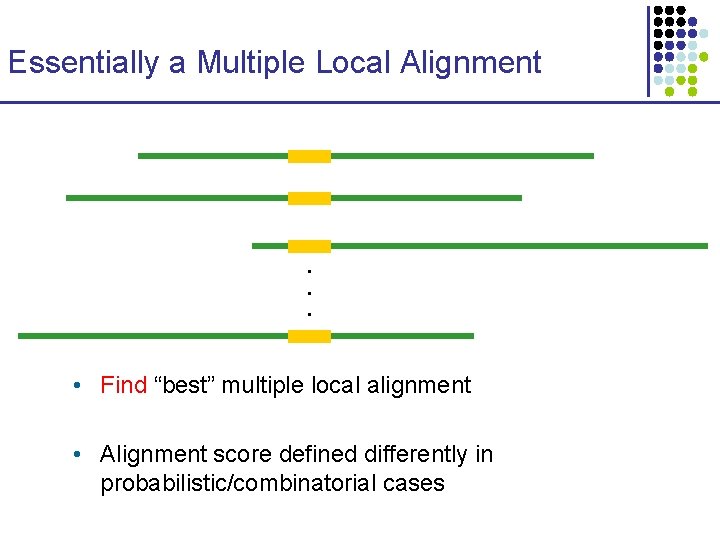
Essentially a Multiple Local Alignment . . . • Find “best” multiple local alignment • Alignment score defined differently in probabilistic/combinatorial cases

Algorithms • Combinatorial CONSENSUS, TEIRESIAS, SP-STAR, others • Probabilistic 1. Expectation Maximization: MEME 2. Gibbs Sampling: Align. ACE, Bio. Prospector

Combinatorial Approaches to Motif Finding

Discrete Formulations Given sequences S = {x 1, …, xn} • A motif W is a consensus string w 1…w. K • Find motif W* with “best” match to x 1, …, xn Definition of “best”: d(W, xi) = min hamming dist. between W and any word in xi d(W, S) = i d(W, xi)

Approaches • Exhaustive Searches • CONSENSUS • MULTIPROFILER • TEIRESIAS, SP-STAR, WINNOWER

Exhaustive Searches 1. Pattern-driven algorithm: For W = AA…A to TT…T Find d( W, S ) Report W* = argmin( d(W, S) ) (4 K possibilities) Running time: O( K N 4 K ) (where N = i |xi|) Advantage: Finds provably “best” motif W Disadvantage: Time

Exhaustive Searches 2. Sample-driven algorithm: For W = any K-long word occurring in some xi Find d( W, S ) Report W* = argmin( d( W, S ) ) or, Report a local improvement of W* Running time: O( K N 2 ) Advantage: Time Disadvantage: If the true motif is weak and does not occur in data then a random motif may score better than any instance of true motif

MULTIPROFILER • Extended sample-driven approach Given a K-long word W, define: Nα(W) = words W’ in S s. t. d(W, W’) α Idea: Assume W is occurrence of true motif W* Will use Nα(W) to correct “errors” in W

MULTIPROFILER Assume W differs from true motif W* in at most L positions Define: A wordlet G of W is a L-long pattern with blanks, differing from W § L is smaller than the word length K Example: K = 7; L = 3 W G = = ACGTTGA --A--CG

MULTIPROFILER Algorithm: For each W in S: For L = 1 to Lmax 1. Find the α-neighbors of W in S 2. Find all “strong” L-long wordlets G in Na(W) 3. For each wordlet G, 1. Modify W by the wordlet G 2. Compute d(W’, S) Report W* = argmin d(W’, S) Step 1 above: Smaller motif-finding problem; Use exhaustive search Nα(W) W’

CONSENSUS Algorithm: Cycle 1: For each word W in S For each word W’ in S Create alignment (gap free) of W, W’ (of fixed length!) Keep the C 1 best alignments, A 1, …, AC 1 ACGGTTG ACGCCTG , , CGAACTT AGAACTA , , GGGCTCT … GGGGTGT …

CONSENSUS Algorithm: Cycle t: For each word W in S For each alignment Aj from cycle t-1 Create alignment (gap free) of W, Aj Keep the Cl best alignments A 1, …, ACt ACGGTTG ACGCCTG … ACGGCTC , , , CGAACTT AGAACTA … AGATCTT , , , GGGCTCT … GGGGTGT … … GGCGTCT …

CONSENSUS • C 1, …, Cn are user-defined heuristic constants § N is sum of sequence lengths § n is the number of sequences Running time: O(N 2) + O(N C 1) + O(N C 2) + … + O(N Cn) = O( N 2 + NCtotal) Where Ctotal = i Ci, typically O(n. C), where C is a big constant

Expectation Maximization in Motif Finding

Expectation Maximization motif All K-long background words Algorithm (sketch): 1. 2. 3. Given genomic sequences find all K-long words Assume each word is motif or background Find likeliest Motif Model Background Model classification of words into either Motif or Background
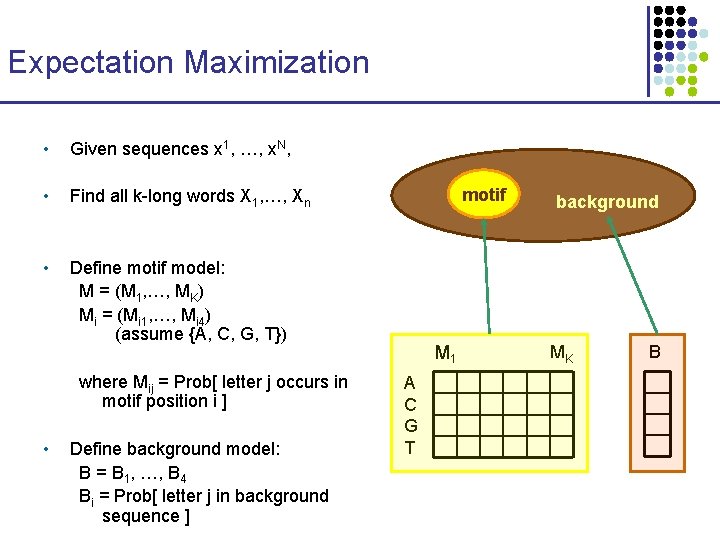
Expectation Maximization • Given sequences x 1, …, x. N, • Find all k-long words X 1, …, Xn • Define motif model: M = (M 1, …, MK) Mi = (Mi 1, …, Mi 4) (assume {A, C, G, T}) where Mij = Prob[ letter j occurs in motif position i ] • Define background model: B = B 1, …, B 4 Bi = Prob[ letter j in background sequence ] motif M 1 A C G T background MK B
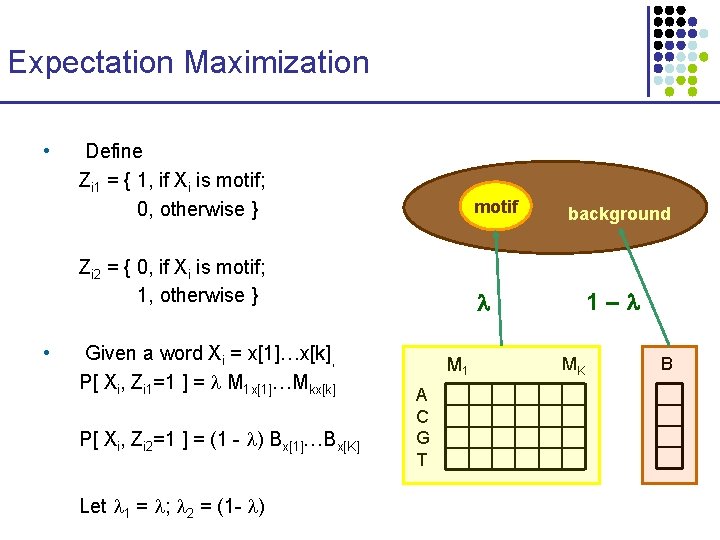
Expectation Maximization • • Define Zi 1 = { 1, if Xi is motif; 0, otherwise } motif Zi 2 = { 0, if Xi is motif; 1, otherwise } Given a word Xi = x[1]…x[k], P[ Xi, Zi 1=1 ] = M 1 x[1]…Mkx[k] P[ Xi, Zi 2=1 ] = (1 - ) Bx[1]…Bx[K] Let 1 = ; 2 = (1 - ) M 1 A C G T background 1– MK B
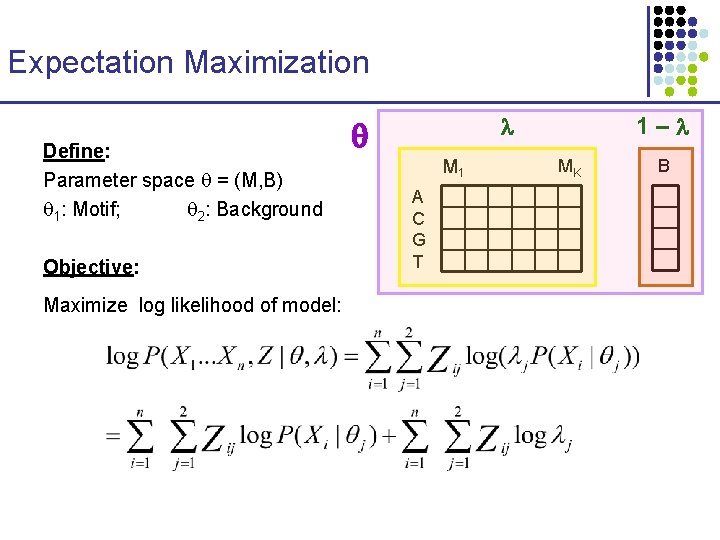
Expectation Maximization Define: Parameter space = (M, B) 1: Motif; 2: Background Objective: Maximize log likelihood of model: 1– M 1 A C G T MK B

Expectation Maximization • Maximize expected likelihood, in iteration of two steps: Expectation: Find expected value of log likelihood: Maximization: Maximize expected value over ,

Expectation Maximization: E-step Expectation: Find expected value of log likelihood: where expected values of Z can be computed as follows:

Expectation Maximization: M-step Maximization: Maximize expected value over and independently For , this is easy:

Expectation Maximization: M-step • For = (M, B), define cjk = E[ # times letter k appears in motif position j] c 0 k = E[ # times letter k appears in background] • cij values are calculated easily from Z* values It easily follows: to not allow any 0’s, add pseudocounts

Initial Parameters Matter! Consider the following “artificial” example: x 1, …, x. N contain: § 212 patterns on {A, T}: A…A, A…AT, ……, T… T § 212 patterns on {C, G}: C…C , C…CG, …… , G…G § D << 212 occurrences of 12 -mer ACTGACTG Some local maxima: ½; B = ½C, ½G; Mi = ½A, ½T, i = 1, …, 12 D/2 k+1; B = ¼A, ¼C, ¼G, ¼T; M 1 = 100% A, M 2= 100% C, M 3 = 100% T, etc.

Overview of EM Algorithm 1. Initialize parameters = (M, B), : § Try different values of from N-1/2 up to 1/(2 K) 2. Repeat: a. Expectation b. Maximization 3. Until change in = (M, B), falls below 4. Report results for several “good”

Overview of EM Algorithm • One iteration running time: O(NK) § Usually need < N iterations for convergence, and < N starting points. § Overall complexity: unclear – typically O(N 2 K) - O(N 3 K) • EM is a local optimization method M, B • Initial parameters matter MEME: Bailey and Elkan, ISMB 1994.
- Slides: 30