Regulation of Gene Expression in Eukaryotes By Amr

Regulation of Gene Expression in Eukaryotes By Amr S Moustafa, MD, Ph. D Medical Biochemistry and Molecular Biology Department Faculty of Medicine, Ain Shams University amrsm@hotmail. com

Intended learning Outcomes (ILOs): By the end of theses 2 lectures, you should be able to: 1. Identify different levels of gene expression 2. Recognize the difference between constitutive Vs regulated genes 3. Describe regulation of gene expression at the level of transcription 4. Identify the regulatory mechanisms at level of posttranscriptional processing of m. RNA 5. Identify the epigenetic mechanisms for control of gene expression
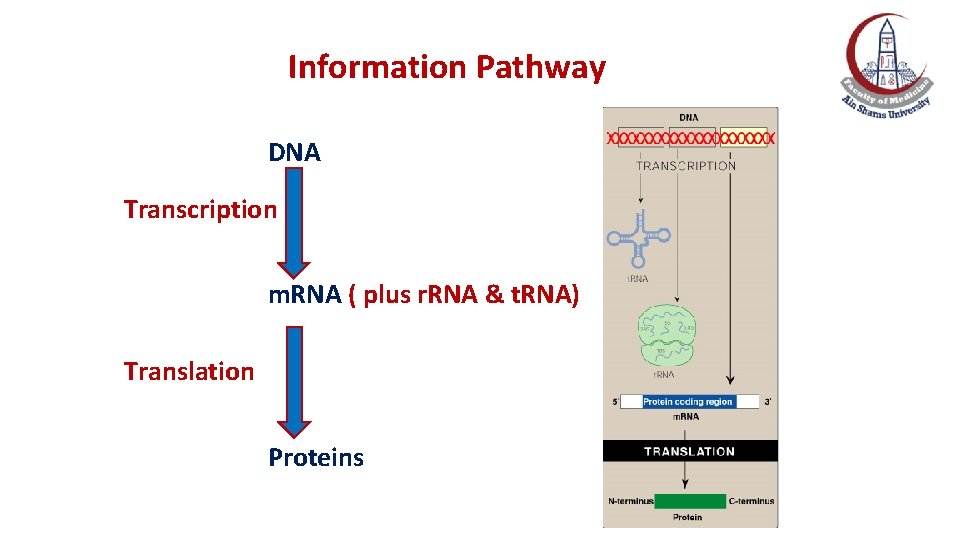
Information Pathway DNA Transcription m. RNA ( plus r. RNA & t. RNA) Translation Proteins

Levels of Gene Expression Level 1: RNA synthesis (transcription) Level 2: Protein synthesis (translation) The final product of gene expression can be RNA or protein: Ø For m. RNA-encoded genes: peptides or proteins are the final products Ø For r. RNA-, t. RNA- and sn. RNA-encoded genes: RNA is the final product
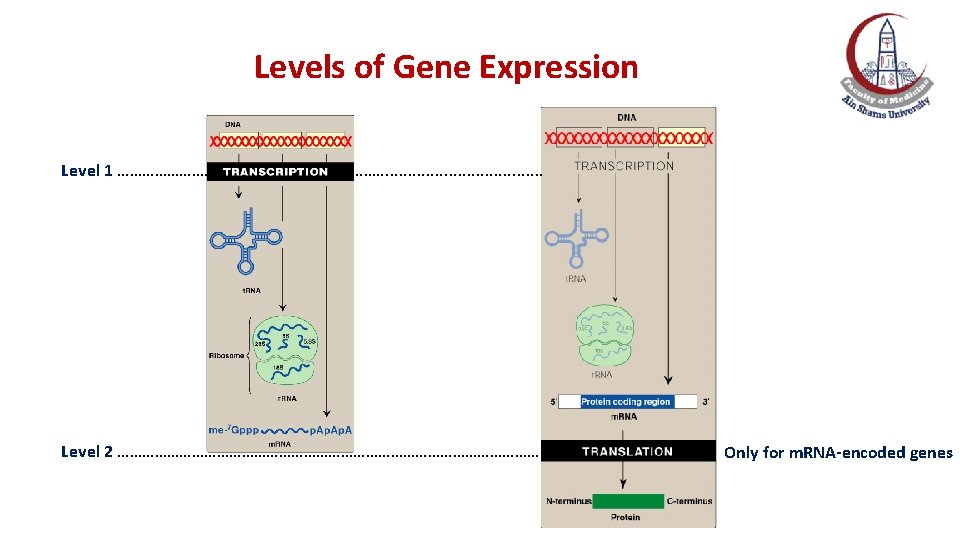
Levels of Gene Expression Level 1 ………… ………. . . . . Level 2 …………………………………………… Only for m. RNA-encoded genes

Constitutive Vs Regulated Genes Not all genes are regulated Constitutive or housekeeping genes: Genes that encode products required for basic cellular functions They are continually expressed (not regulated) Regulated genes: They are expressed only under certain conditions They maybe expressed in all cells or only in a subset of cells (differential or cell specific expression) Their expression is regulated (Where? , When? , and How much? ) Their regulated expression is the basis for: cellular differentiation, morphogenesis and adaptability of organisms

Regulation of Gene Expression: Different Control Sites 1. Control at the level of transcription 2. Control at the level of posttranscriptional processes of m. RNA 3. Control by nc. RNA-related mechanisms 4. Control by DNA-related mechanisms in eukaryotes

1. Control at the Level of Transcription q Main regulatory process q Interaction between cis-acting DNA sequences and trans-acting proteins (basal and specific transcription factors)
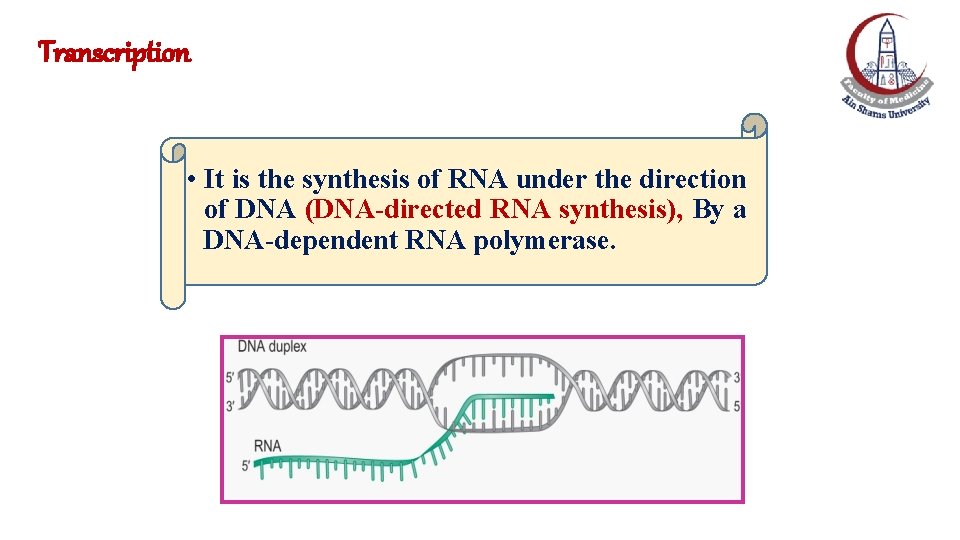
Transcription • It is the synthesis of RNA under the direction of DNA (DNA-directed RNA synthesis), By a DNA-dependent RNA polymerase.

Glossary: Cis-acting dna sequences • Cis-acting elements: DNA sequences on the same chromosome as the gene whose transcription they stimulate: 1. Promotor sequences: Upstream of the transcription start site (+1 position), serve as binding sites for basal transcription protein factors and RNA polymerase. 2. Enhancer sequences: Upstream or downstream of the transcription start site (+1 position), close or thousands base pairs away from the promoter, serve as binding sites for specific transcription factors (activators). 3. Silencer sequences: Similar to enhancer sequences but serve as binding sites for specific transcription factors (repressors).

Glossary: trans-acting transcription Factors (protein factors) • Trans-acting elements: Protein factors that are encoded by different genes, synthesized in the cytosol and then translocated into the nucleus (hence the name as “trans”) to control transcription of genes. 1. General (basal) transcription factors: protein factors that recognize and bind to promotor sequences for the initiation of transcription through recruitment of RNA polymerase (i. e. , both protein/DNA and protein/protein interactions). 2. Specific transcription factors: Protein factors (activators or repressors) that bind to DNA sequences. Activators bind to enhancer sequences & repressors bind to silencers DNA sequences. They regulate transcription in response to different signals, e. g. , hormones. Through bending of DNA, specific TFs can also interact with the basal TFs and RNA polymerase to modulate the efficiency of initiation of transcription. Note: Co-activators: are proteins that bind to and interact with the specific TFs and control transcription. In contrast to activators, coactivators does not bind to DNA (have no DNA-binding domain), e. g. , HAT.

General or basal TFs Bind to consensus sequences of promoter; e. g. , CTF, SP 1 and TFIID Facilitate assembly of the initiation complex and recruitment of RNA polymerase II, e. g. , TFIIF Catalyze basal or constitutive transcription

Specific or Regulatory TFs • Bind to regulatory sequences of DNA known as enhancers or silencers • Modulate the efficiency of initiation of transcription • Mediate the response to signals, e. g. , hormones • Regulate which genes are expressed at a given time

Specific or Regulatory TFs

Specific or Regulatory TFs • Specific TFs interact with: DNA sequences (enhancer or silencer) by DNA-binding domain General TFs and RNAP at the initiation complex by protein-binding domain Coactivator proteins (e. g. , histone acetyltransferase) by transcription activation domain
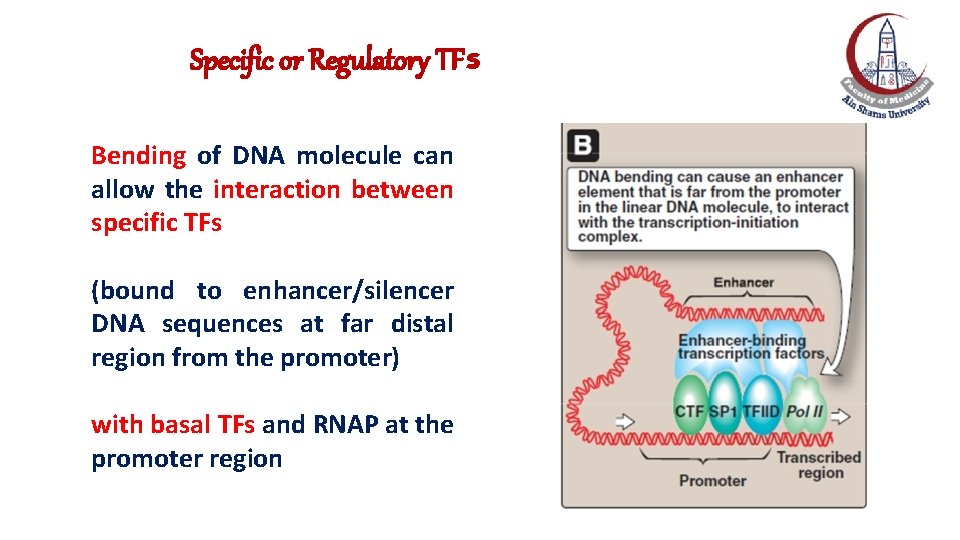
Specific or Regulatory TFs Bending of DNA molecule can allow the interaction between specific TFs (bound to enhancer/silencer DNA sequences at far distal region from the promoter) with basal TFs and RNAP at the promoter region

Cis-acting element: DNA Enhancers v. An enhancer is a special DNA sequence which increases the rate of transcription of a specific gene. v They must be located on the same chromosome as the gene to be transcribed. v. An enhancer can be located close to the genes it acts on, or far away from the promoter. v. Are located either upstream (to the 5`-side) or downstream (to the 3`-side) to the transcription start site.
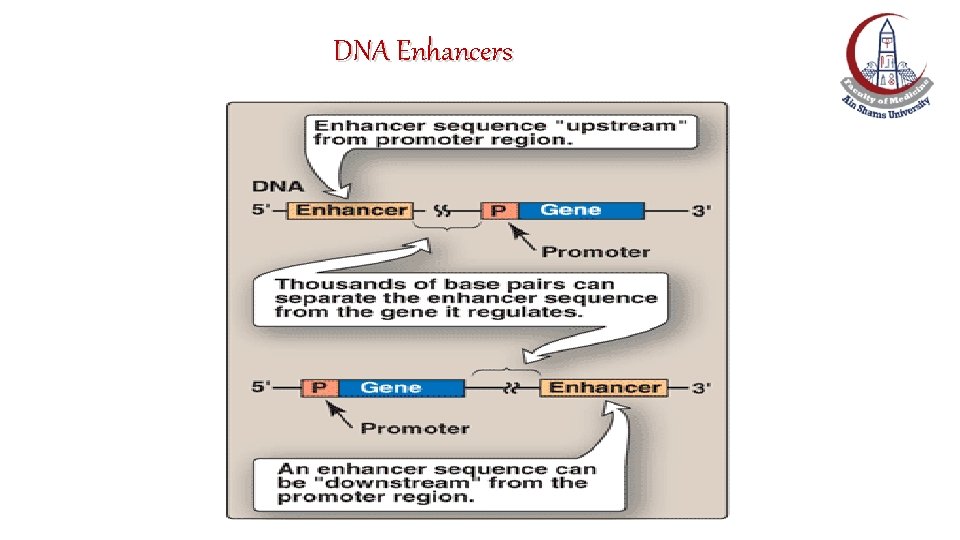
DNA Enhancers
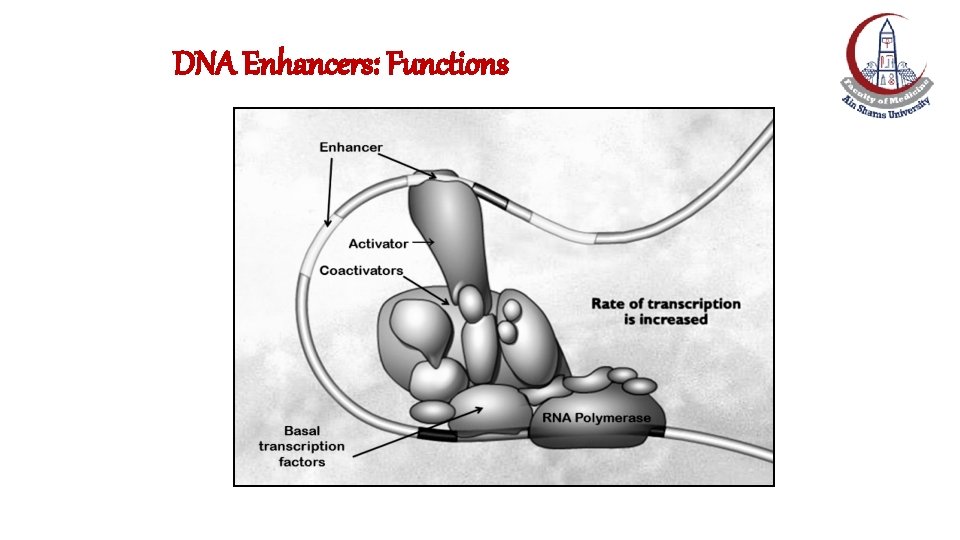
DNA Enhancers: Functions
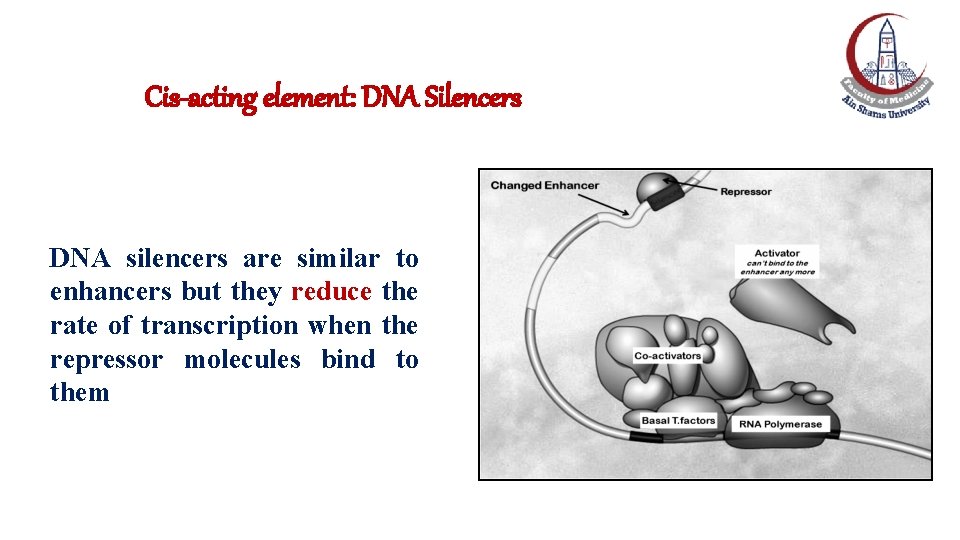
Cis-acting element: DNA Silencers DNA silencers are similar to enhancers but they reduce the rate of transcription when the repressor molecules bind to them

2. Regulation of Gene Expression at m. RNA Posttranscriptional Level q Splice-site choice and differential tissue expression q m. RNA editing: modifications for m. RNA after it is fully processed q m. RNA stability and translation repression (si. RNA/mi. RNA)

Splice-site choice: Alternative splicing of m. RNA molecules • It produces multiple variations of the m. RNA and, therefore, of its protein product. • This appears to be a mechanism for producing a diverse set of proteins (>100, 000 proteins) from a limited set of genes (~25, 000 genes).
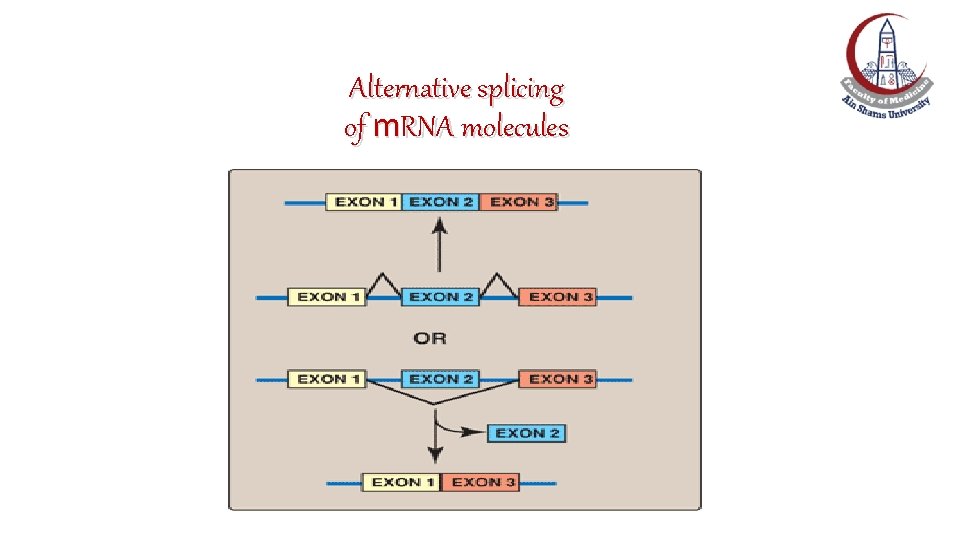
Alternative splicing of m. RNA molecules
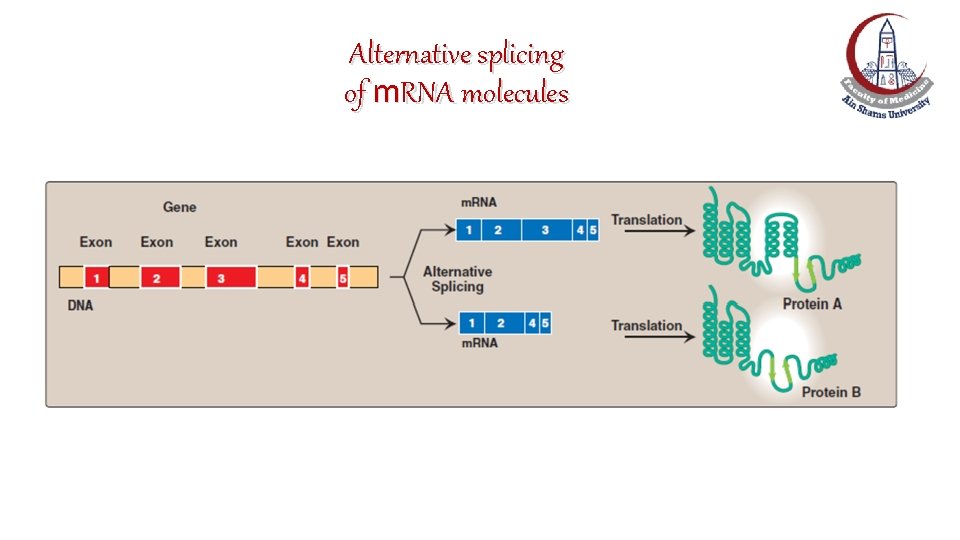
Alternative splicing of m. RNA molecules

Alternative splicing of m. RNA molecules: e. g. , Tropomyosin m. RNA • For example, different types of muscle cells all produce the same primary transcript from the tropomyosin gene. • However, different patterns of splicing in the different cell types produce seven tissue-specific tropomyosin protein molecules (isoforms).

The Editing of m. RNA: Modifications for m. RNA after it is Fully Processed, e. g. , Apo B m. RNA (4563 codons) is expressed in both hepatocytes and small intestinal cells. Two different-size proteins are produced that are important components of chylomicrons and very low density lipoproteins (VLDLs): Ø Apo B-100 protein of VLDLs in the liver (full length protein of 512 k. Da). Ø Apo B-48 protein of chylomicrons in the small intestine (shorter protein of 250 k. Da, only 48% of apo B-100 protein).

The Editing of m. RNA: Apo B m. RNA Apo B gene, its primary m. RNA transcript, and the posttranscriptionally-modified apo-B m. RNA are the same in both hepatocytes and intestinal cells. The editing of fully processed m. RNA occurs only in the intestinal cells by the deamination of cytosine (C) into uracil (U) in the codon number 2153 so that CAA (glutamine) UAA (stop codon). A shorter apo B-48 protein product is produced which represents only 48% of the full length apo B 100 protein. Apo B-48 m. RNA editing in Small intestine

The m. RNA Stability and Translation Repression (si. RNA/mi. RNA) a) m. RNA degradation (in case of perfect base-pair match between si. RNA and m. RNA) b) inhibition (repression) of m. RNA translation (in case of imperfect base-pair match between si. RNA & m. RNA)

Different Species of Eukaryotic RNAs can be classified into protein-coding m. RNA and non-protein coding or simply non-coding (nc) RNA. The nc. RNA can be further subcategorized based on the size into large and small nc. RNAs. Both long nc. RNA (lnc. RNA) & short interference/micro. RNA (si. RNA/mi. RNA) contribute to the control of gene expression.

Different Types of Eukaryotic RNAs Protein coding RNA species Size Abundance Messenger RNA (m. RNA) Heterogeneous 2 -5% Ribosomal r. RNA (r. RNA) 28 S, 18 S, 5. 8 S & 5 S 80% Long noncoding RNA (lnc. RNA) >200 bases 1 -2% Transfer RNA (t. RNA) 4 S 15% Small nuclear RNA (sn. RNA) 150 bases <1% Silencing/micro RNA si. RNA/mi. RNA 22 -24 bases <1% Protein non-coding nc. RNA Large nc. RNA Small nc. RNA

Regulation of Gene Expression at m. RNA Stability Level: RNA interference (RNAi) RNAi is triggered by ds. RNA in eukaryotic cells: 1. Recognition of ds. RNA by endonuclease (Dicer) that is cleaved into short interference RNA (si. RNA) of 21 -24 nucleotides 2. Formation of RNA-induced silencing complex (RISC) by the association of the antisense strand with proteins (containing Slicer endonuclease) 3. Hybridization of antisense strand to the complementary target m. RNA strand with consequent decrease level of gene expression via: a) m. RNA degradation by Slicer (in case of perfect base-pair match between si. RNA and m. RNA) b) inhibition (repression) of m. RNA translation (in case of imperfect base-pair match between si. RNA and m. RNA)

Applications of RNA interference (RNAi) 1. Natural defense against retroviruses infection, e. g. , HIV 2. RNAi therapy: promising approach with some limitations

3. Regulation of Gene Expression by nc. RNA Trans-acting Mechanisms Small nc. RNA: si. RNA/mi. RNA: See above Large nc. RNA: Long nc. RNA (lnc. RNA): q Control at transcription level q Control at posttranscription level q Control through epigenetic mechanisms

4. Regulation of Gene Expression by DNA-related Mechanisms in Eukaryotes q DNA accessibility by transcription machinery e. g. , chromatin remodeling and DNA methylation q Amount of DNA: DNA copy number or DNA amplification e. g. , drug resistance to methotrexate due to gene amplification q DNA rearrangements e. g. , production of immunoglobulins by B-lymphocytes

Regulation of transcription by epigenetic mechanisms DNA accessibility by transcription machinery Chromatin Remodeling by acetylation/deacetylation DNA methylation on cytosine of Cp. G dinucleotide at the promoter region Epigenetic changes: Control of gene expression by other mechanisms than regulatory DNA sequences.
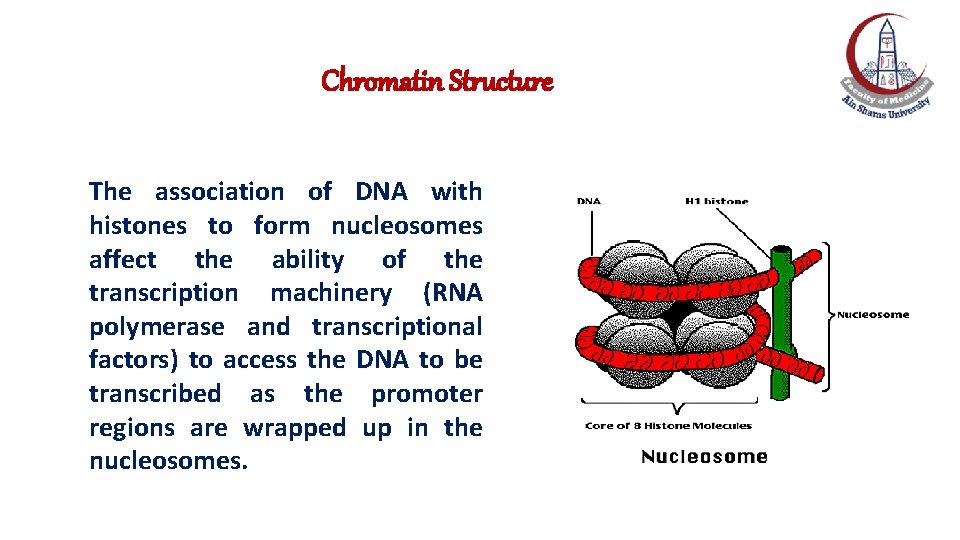
Chromatin Structure The association of DNA with histones to form nucleosomes affect the ability of the transcription machinery (RNA polymerase and transcriptional factors) to access the DNA to be transcribed as the promoter regions are wrapped up in the nucleosomes.

Chromatin Remodeling • The interconversion between active and inactive forms of chromatin is called chromatin remodeling. • Most actively transcribed genes are found in a relatively relaxed form of chromatin called euchromatin (without nucleosomes), whereas most inactive segments of DNA are found in highly condensed heterochromatin (with nucleosomes).

Chromatin remodeling by Histone Acetylation/deacetylation • Acetylation of lysine residues at the amino terminus of histone proteins by acetyl transferase. • Acetylation eliminates the positive charge on the lysine and thereby decreases the interaction of the histone with the negatively charged DNA, forming the relaxed euchromatin. • Removal of the acetyl group by histone deacetylase reverses the process and produces the condensed heterochromatin.

Euchromatin and its effect on transcription Following acetylation and nucleosomal removal: The promoter is opened (accessible by transcription machinery) and the transcription is active
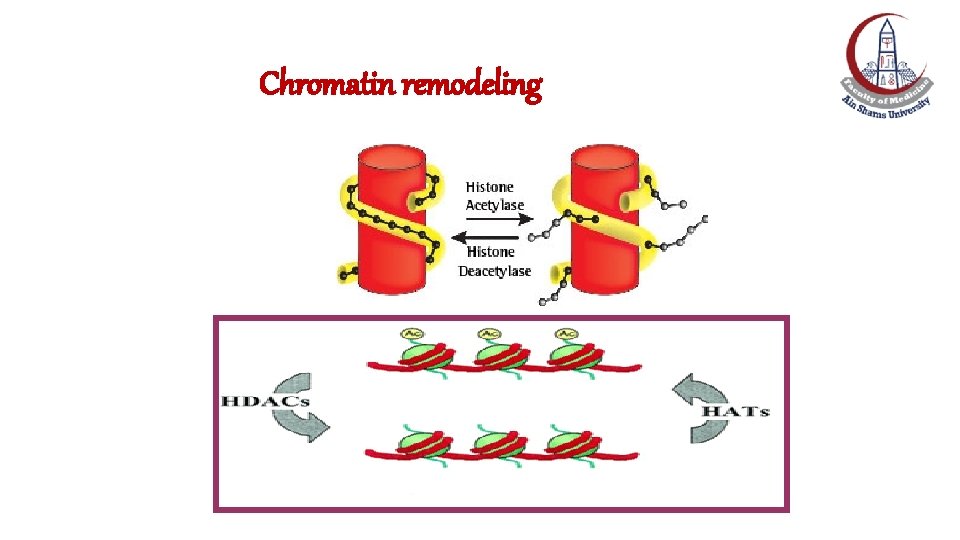
Chromatin remodeling
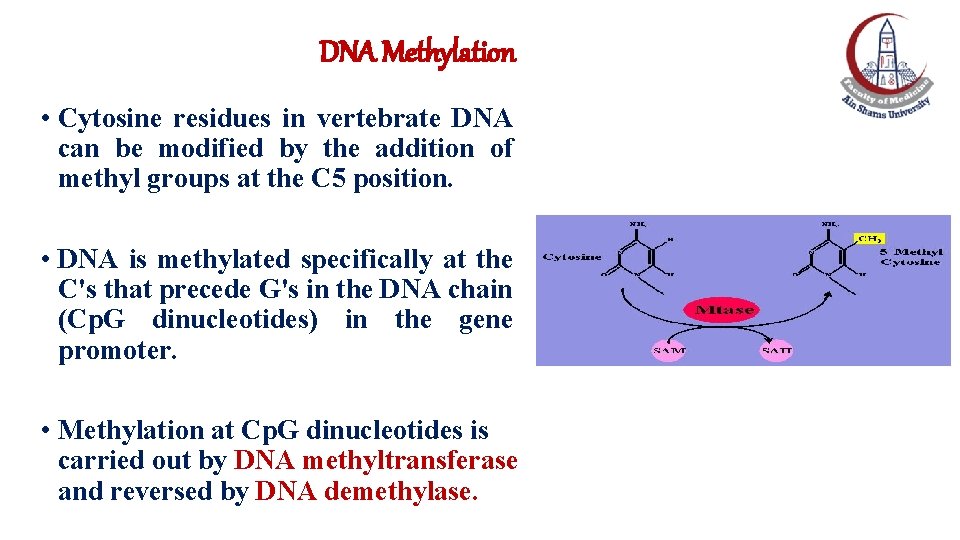
DNA Methylation • Cytosine residues in vertebrate DNA can be modified by the addition of methyl groups at the C 5 position. • DNA is methylated specifically at the C's that precede G's in the DNA chain (Cp. G dinucleotides) in the gene promoter. • Methylation at Cp. G dinucleotides is carried out by DNA methyltransferase and reversed by DNA demethylase.
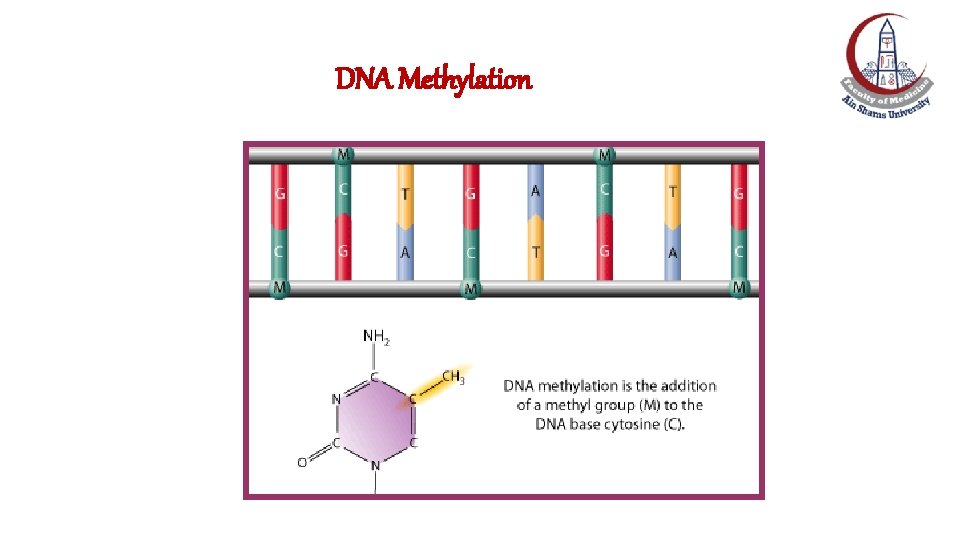
DNA Methylation

Regulation of transcription by DNA Methylation: Mechanisms 1 - The most direct mechanism by which DNA methylation can interfere with transcription is to prevent the binding of basal transcriptional factors with promoter cytosines. 2 - Recruiting histone deacetylase and formation of heterochromatin (inactive)
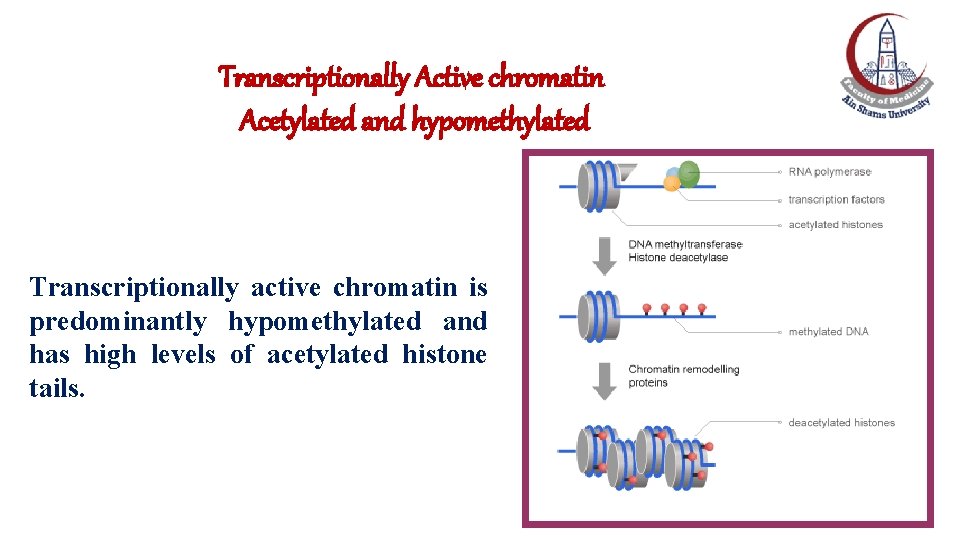
Transcriptionally Active chromatin Acetylated and hypomethylated Transcriptionally active chromatin is predominantly hypomethylated and has high levels of acetylated histone tails.

Regulation of gene expression by the Amount of DNA: DNA copy number or DNA amplification • A change in the number of copies of a gene can affect the amount of its product. • An increase in copy number or gene amplification is a normal developmental process in certain mammalian species.

DNA amplification & Methotrexate DRUg resistance • Methotrexate (Cytotoxic drug): An inhibitor of dihydrofolate reductase (DHFR) that is required for thymidine monophosphate (d. TMP) synthesis. • Amplification of DHFR gene results in increased expression of DHFR and the increased synthesis of d. TMP and consequently the development of drug resistance to methotrexate, i. e. , methotrexate becomes less efficient in treatment of cancer.
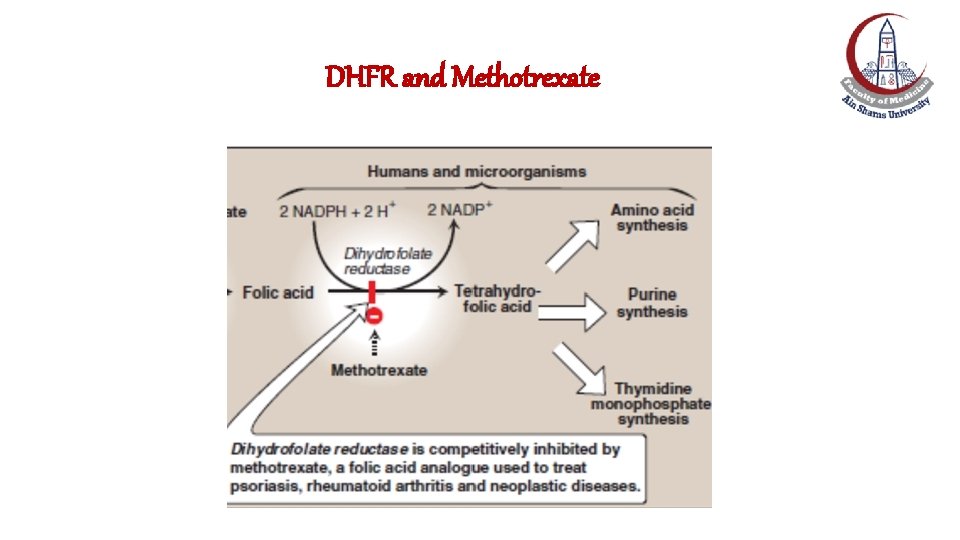
DHFR and Methotrexate
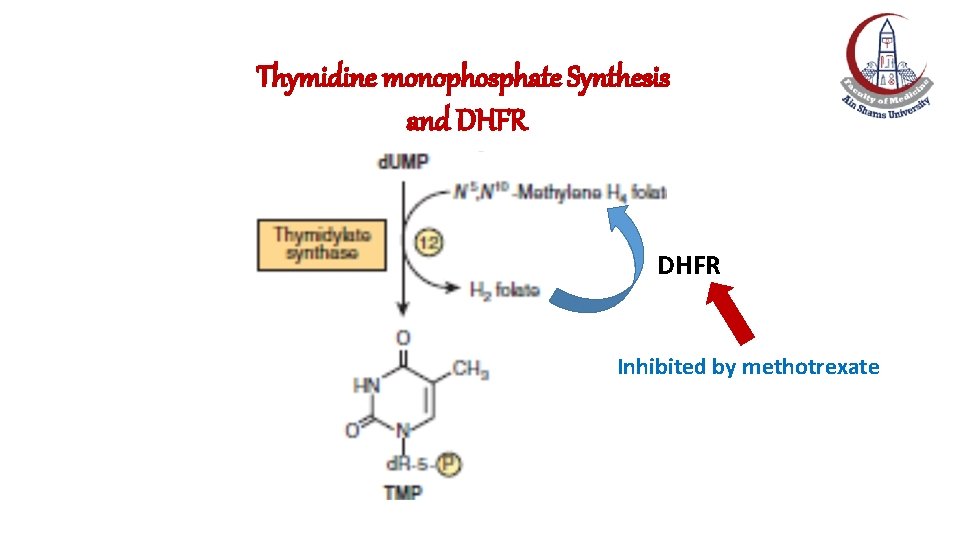
Thymidine monophosphate Synthesis and DHFR Inhibited by methotrexate

Regulation of gene expression by Re-arrangement of DNA e. g. , antibody diversity • Permanent rearrangement of DNA occurs in B-lymphocytes during production of immunoglobulins. • The immunoglobulin consists of heavy and light chains, with each chain containing regions of variable and constant amino acid sequence. • The variable region is the result of somatic recombination of segments (variable, diversity and joining) within both light and heavy chain genes. • DNA arrangements allow generation of 109 -1011 different antibodies from a single gene, providing the diversity of antibodies.
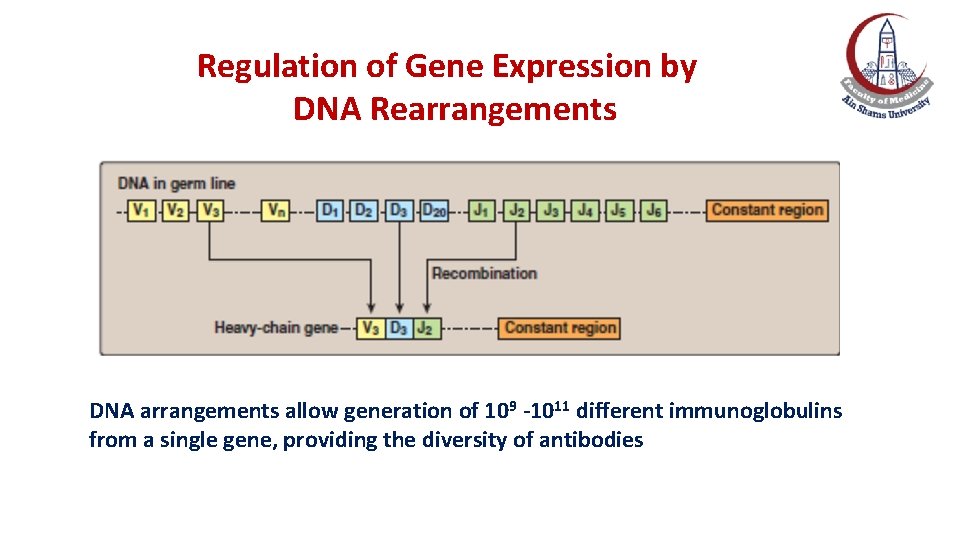
Regulation of Gene Expression by DNA Rearrangements DNA arrangements allow generation of 109 -1011 different immunoglobulins from a single gene, providing the diversity of antibodies

- Slides: 51