Regulation of gene expression Haixu Tang School of






- Slides: 74

Regulation of gene expression Haixu Tang School of Informatics


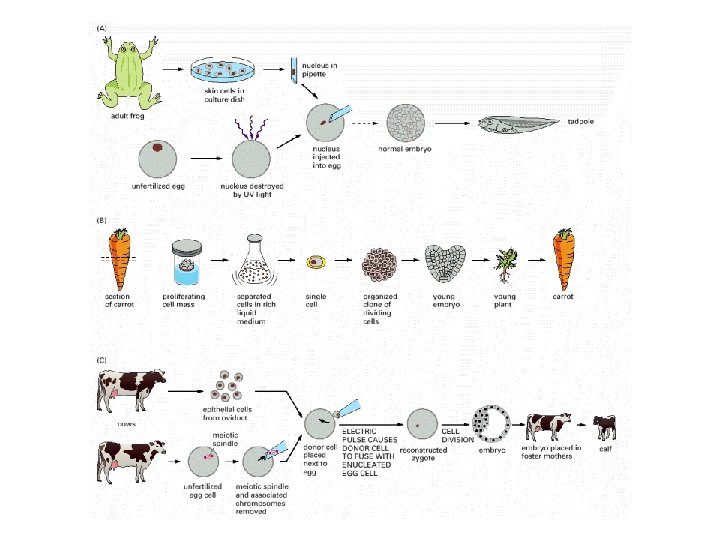
Genetic material are not lost

Different Cell Types Synthesize Different Sets of Proteins • Many processes are common to all cells, and any two cells in a single organism therefore have many proteins in common. • Some proteins are abundant in the specialized cells in which they function and cannot be detected elsewhere, even by sensitive tests. Hemoglobin, for example, can be detected only in red blood cells. • Studies of the number of different m. RNAs suggest that, at any one time, a typical human cell expresses approximately 10, 000 20, 000 of its approximately 30, 000 genes • Although the differences in m. RNAs among specialized cell types are striking, they nonetheless underestimate the full range of differences

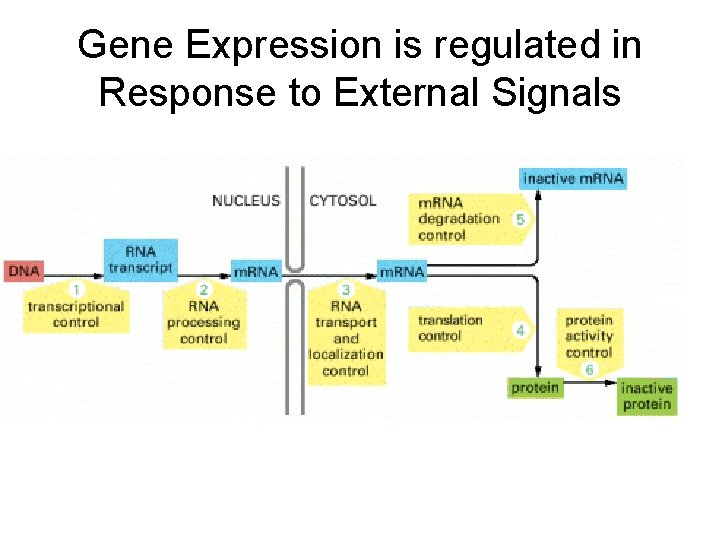
Gene Expression is regulated in Response to External Signals

Switching devices for gene regulation • Short stretches of DNA of defined sequence (cis-elements) • gene regulatory proteins that recognize and bind to them (trans-factors)


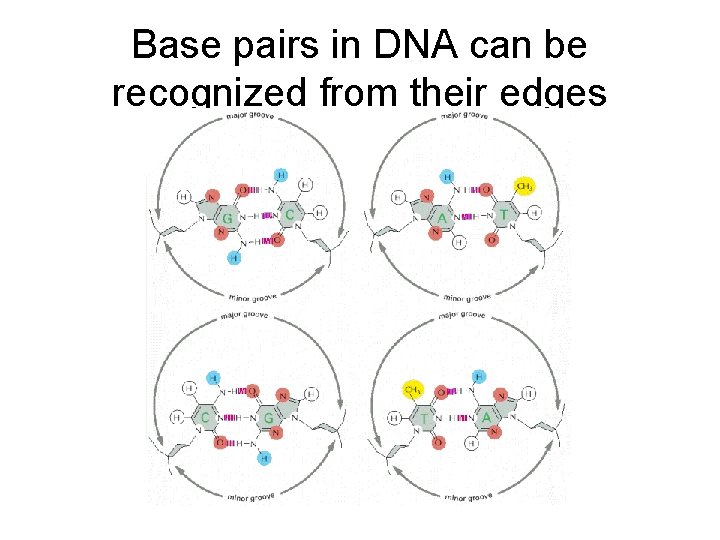
Base pairs in DNA can be recognized from their edges

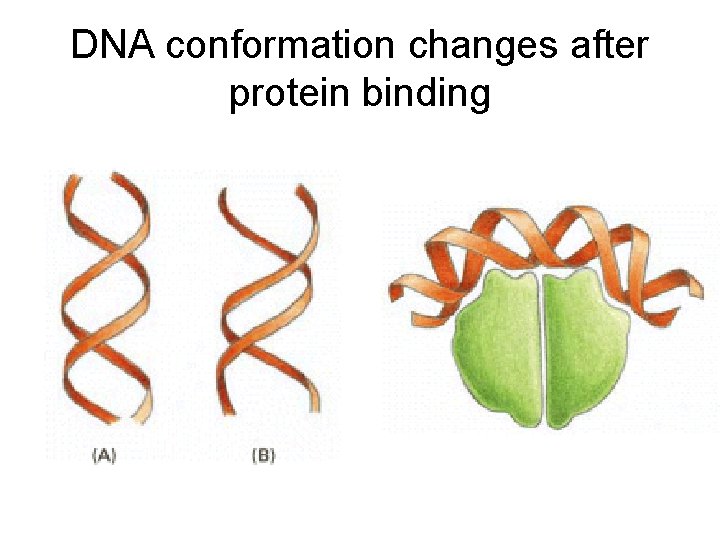
DNA conformation changes after protein binding

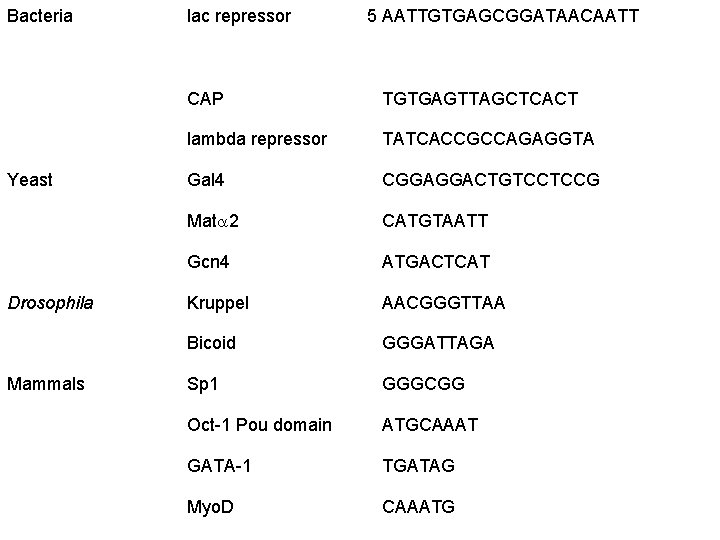
Bacteria Yeast Drosophila Mammals lac repressor 5 AATTGTGAGCGGATAACAATT CAP TGTGAGTTAGCTCACT lambda repressor TATCACCGCCAGAGGTA Gal 4 CGGAGGACTGTCCTCCG Mata 2 CATGTAATT Gcn 4 ATGACTCAT Kruppel AACGGGTTAA Bicoid GGGATTAGA Sp 1 GGGCGG Oct-1 Pou domain ATGCAAAT GATA-1 TGATAG Myo. D CAAATG

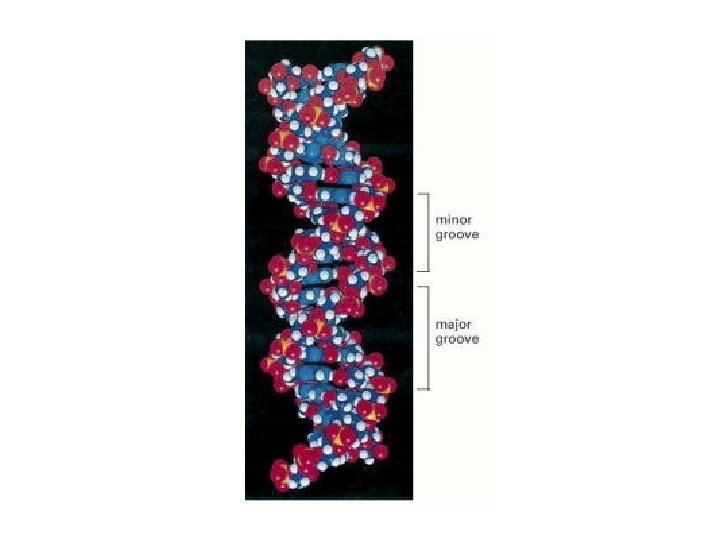
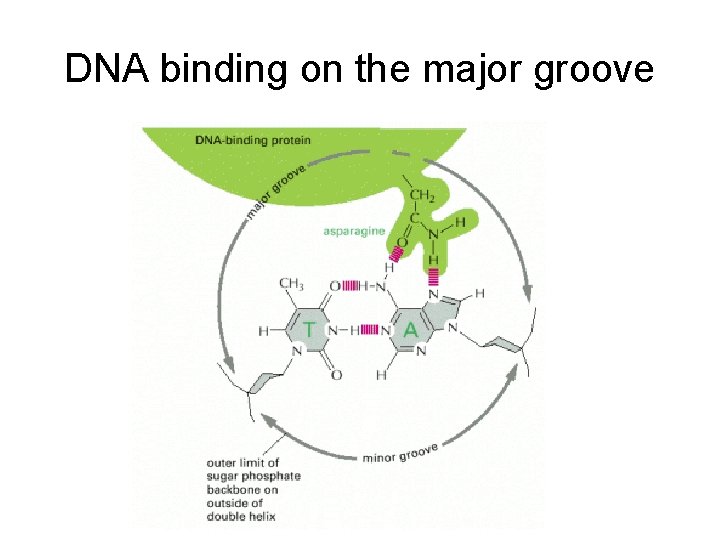
DNA binding on the major groove

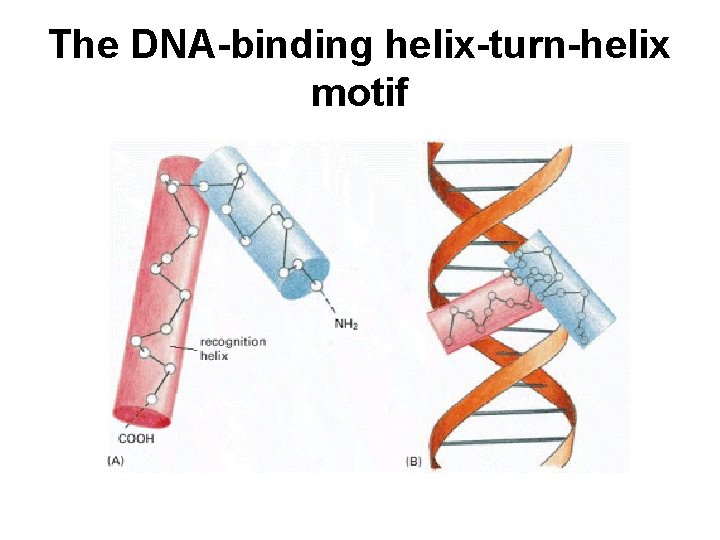
The DNA-binding helix-turn-helix motif

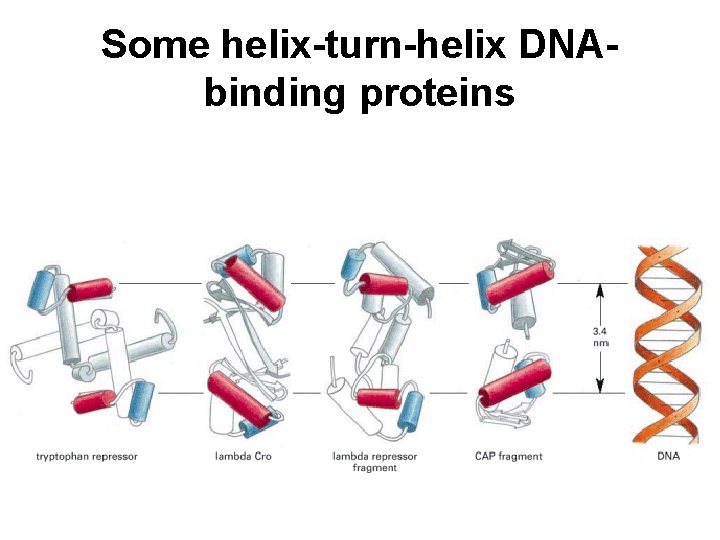
Some helix-turn-helix DNAbinding proteins

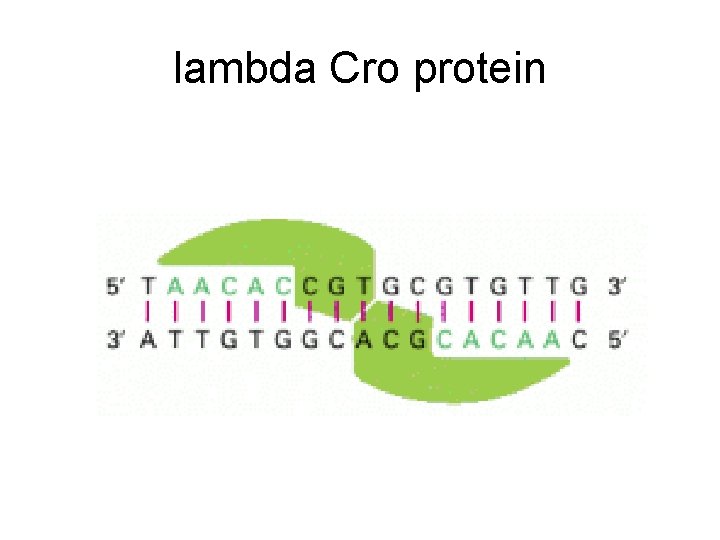
lambda Cro protein

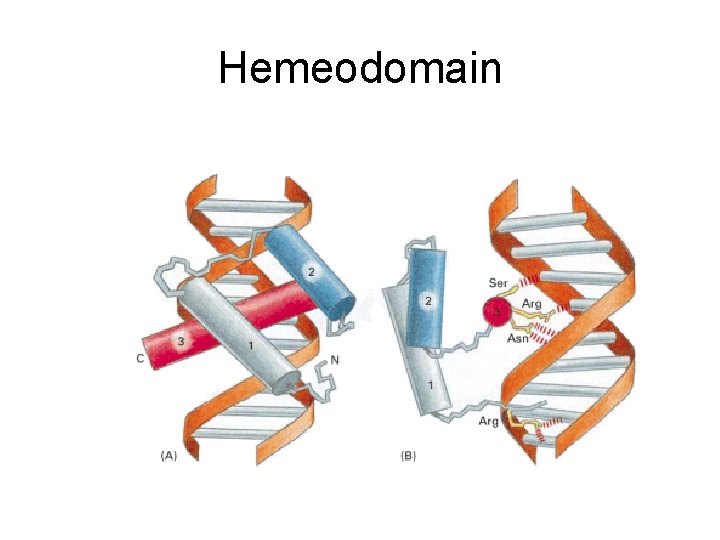
Hemeodomain

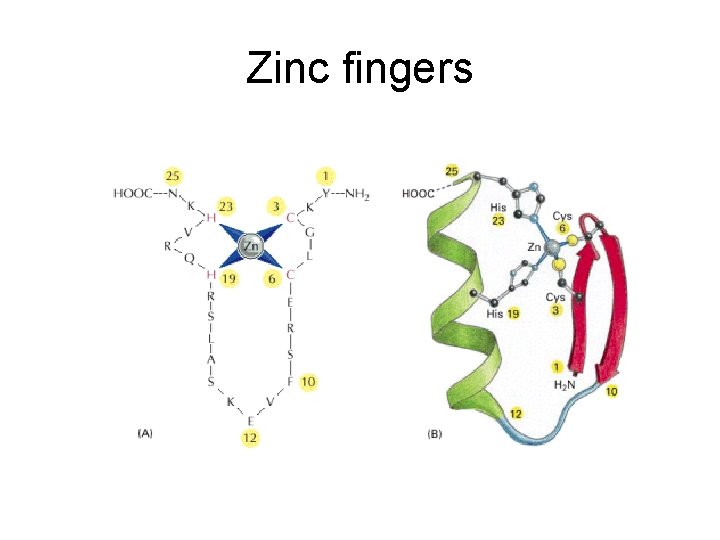
Zinc fingers

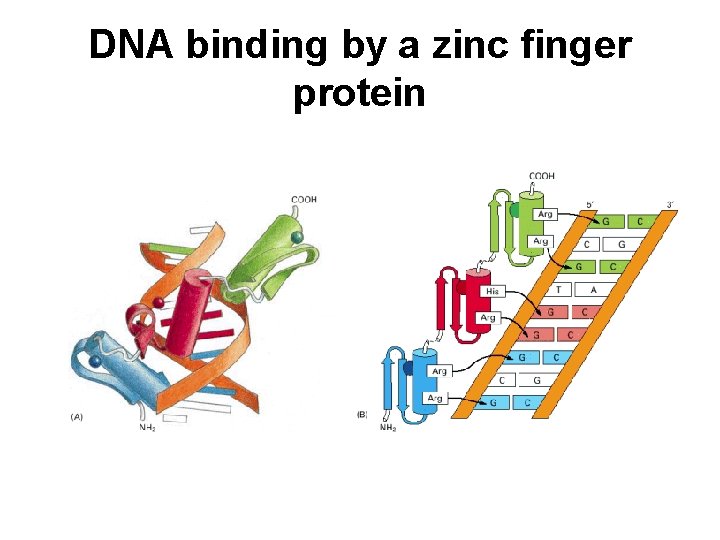
DNA binding by a zinc finger protein

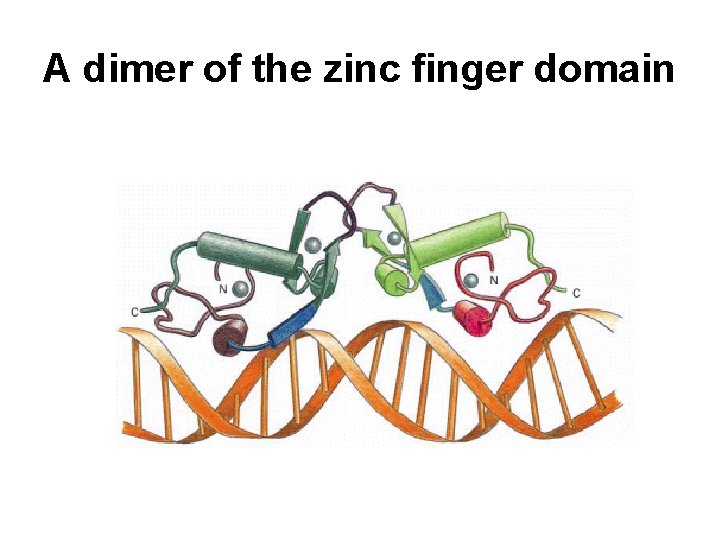
A dimer of the zinc finger domain

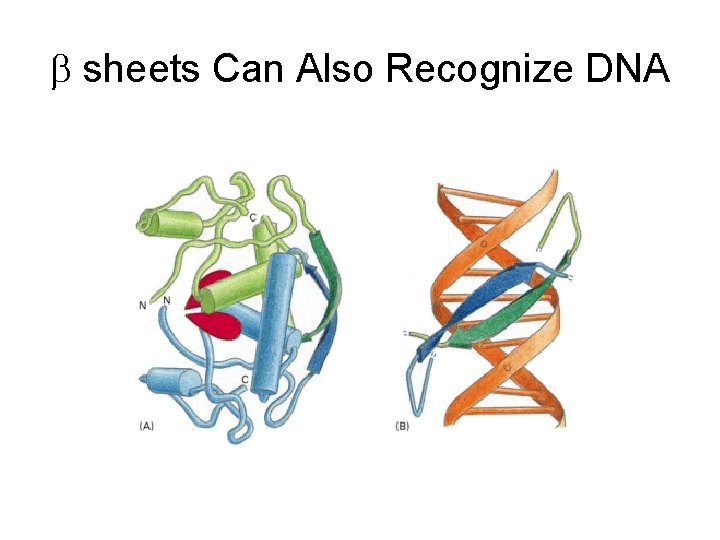
b sheets Can Also Recognize DNA

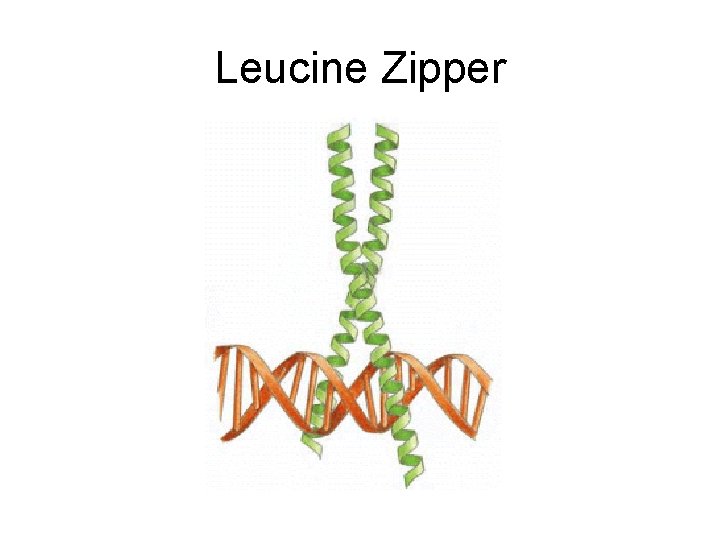
Leucine Zipper

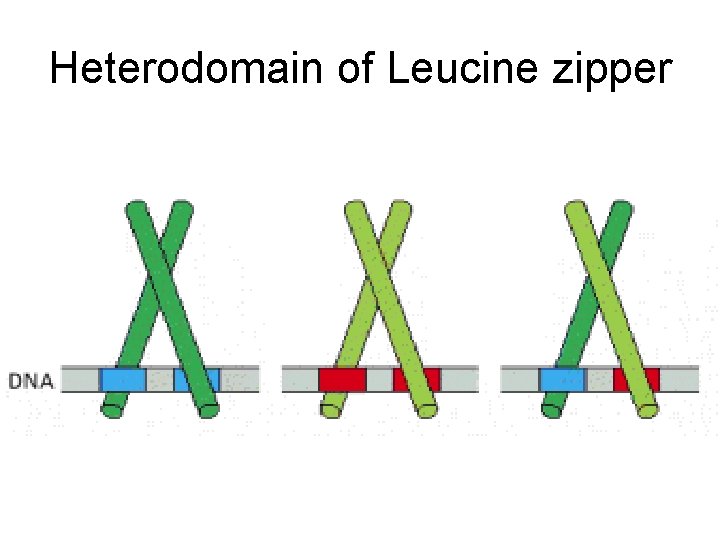
Heterodomain of Leucine zipper

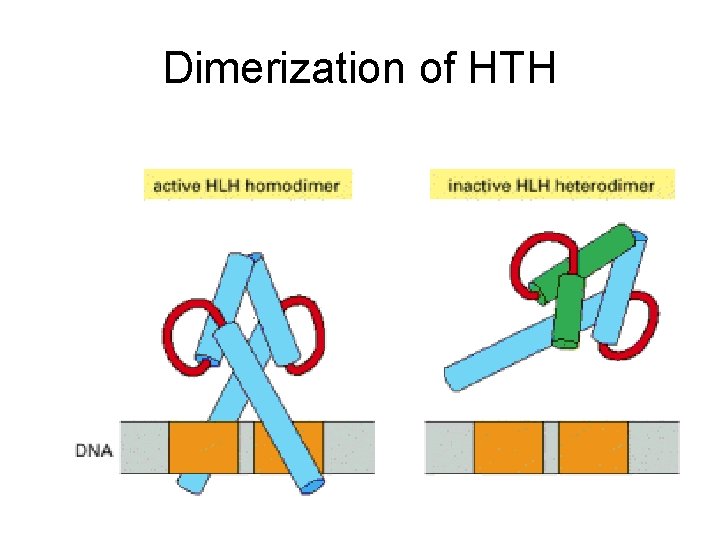
Dimerization of HTH

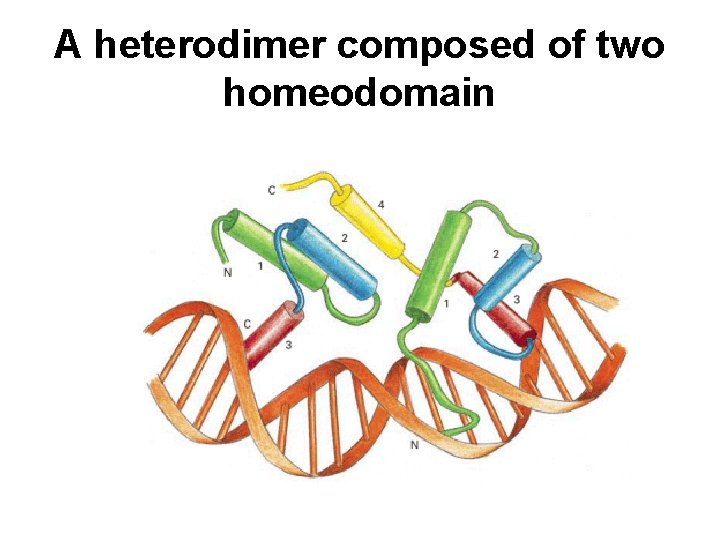
A heterodimer composed of two homeodomain

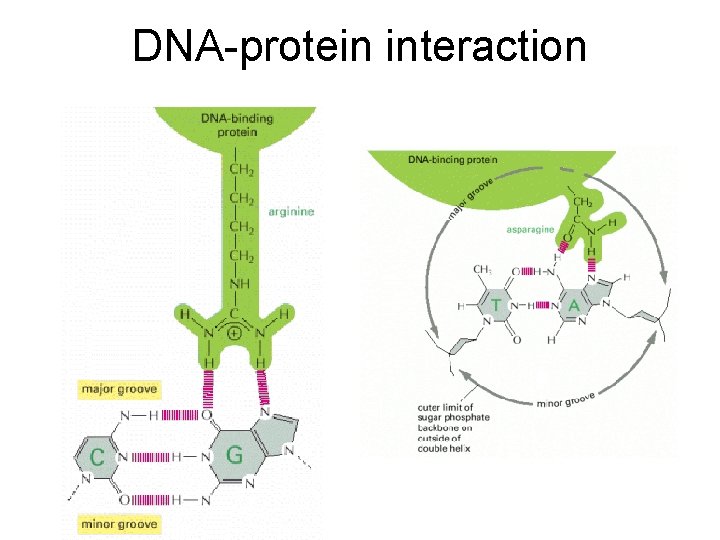
DNA-protein interaction

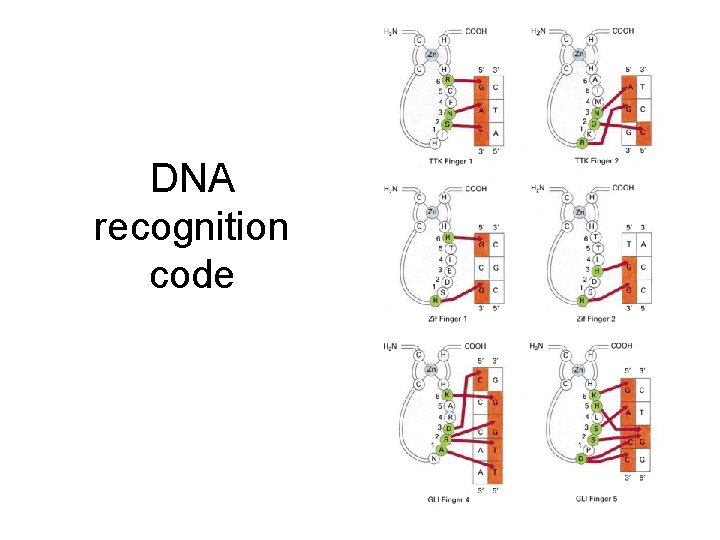
DNA recognition code

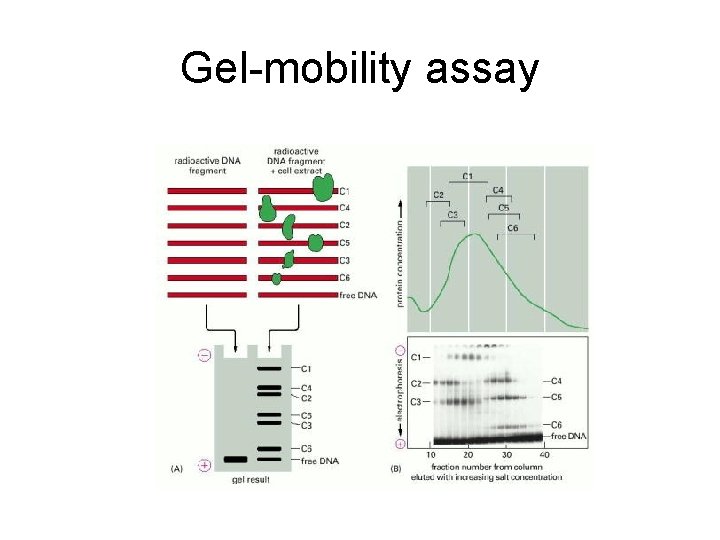
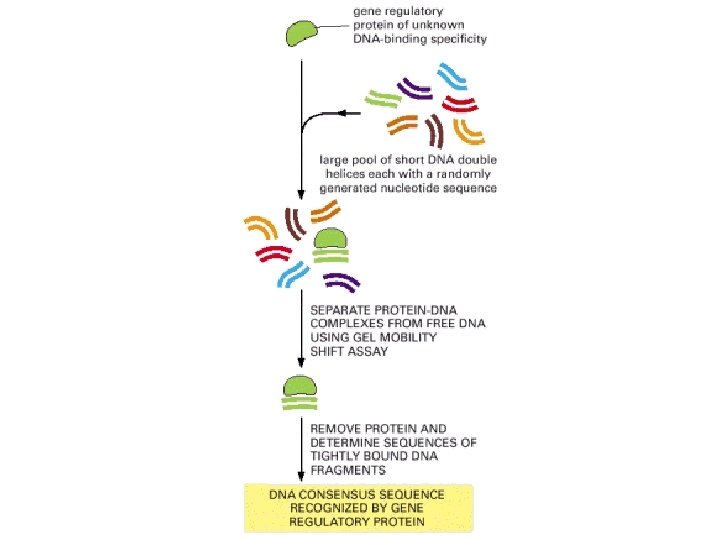
Gel-mobility assay

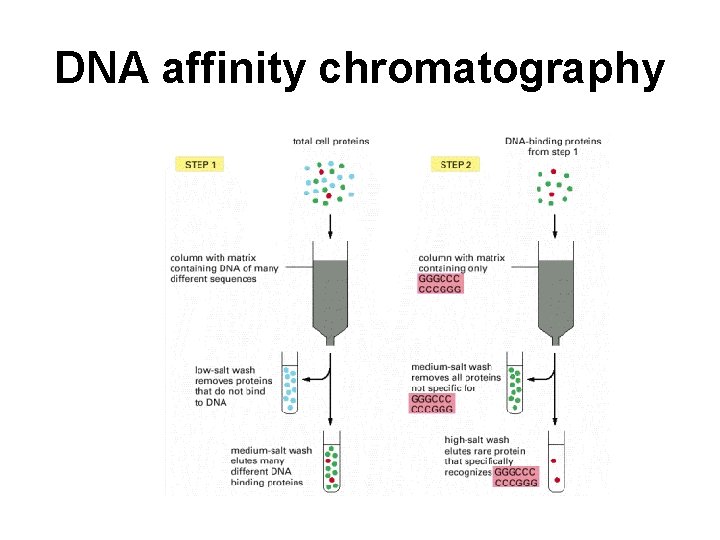
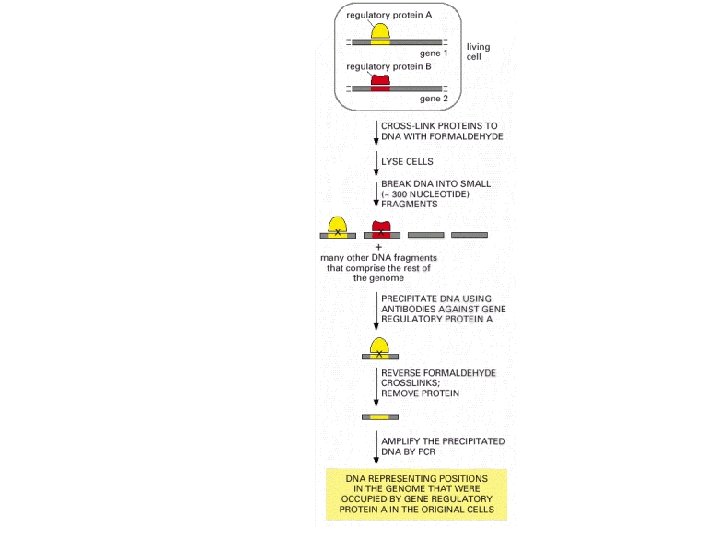
DNA affinity chromatography



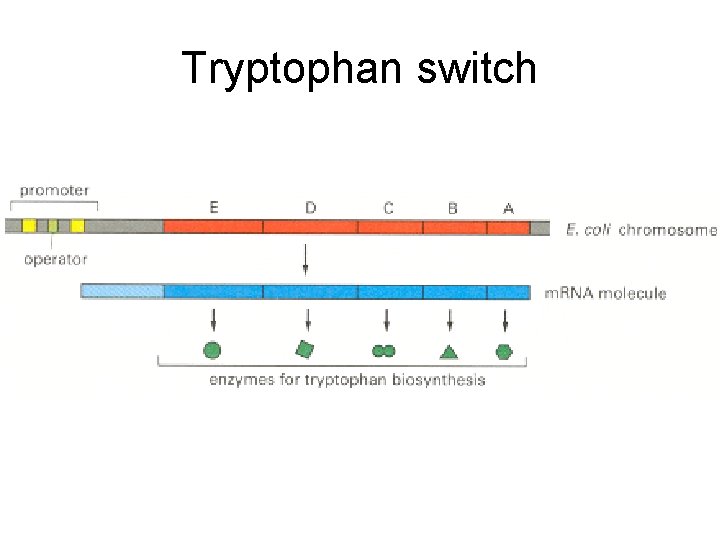
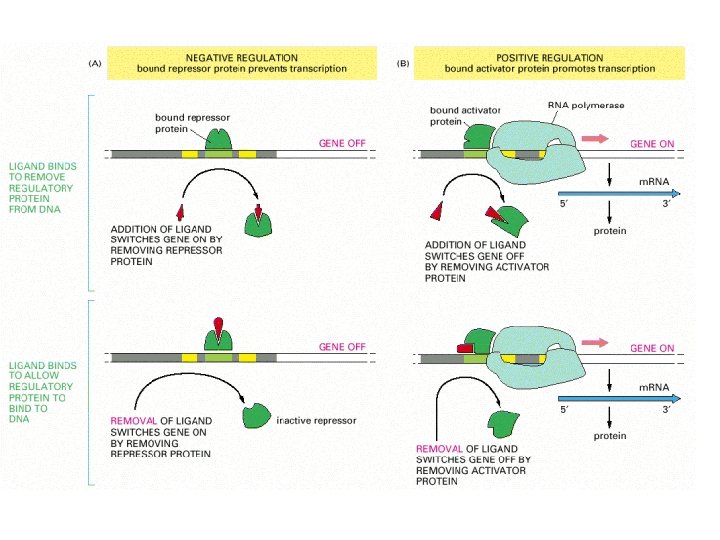
Tryptophan switch

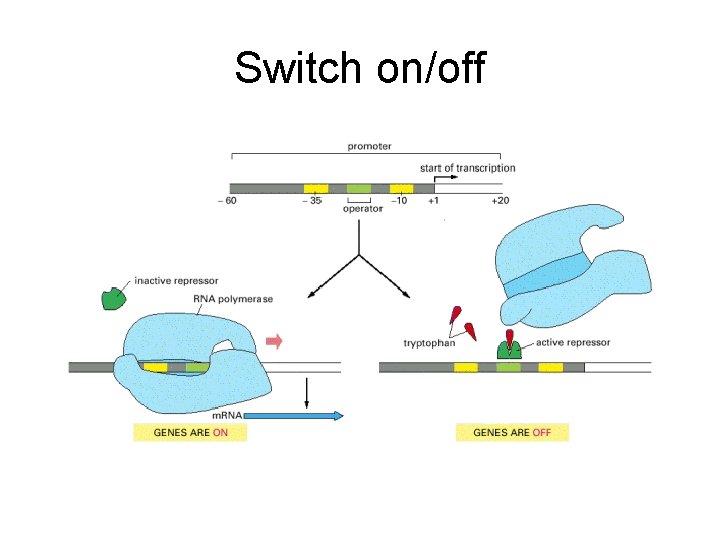
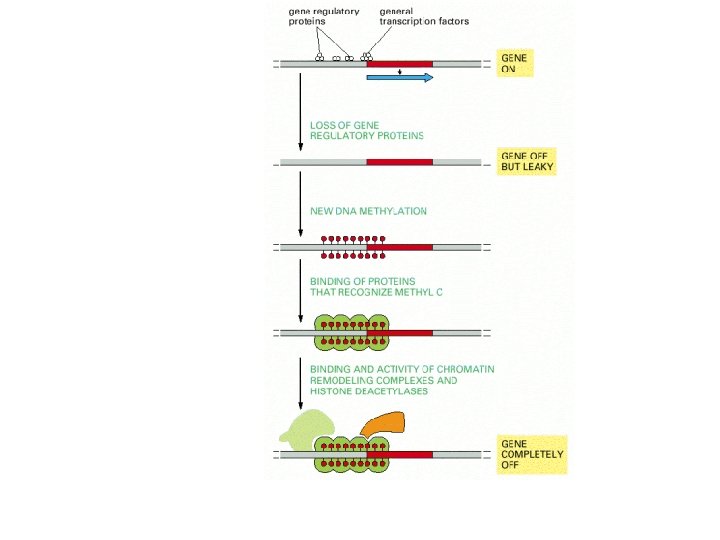
Switch on/off

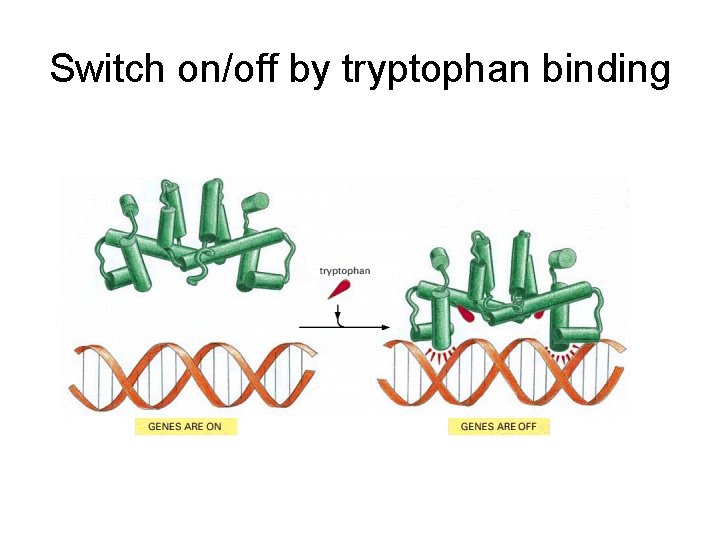
Switch on/off by tryptophan binding


Dual control of the lac operon

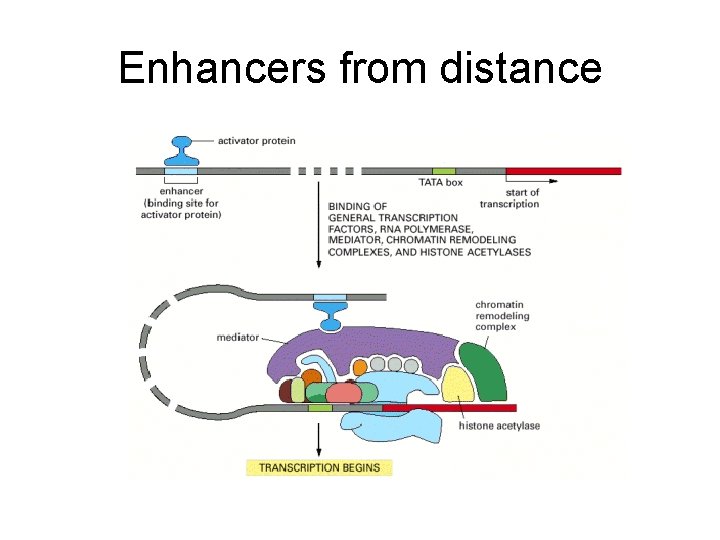
Enhancers from distance

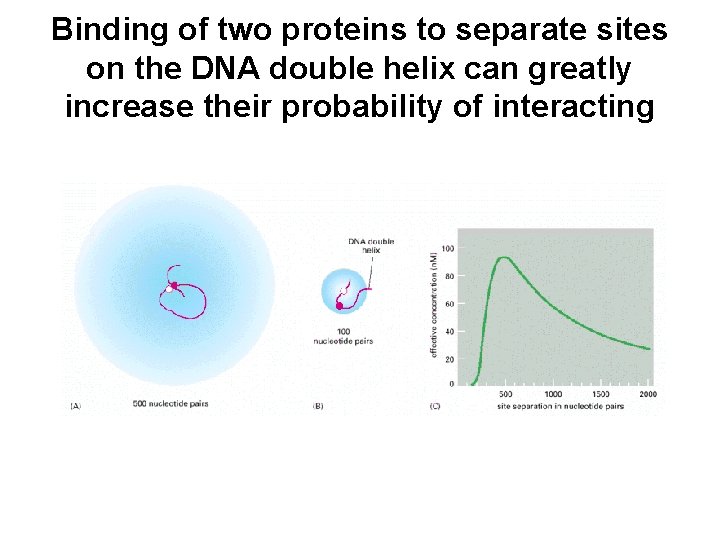
Binding of two proteins to separate sites on the DNA double helix can greatly increase their probability of interacting

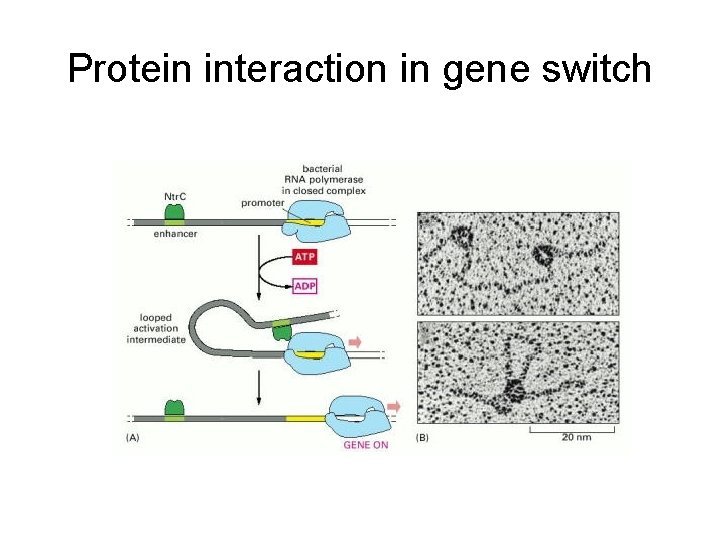
Protein interaction in gene switch

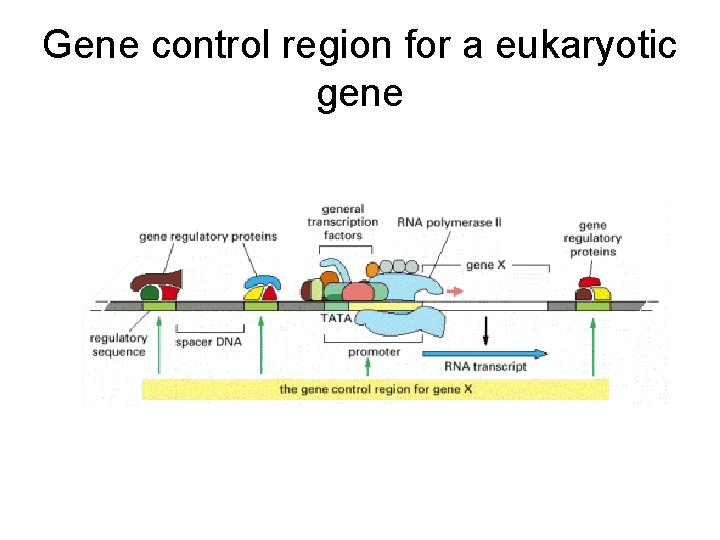
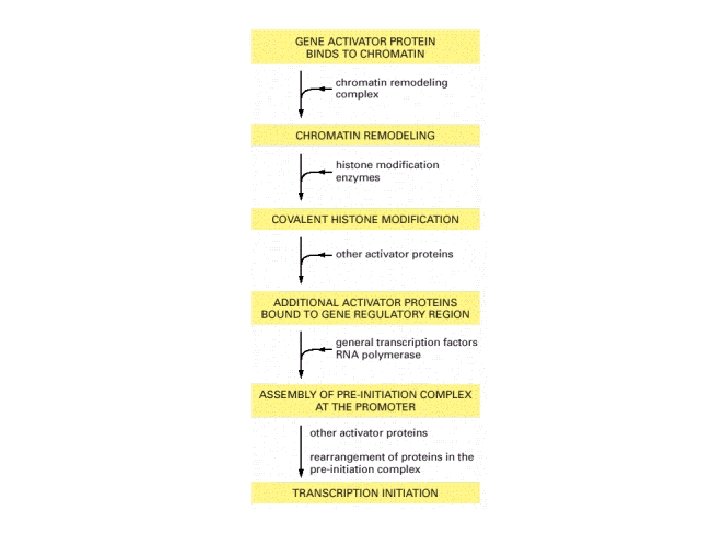
Gene control region for a eukaryotic gene

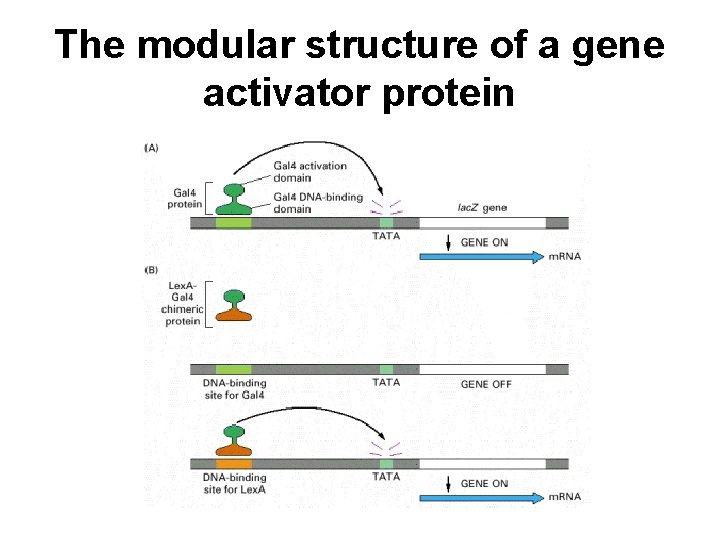
The modular structure of a gene activator protein


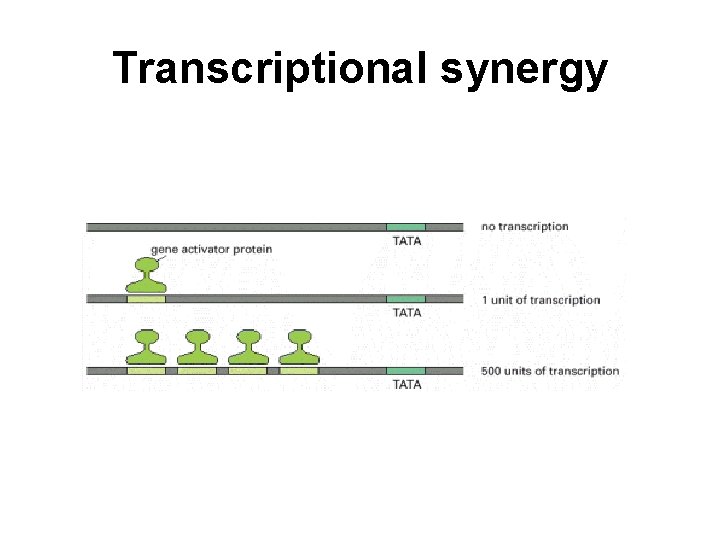
Transcriptional synergy


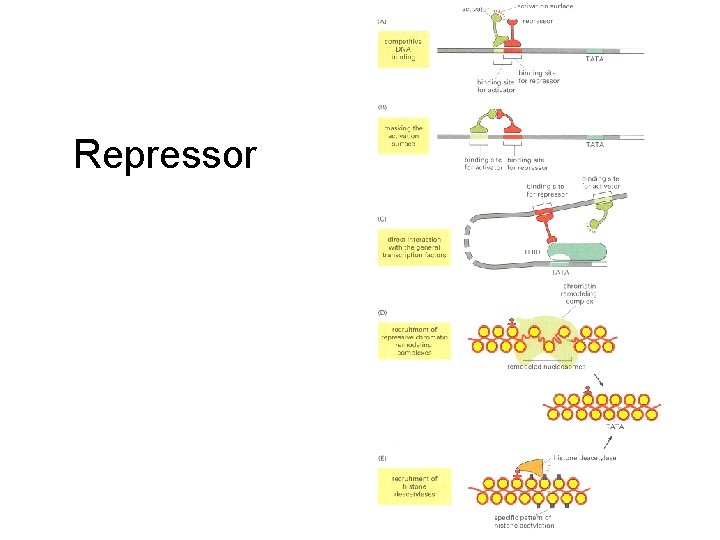
Repressor

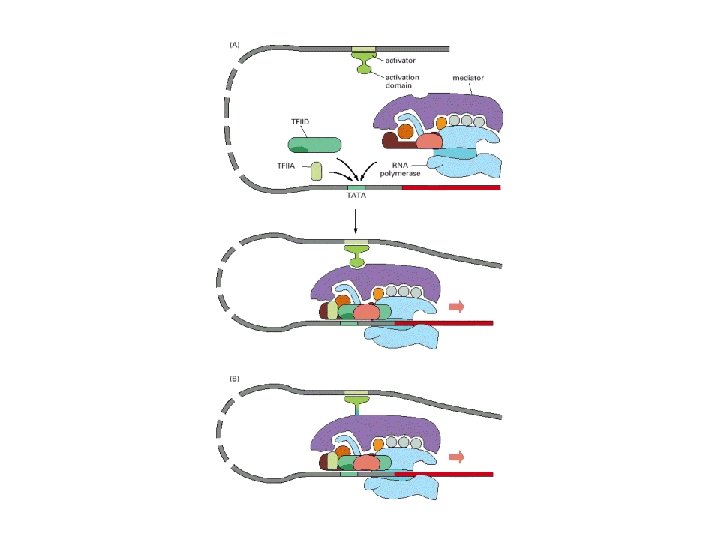
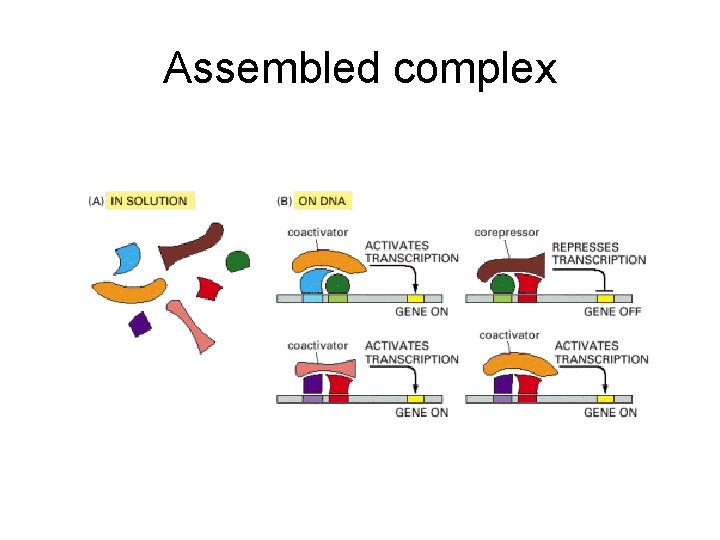
Assembled complex

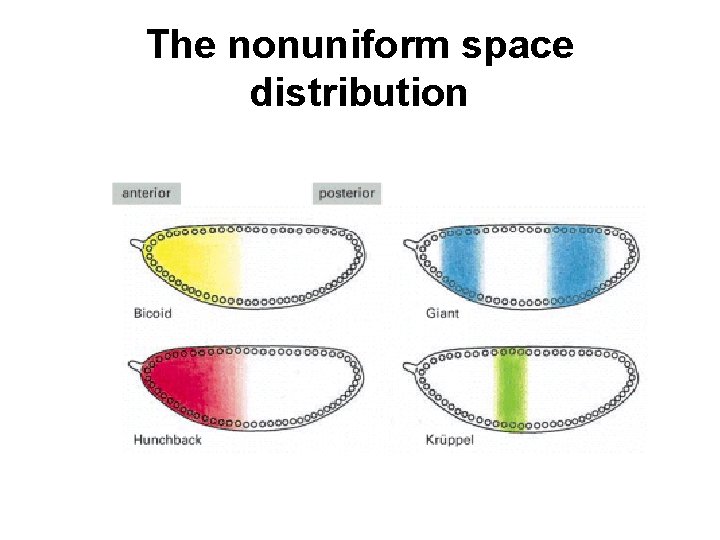
The nonuniform space distribution

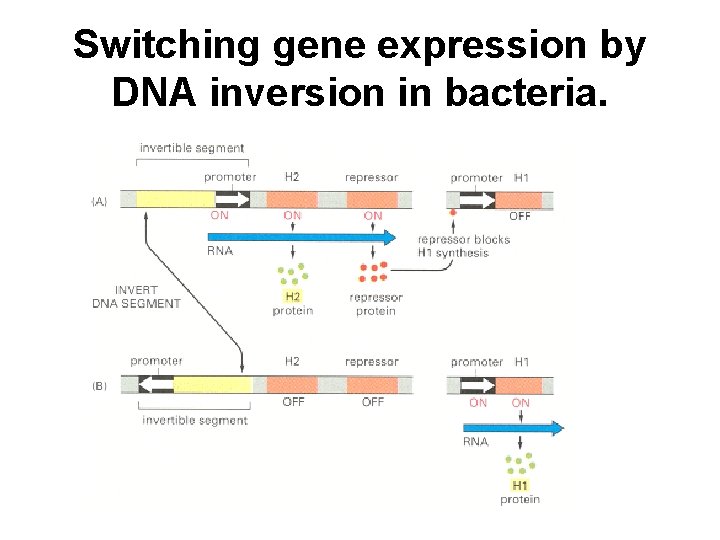
Switching gene expression by DNA inversion in bacteria.

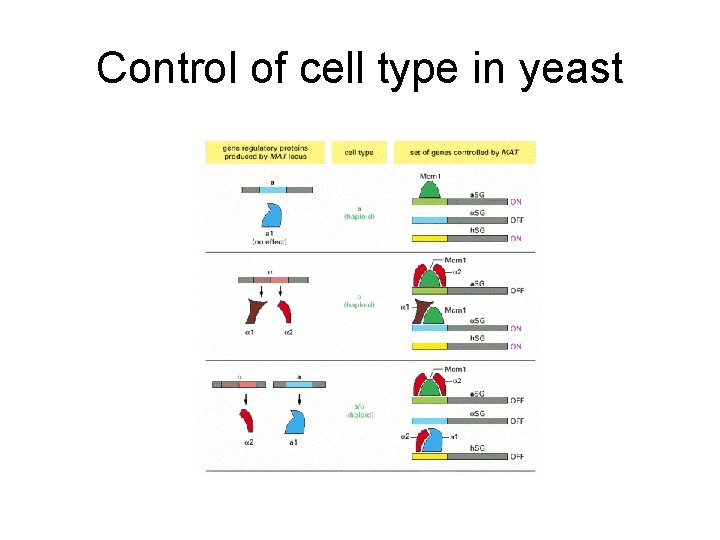
Control of cell type in yeast

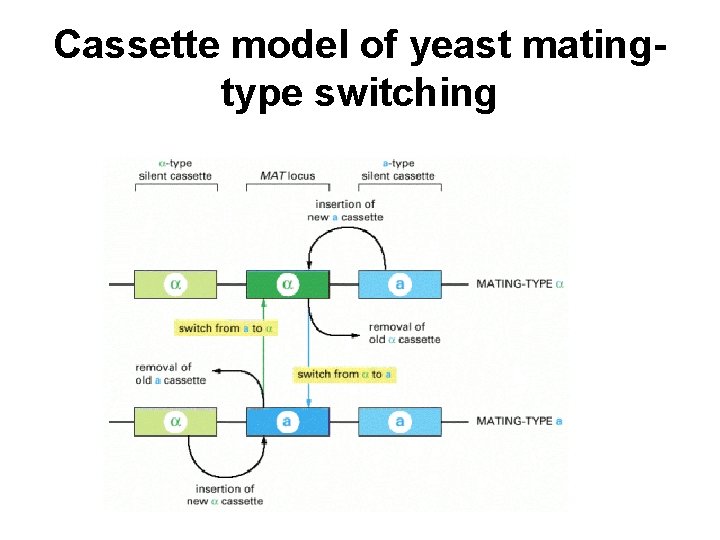
Cassette model of yeast matingtype switching

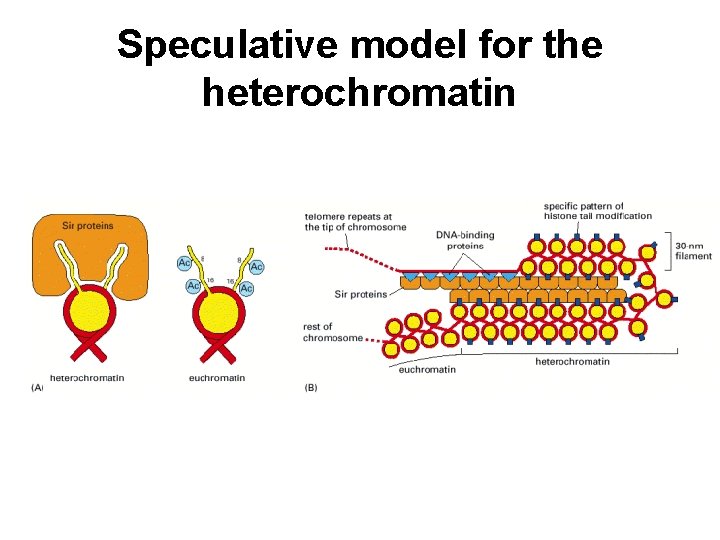
Speculative model for the heterochromatin

A positive feedback loop

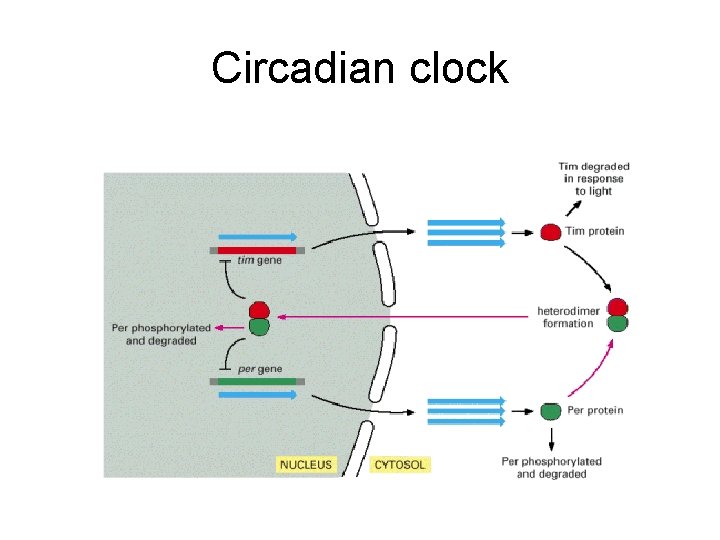
Circadian clock

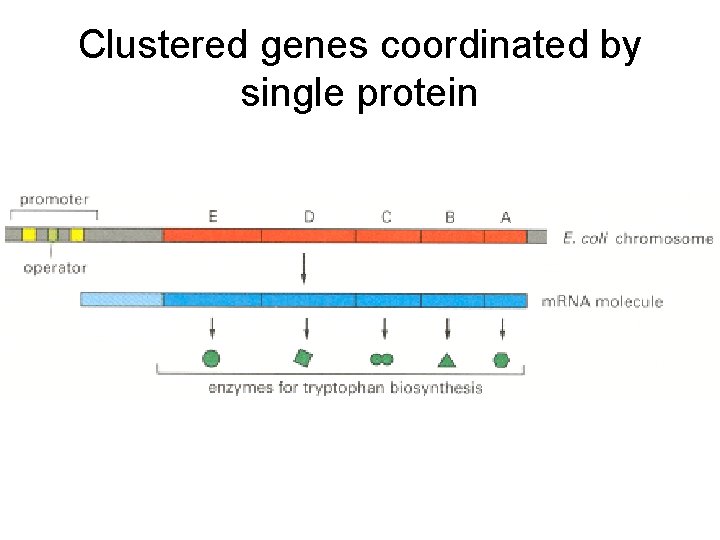
Clustered genes coordinated by single protein

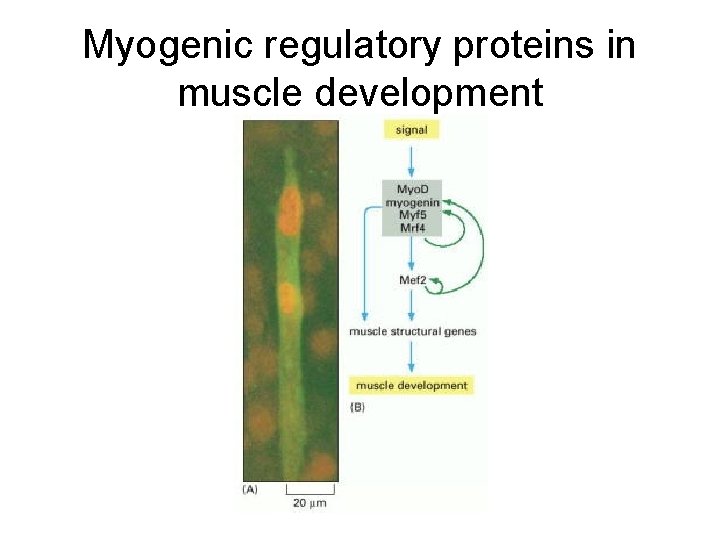
Myogenic regulatory proteins in muscle development

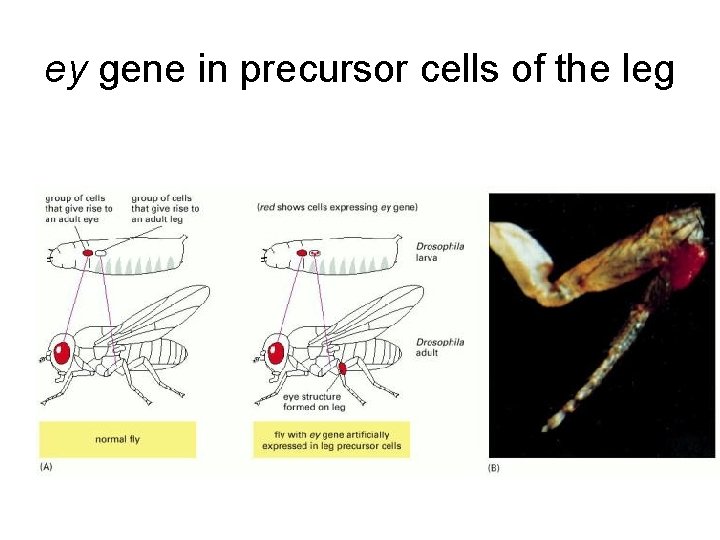
ey gene in precursor cells of the leg

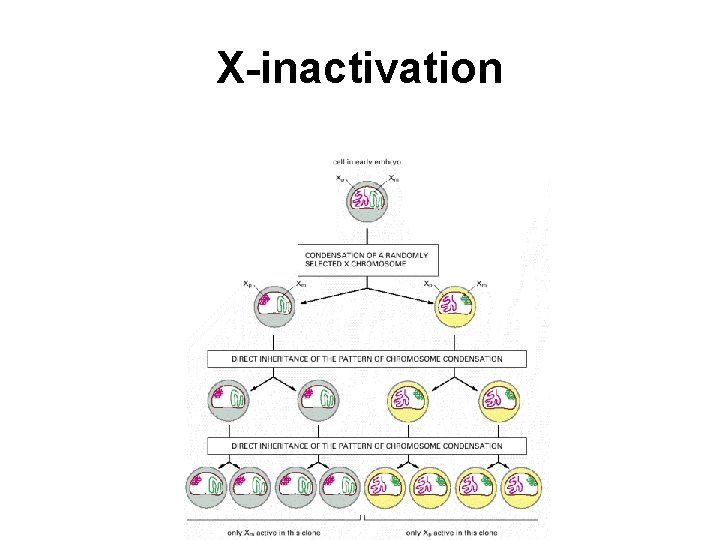
X-inactivation

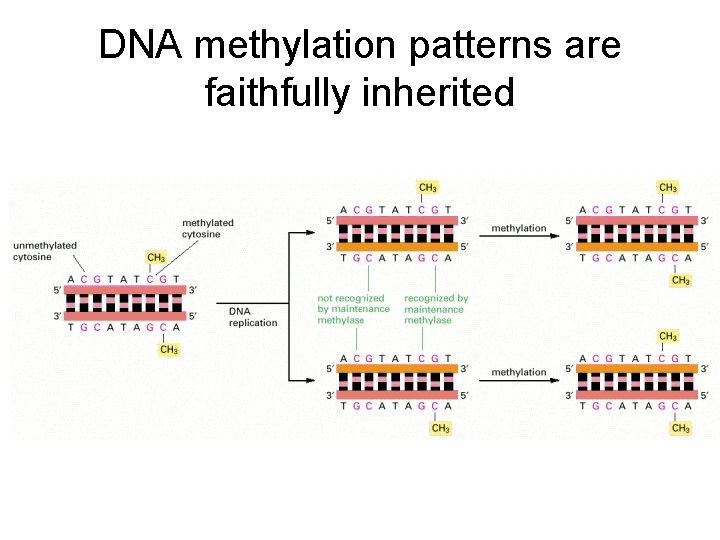
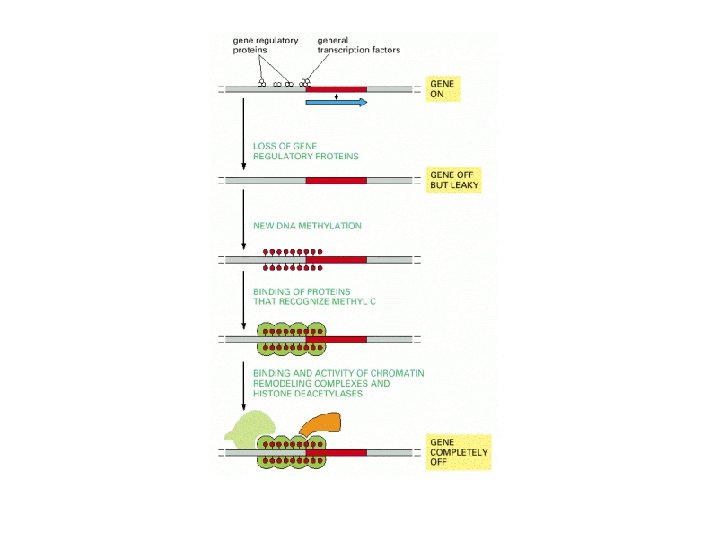
DNA methylation patterns are faithfully inherited


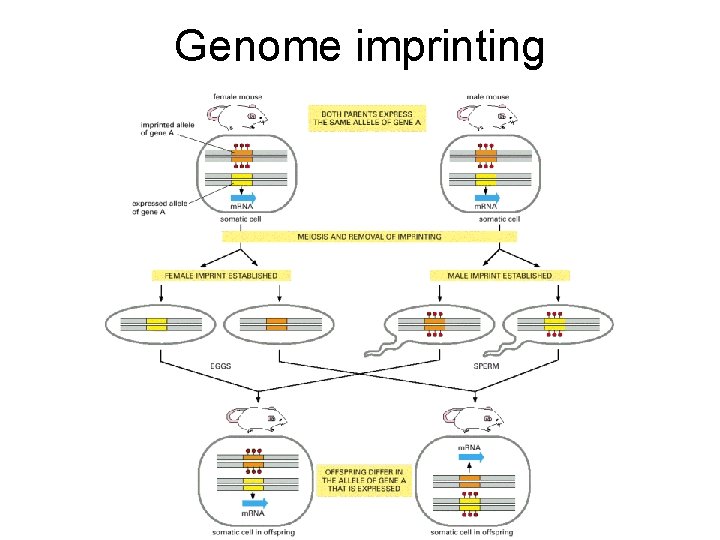
Genome imprinting

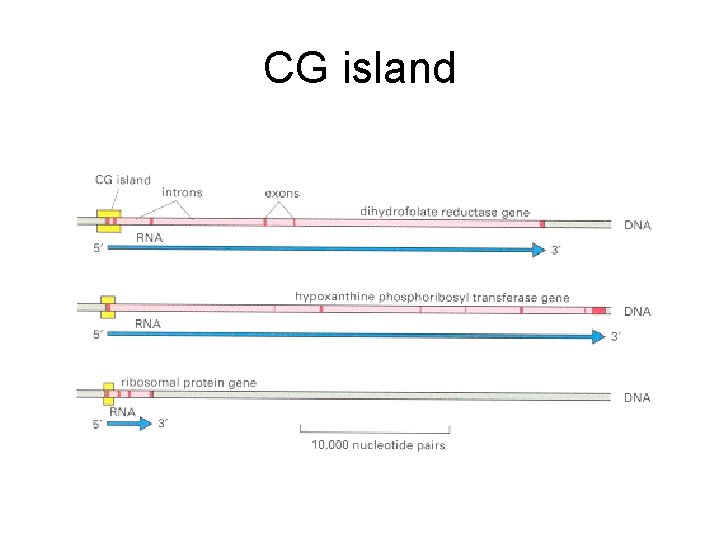
CG island


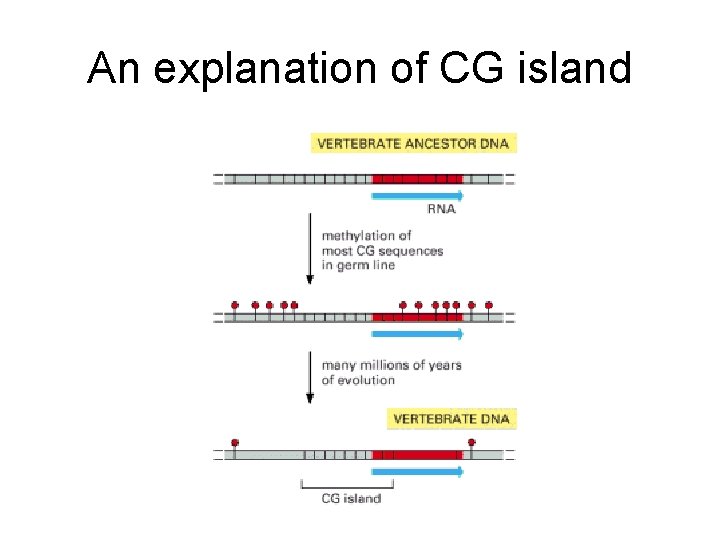
An explanation of CG island

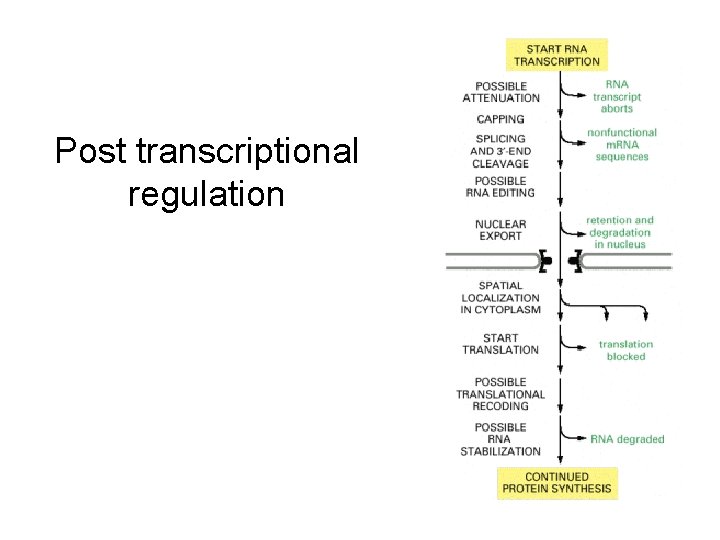
Post transcriptional regulation

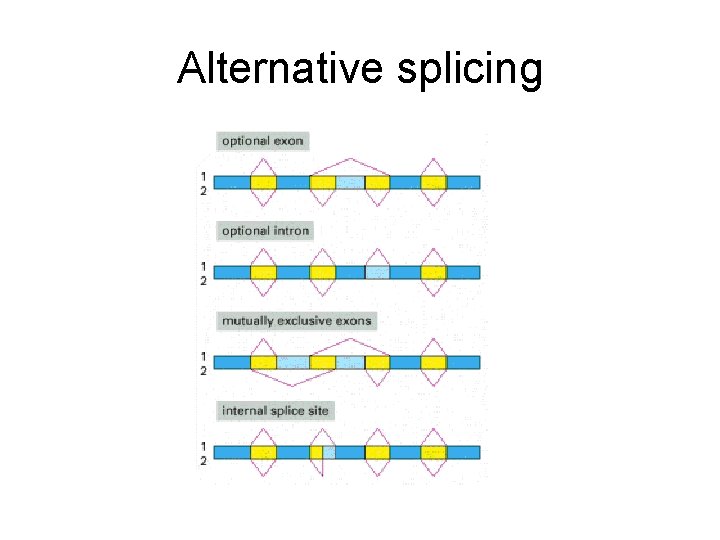
Alternative splicing

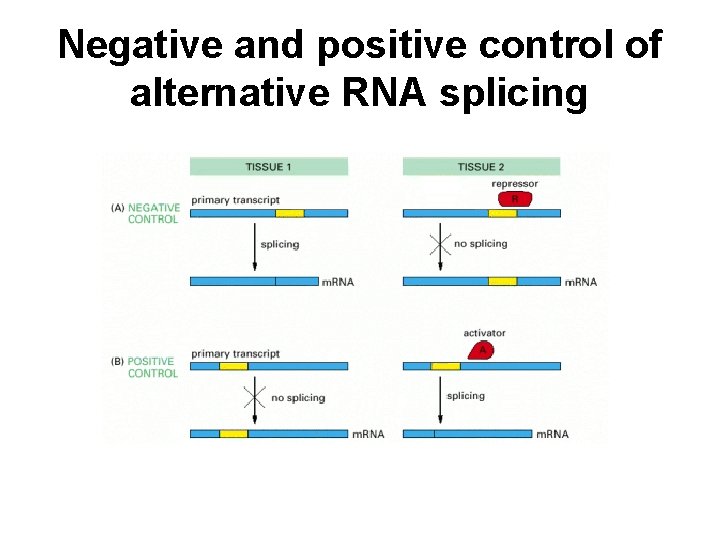
Negative and positive control of alternative RNA splicing

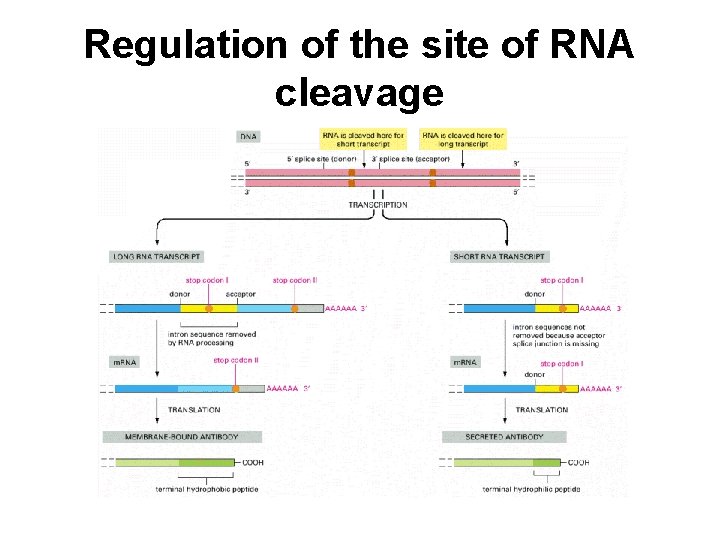
Regulation of the site of RNA cleavage

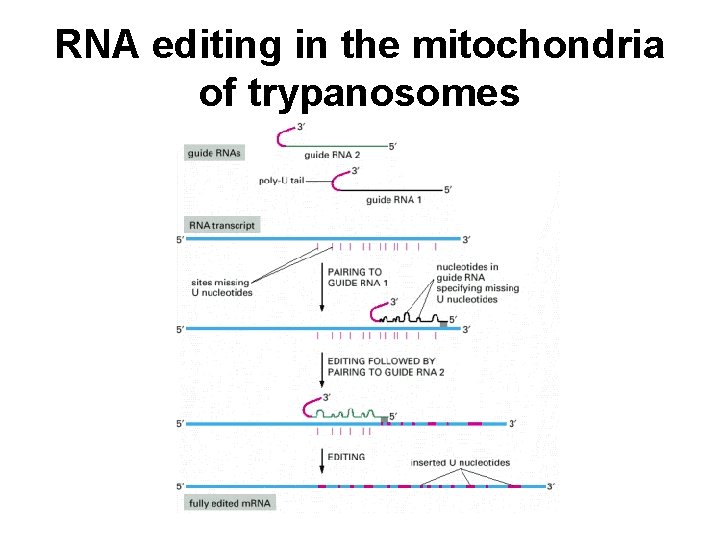
RNA editing in the mitochondria of trypanosomes

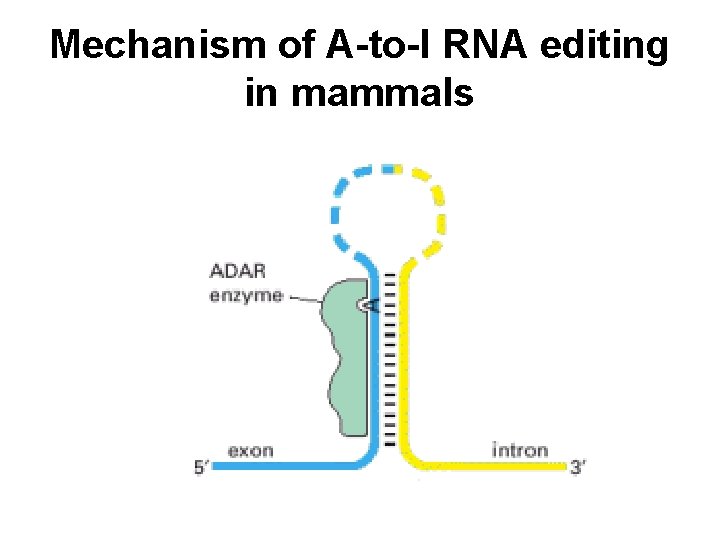
Mechanism of A-to-I RNA editing in mammals

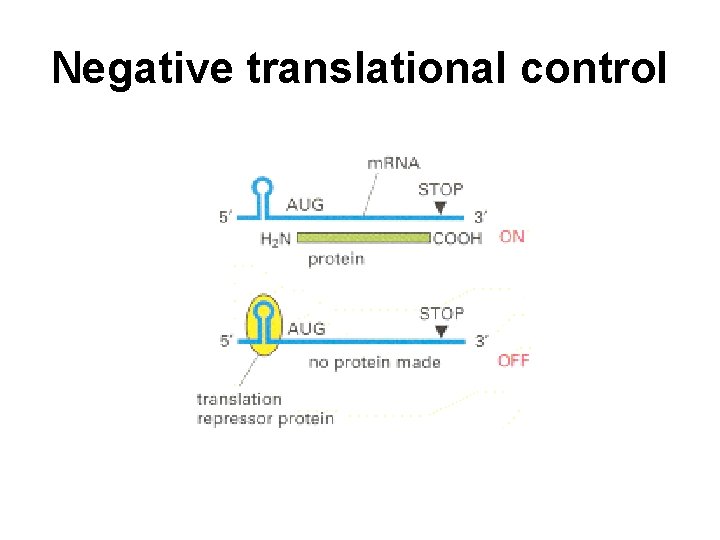
Negative translational control

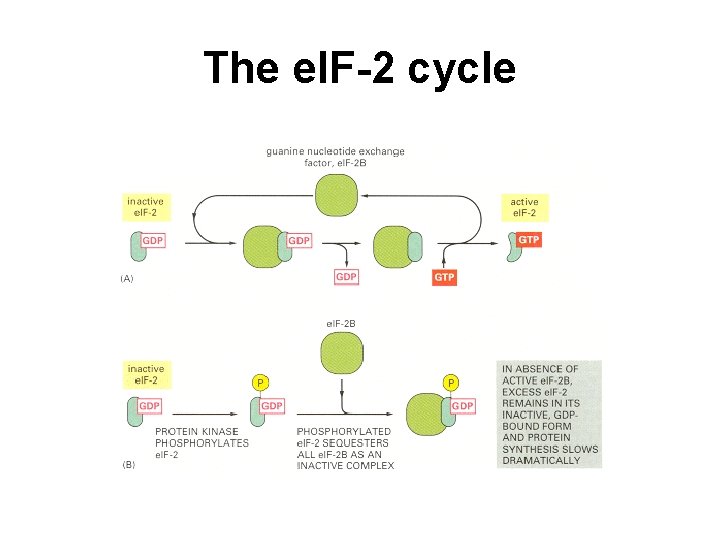
The el. F-2 cycle

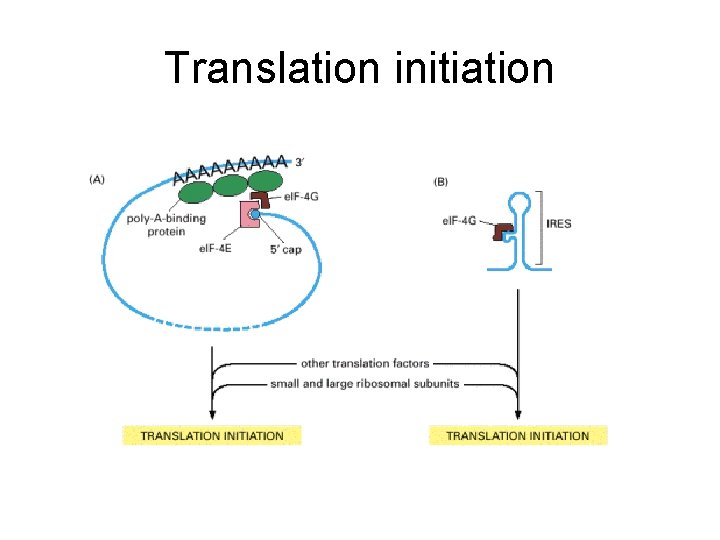
Translation initiation

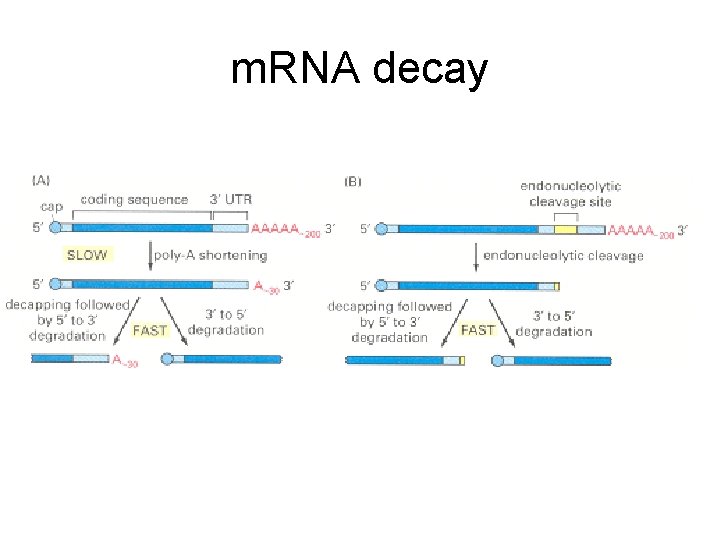
m. RNA decay

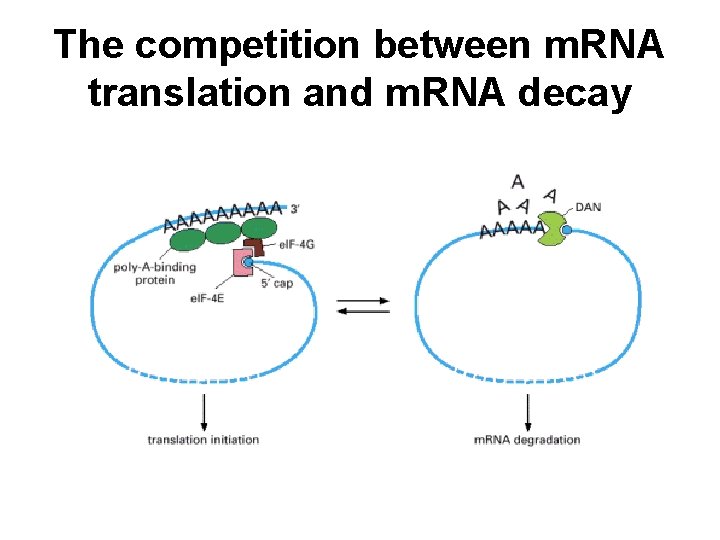
The competition between m. RNA translation and m. RNA decay

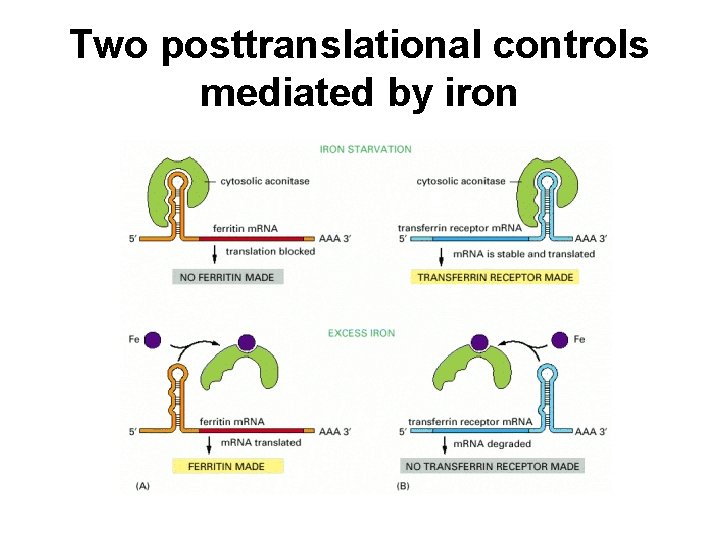
Two posttranslational controls mediated by iron

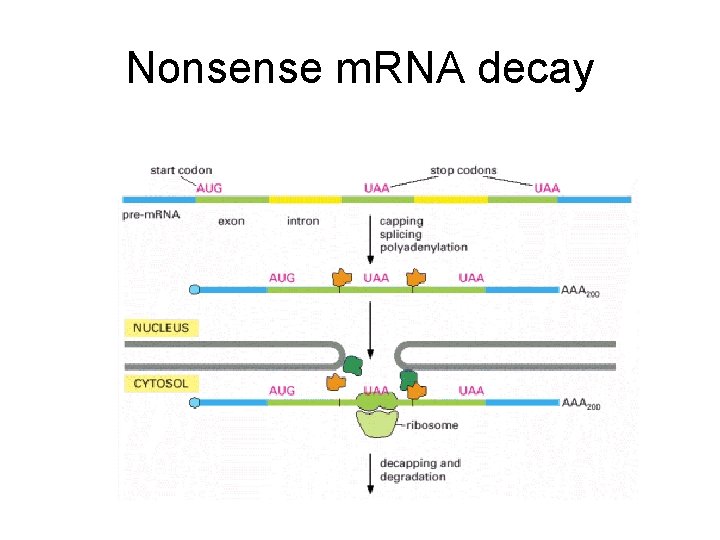
Nonsense m. RNA decay