Regular Expressions A simple and powerful way to


-x(0, 1)-{V} Perl Regexp: ^A. Example: PROSITE Patterns are regular expressions n n Pattern: <A-x-[ST](2)-x(0, 1)-{V} Perl Regexp: ^A.](https://slidetodoc.com/presentation_image/18b81cac829bcb963ae37fbdd3905731/image-3.jpg)



![Solution Reg. Exp n /[d{1, 3}. ]{3}d{1, 3}/ n /w+@w+-? w+. [a-z]{2, 4}/ n Solution Reg. Exp n /[d{1, 3}. ]{3}d{1, 3}/ n /w+@w+-? w+. [a-z]{2, 4}/ n](https://slidetodoc.com/presentation_image/18b81cac829bcb963ae37fbdd3905731/image-8.jpg)





















- Slides: 29

Regular Expressions A simple and powerful way to match characters Laurent Falquet, EPFL March, 2005 Swiss Institute of Bioinformatics Swiss EMBnet node

Regular Expressions What is a regular expression? Literal (or normal characters) Alphanumeric abc…ABC… 0123. . . Punctuation -_ , . ; : =()/+ *%&{}[]? !$’^|<>"@# Metacharacters Ex: ls *. java Flavors… awk, egrep, Emacs, grep, Perl, POSIX, Tcl, PROSITE !
-x(0, 1)-{V} Perl Regexp: ^A.](https://slidetodoc.com/presentation_image/18b81cac829bcb963ae37fbdd3905731/image-3.jpg)
Example: PROSITE Patterns are regular expressions n n Pattern: <A-x-[ST](2)-x(0, 1)-{V} Perl Regexp: ^A. [ST]{2}. ? [^V] Text: The sequence must start with an alanine, followed by any amino acid, followed by a serine or a threonine, two times, followed by any amino acid or nothing, followed by any amino acid except a valine. Simply the syntax differ…

Regular Expressions (1) n In Perl: /…/ ¨ Start and End of line n ¨ Match any of several n ¨ ^ start, $ end […] or (…|…) Match 0, 1 or more n n n . 1 of any ? 0 or 1 + 1 or more * 0 or more {m, n} range ! negation Examples Match every instance of a Swiss. Prot AC m/[OPQ][0 -9][A-Z 0 -9]{3}[0 -9]/; m/ [OPQ]d[A-Z 0 -9]{3}d/; Match every instance of a Swiss. Prot ID m/[A-Z 0 -9]{2, 5}_[A-Z 0 -9]{3, 5}/;

Regular Expressions (2) n Escape character or back reference ¨ n char or num Match operator ¨ d digit [0 -9] ¨ s whitespace [spacefnrt] ¨ w character [a-z. A-Z 0 -9_] ¨ DSW complement of dsw Byte notation num character in octal ¨ xnum character in hexadecimal ¨ cchar control character m/…/ n Shorthand ¨ n n Substitution operator ¨ s/…/…/ n n tr/…/…/ n n $var =~ s/colou? r/couleur/; Translate operator ¨ ¨ $var =~ m/colou? r/; $var !~ m/colou? r/; $revcomp =~ tr/ACGT/tgca/; Modifiers /…/# n n n /i case insensitive /g global match Many other /s, /m, /o, /x. . .

Regular Expressions (3) n Grouping ¨ External reference $var =~ s/sp: (wd{5})/swissprot AC=$1/; ¨ Internal reference $var =~ s/tr: (wd{5})|1/trembl AC=$1/; ¨ Numbering n n $1 to $9 $10 to more if needed. . . n Exercises Create a regexp to recognize any pseudo IP address: 012. 345. 678. 912 ¨ Create a regexp to recognize any email address: Jean. Dupond@isb-sib. ch ¨ Create a regexp to change any HTML tag to another ¨ n <address> -> <pre> On sib-dea: use visual_regexp-1. 2. tcl to check your regular expressions (requires X-windows) ¨

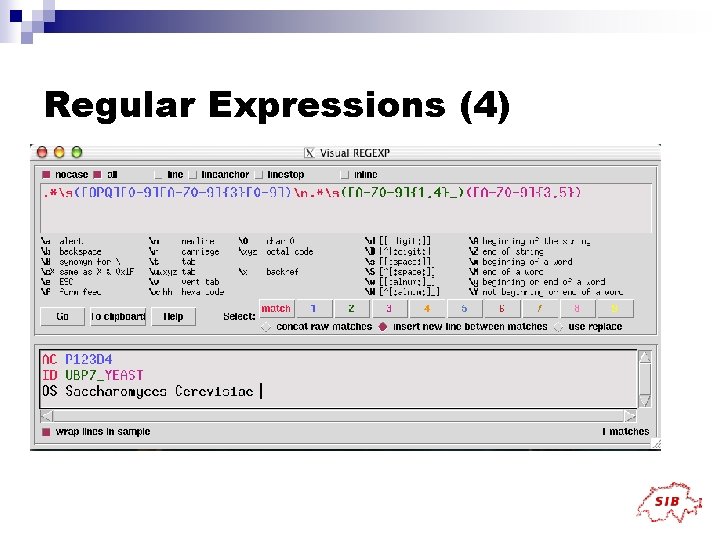
Regular Expressions (4)
![Solution Reg Exp n d1 3 3d1 3 n ww w az2 4 n Solution Reg. Exp n /[d{1, 3}. ]{3}d{1, 3}/ n /w+@w+-? w+. [a-z]{2, 4}/ n](https://slidetodoc.com/presentation_image/18b81cac829bcb963ae37fbdd3905731/image-8.jpg)
Solution Reg. Exp n /[d{1, 3}. ]{3}d{1, 3}/ n /w+@w+-? w+. [a-z]{2, 4}/ n /<(/? )address>/<$1 pre>/ generalized: n ¨ address = w+

Perl In-liners

In-liners: some options n n n n n -a autosplit (only with -n or -p) -c check syntax -d debugger -e pass script lines -h help -i direct editing of a file -n loop without print -p loop with print -v version … n Example: perl -e 'print qq(hello worldn); '

In-liners: -n and -p § perl -pe ‘s/r/n/g’ <file> n is equivalent to: open READ, “file”; while (<READ>) { s/r/n/g; print; } close(READ); n n n perl -i -pe ‘s/r/n/g’ <file> Warning: the -i option modifies the file directly perl -ne is the same without the “print”

In-liners: -a (only with -n or -p) n perl -ane ‘print @F, “n”; ’ <file> n Example: n is equivalent to: open READ, “file”; while (<READ>) { @F = split(‘ ‘); print @F, “n”; } close(READ); hits -b 'sw' -o pff 2 prf: CARD | perl -ane 'print join("t", reverse(@F)), "n"; '

In-liners: -a (only with -n or -p) n hits -b 'sw' -o pff 2 prf: CARD sw: ICEA_XENLA 1 90 prf: CARD 5 -1 18. 553 sw: RIK 2_MOUSE 435 513 prf: CARD 5 -11 15. 058 sw: CARC_HUMAN 1 88 prf: CARD 6 -1 15. 395 sw: NAL 1_HUMAN 1380 1463 prf: CARD 7 -1 15. 058 sw: ASC_HUMAN 113 195 prf: CARD 7 -2 15. 374 sw: CAR 8_HUMAN 347 430 prf: CARD 8 -1 18. 343 sw: CARF_HUMAN 134 218 prf: CARD 9 -1 12. 932 n hits -b 'sw' -o pff 2 prf: CARD | perl -ane 'print join("t", reverse(@F)), "n"; ' 18. 553 15. 058 15. 395 15. 058 15. 374 18. 343 12. 932 -1 -1 -1 -2 -1 -1 5 5 6 7 7 8 9 prf: CARD prf: CARD 90 513 88 1463 195 430 218 1 sw: ICEA_XENLA 435 sw: RIK 2_MOUSE 1 sw: CARC_HUMAN 1380 sw: NAL 1_HUMAN 113 sw: ASC_HUMAN 347 sw: CAR 8_HUMAN 134 sw: CARF_HUMAN

In-liners: examples n perl -e ‘print int(rand(100)), "n" for 1. . 100' | perl -e '$x{$_}=1 while <>; print sort {$a<=>$b} keys %x' for($i=0; $i<100; $i++) { $nb = int(rand(100)); $hash{$nb} = 1; } print sort {$a<=>$b} keys %hash;

In-liners: extract FASTA from SP open (READ, “/db/proteome/ECOLI. dat”); # open file while ($line=<READ>) { # read line by line until the end if($line=~ /^ID +(w+)/) { print “>$1n”; } # print fasta header if($line=~ /^ /) { $line =~ s/ //g; # remove spaces print $line; # print sequence line } } close(READ); n cat /db/proteome/ECOLI. dat | perl -ne ‘if (/^ID +(w+)/) {print">$1n"; } if(/^ /) {s/ //g; print}’

In-liners: your turn… n Create an In-liner that extracts non-redundant FASTA format sequences from a redundant database in Swiss. Prot format cat /db/proteome/ECOLI. dat | perl -ne ' if (/^ID +(w+)/) {print ">$1n”; } if(/^ /) {s/ //g; print}' | perl -e 'while(<>) { if (/>/) { $i=$_; $x{$i}=""} $x{$i}. =$_} print values %x’

Patterns in Biology n n Biology is all about patterns DNA patterns restriction sites n promoters/transcription factor binding sites n intron splice site n n Protein patterns conserved domains (motifs) n active sites n structural motifs (membrane spanning, signal peptide, etc. ) n

Computers are good at finding Patterns n n n ‘Find’ command in your word processor, “Find File” in your computer’s operating system Based on an underlying concept called a “Regular Expression” (regexp) a regexp is a text string, such as: “aatcg” ¨ can also have variable characters: “aa[ta]cg” ¨ or a wildcard: “aa[x]cg” ¨ or a variable spacer: “aa[x](120)cg”

grep n n grep is a handy Unix tool to “get regular expresssions” it is powerful and moderately complex tool (has one of the longest ‘man’ pages in the online Unix help system) n does not require it’s own O’Reilly book, but is a solid chapter in Intro and intermediate Unix/Linux books

Perl n n tr/// function tr means transliterate – replaces a character with another character $dna =~ tr/a/c/ replaces all “a” with “c” in in $dna It also works on a range: $dna =~ tr/a-z/A-Z/ replaces all lower case letters with upper case tr also counts $count = ($string =~ tr/A//) (you might think this also deletes all “A” from the string, but it doesn’t)

Perl Regular Expressions n n Perl Regular Expressions are more complex and more powerful than grep Can find and substitute bits of text in a single command Various options for “fuzzy” matches Perl regular expressions can get extremely complex - goes way beyond the scope of this course > man perlrequick

The Match Operator: =~ / / n Perl uses a special type of operator to do text matching with regular expressions: =~ / / n The =~ symbol is a pattern match comparison operator - n it can be translated as “contains” The forward slashes contain the pattern to be matched, like this: print “Eco. RI site found!” if $dna =~ /GAATTC/

Alternative Characters n Square brackets within the match expression allow for alternative characters: if $dna =~ /GGG[GATC]CCC/ n n This will match an DNA string that starts with GGG; has G, A, T, or C in the 4 th position, followed by CCC A vertical line within the /expression/ allows you to look for either of two completely different patterns: if $dna =~ /GAATTC|AAGCTT/

Wildcards n Perl has a set of wildcard characters for Reg. Exps. that are completely different than the ones used by Unix . ¨ the dot ( ) matches any character ¨ d matches any digit (a number from 0 -9) ¨ w matches any text character (a letter or number, not punctuation or space) ¨ s matches white space (any amount) ¨ ^ matches the beginning of a line ¨ $ matches the end of a line (Yes, this is very confusing!)

Repeat for a count n n Use curly brackets to show that a character repeats a specific number (or range) of times: find an Eco. RI fragment of 100 -500 bp length (two Eco. RI sites with any other sequence between): if $ecofrag =~ /GAATTC[GATC]{100, 500}GAATTC/ n The + sign is used to indicate an unlimited number of repeats (occurs 1 or more times)

It gets worse… n What if you need to match text that contains a special character? n n (the dot shows up all the time in Gen. Bank IDs, filenames, etc. ) Now you have to use a backslash () to “escape” the wildcard meaning of that character: if $seqname =~ /w+ . d/ -This would match any sequence ID that has some text characters, a dot, followed by a single digit: M 65783. 2

Grabbing parts of a string n n Regular expressions can do more than just ask ‘if” questions They can be used to extract parts of a line of text into variables; Check this out: /^>(w+)s(. +)$/; right? n Complete gibberish, It means: -look for the > sign at the beginning of a FASTA formatted sequence file -dump the first word (w+) into variable $1 (the sequence ID) -after a space, dump the rest of the line (. +), until you reach the end of line $, into variable $2 (the description)

You can also do Substitution n n To replace one string with another, use the tricky s/// function It works like this: s/expression/replacement/ $text =~ s/C-/A+/; (If only life were as easy as Perl)

n You know enough now to learn more Perl from Perl documentation, a book, or a website n > man perlintro > man perlrequick n Other cool websites n n http: //www. troubleshooters. com/codecorn/littperl/perlreg. htm http: //www. comp. leeds. ac. uk/Perl/oldindex. htm?