Referencebased Indexing of Sequence Databases Jayendra Venkateswaran Deepak

Reference-based Indexing of Sequence Databases Jayendra Venkateswaran, Deepak Lachwani, Tamer Kahveci, Christopher Jermaine University of Florida-Gainesville www. cise. ufl. edu/~jgvenkat VLDB 2006
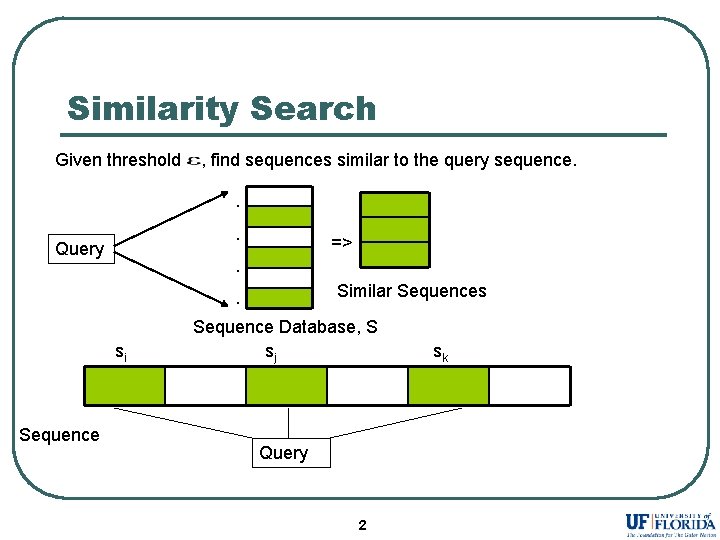
Similarity Search Given threshold , find sequences similar to the query sequence. . . Query => . Similar Sequences . si Sequence Database, S sj Query 2 sk

Measure: Edit Distance Edit Operations: Insert, Delete and Replace. Example: P: ACGTAC_GT | ||| || Q: A_GTACCGT Sequence Length: 12 3 Edit Operations: 2 Insertions and 1 Replace Edit Distance is the minimum number of edit operations needed to transform one sequence to another. 3

Edit Distance: Complexity Time and space complexity for computing Edit Distance between two sequences is O(n 2). Query . Sequence Database, S . |S| = 100, 000 . One Sequence Comparison: 0. 25 second. Time taken for single query: 7 hours. 4
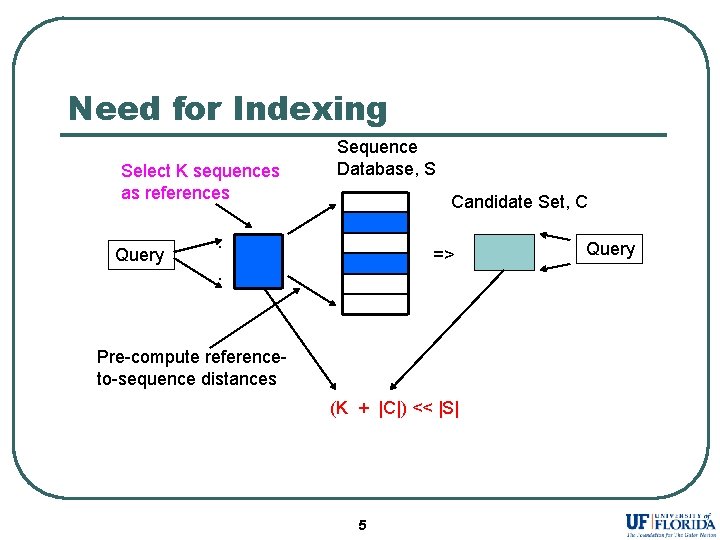
Need for Indexing Select K sequences as references Query Sequence Database, S Candidate Set, C . => . Pre-compute referenceto-sequence distances (K + |C|) << |S| 5 Query

Existing Methods • Hierarchical Methods, • VP-Tree (Yianilos, 1993) • MVP-Tree (Bozkaya et al. , 1997) • M-Tree (Ciaccia et al. , 1997), • Slim-Tree (Traina et al. , 2000), • DF-Tree (Traina et al. , 2002). • DBM-Tree (Vieira et al. , 2004) • Omni (Filho et al. , 2001) • Frequency Vector (Kahveci et al. , 2004). 6

Reference-based Indexing Reference Circle Including Query Database Sequences Reference Sequence 7 Query
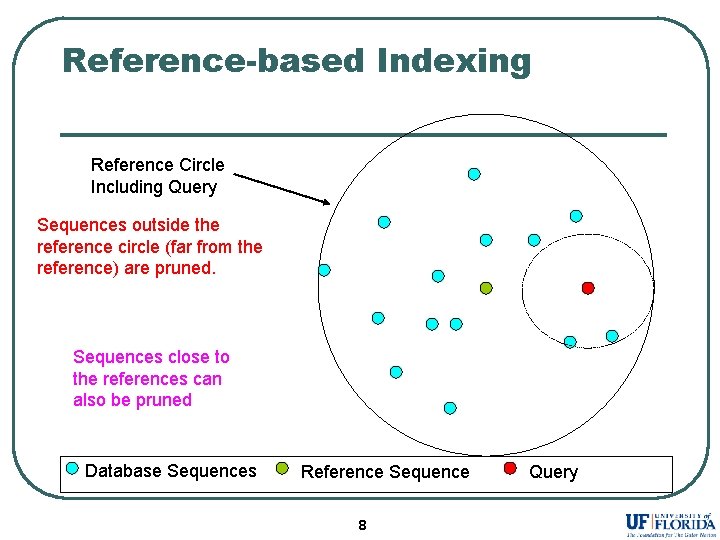
Reference-based Indexing Reference Circle Including Query Sequences outside the reference circle (far from the reference) are pruned. Sequences close to the references can also be pruned Database Sequences Reference Sequence 8 Query
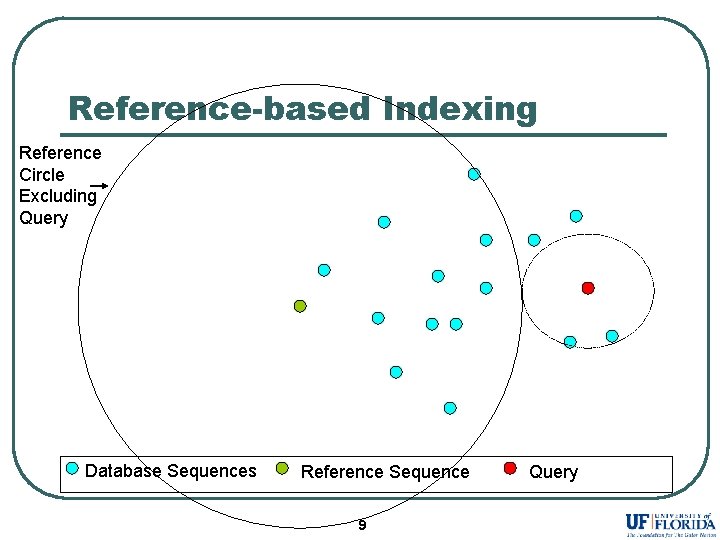
Reference-based Indexing Reference Circle Excluding Query Database Sequences Reference Sequence 9 Query
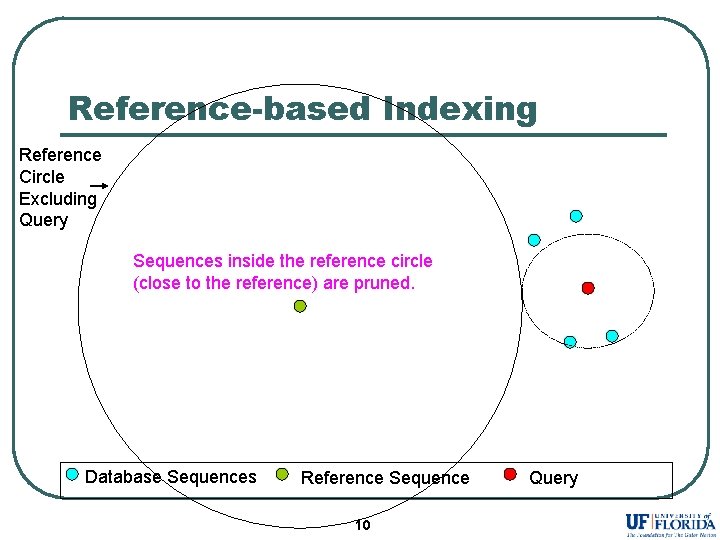
Reference-based Indexing Reference Circle Excluding Query Sequences inside the reference circle (close to the reference) are pruned. Database Sequences Reference Sequence 10 Query
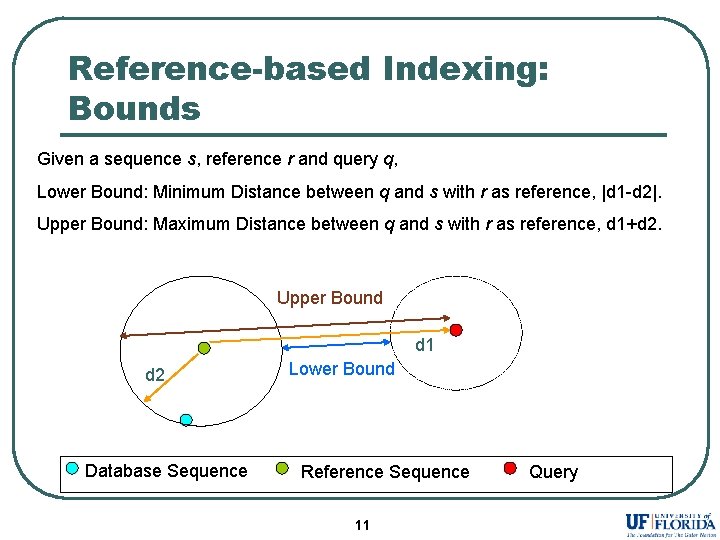
Reference-based Indexing: Bounds Given a sequence s, reference r and query q, Lower Bound: Minimum Distance between q and s with r as reference, |d 1 -d 2|. Upper Bound: Maximum Distance between q and s with r as reference, d 1+d 2. Upper Bound d 1 d 2 Database Sequence Lower Bound Reference Sequence 11 Query

Observations l l l Two types of pruning: • • Sequences close to references. Sequences far from references. A good reference set should be able to use both kinds of pruning effectively. Each reference should prune some part of the database not pruned by other references. 12

Outline l l l Selection of References Reference Assignment Search Algorithm Experimental Results Conclusions 13

Our Contributions l Selection of References: 1. Maximum Variance Selection: Reference with high variance of distance distributions with other sequences in the database. 2. Maximum Pruning: A Combinatorial approach of selecting the best reference set. l Assignment of References: • Each sequence has different set of references. 14
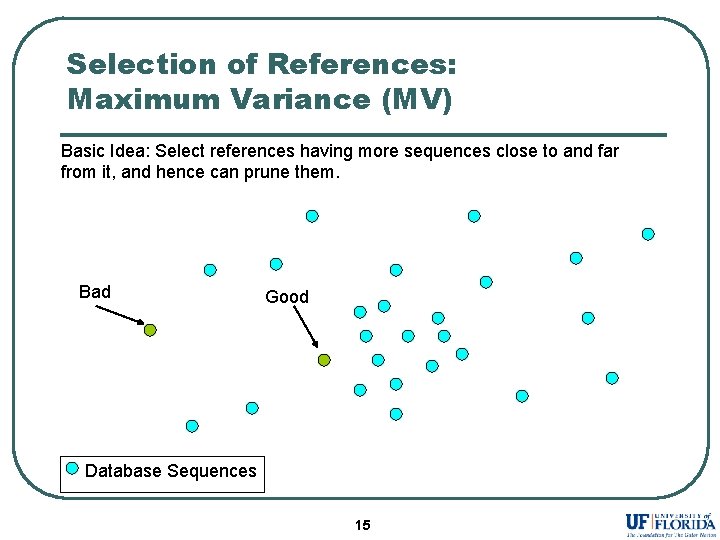
Selection of References: Maximum Variance (MV) Basic Idea: Select references having more sequences close to and far from it, and hence can prune them. Bad Good Database Sequences 15

Selection of References: Maximum Variance (MV) l l l Select references having sequences close to and far away from them. References have maximum variance of distance distributions with other sequences in the database. New reference prunes some part of the database not pruned by existing set of references. 16
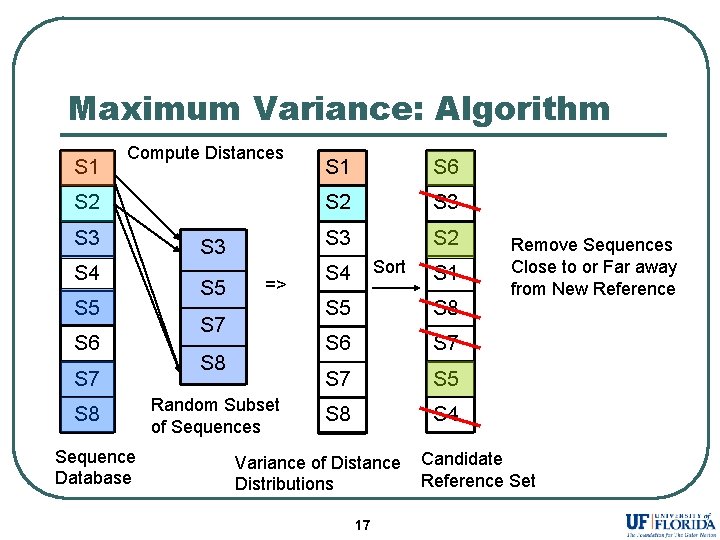
Maximum Variance: Algorithm S 1 Compute Distances S 2 S 3 S 4 S 5 S 6 S 7 S 8 Sequence Database S 3 S 5 => S 7 S 8 Random Subset of Sequences S 1 S 6 S 2 S 3 S 2 Sort S 4 S 1 S 5 S 8 S 6 S 7 S 5 S 8 S 4 Variance of Distance Distributions 17 Remove Sequences Close to or Far away from New Reference Candidate Reference Set
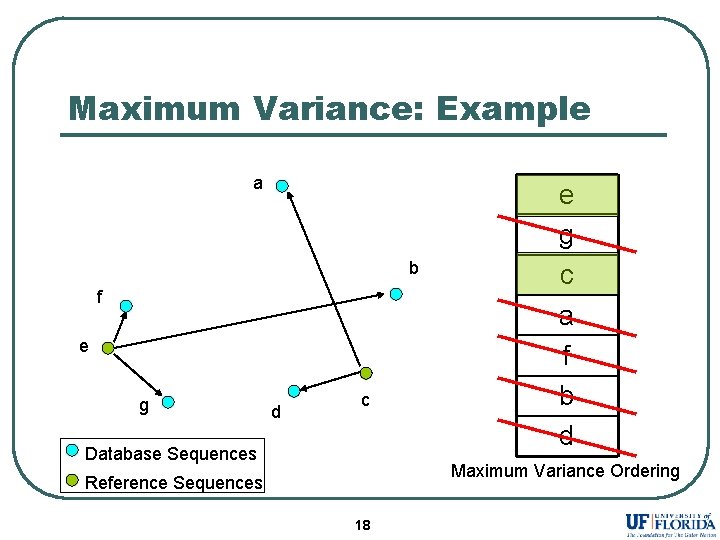
Maximum Variance: Example a b f e g d c Database Sequences e g c a f b d Maximum Variance Ordering Reference Sequences 18

Selection of References: Maximum Pruning (MP) l l Combinatorial approach to select the best reference set for given query set. Select reference set that can prune more sequences over all queries. Sample query set Q’ following the actual query distribution is given. Sampling techniques to reduce the complexity of this method. 19
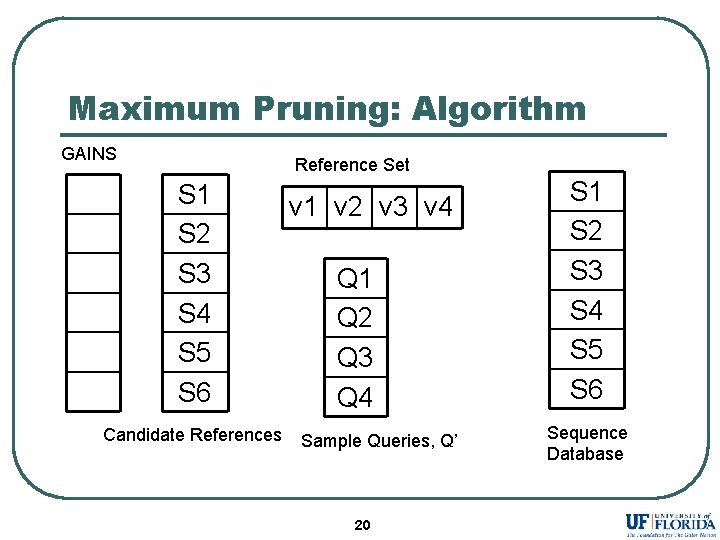
Maximum Pruning: Algorithm GAINS Reference Set S 1 S 2 S 3 S 4 S 5 S 6 Candidate References v 1 v 2 v 3 v 4 Q 1 Q 2 Q 3 Q 4 Sample Queries, Q’ 20 S 1 S 2 S 3 S 4 S 5 S 6 Sequence Database
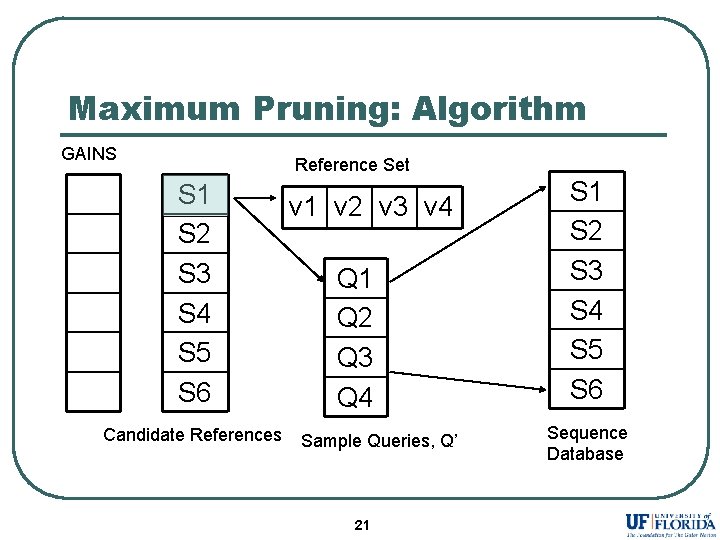
Maximum Pruning: Algorithm GAINS Reference Set S 1 S 2 S 3 S 4 S 5 S 6 Candidate References v 1 v 2 v 3 v 4 Q 1 Q 2 Q 3 Q 4 Sample Queries, Q’ 21 S 2 S 3 S 4 S 5 S 6 Sequence Database
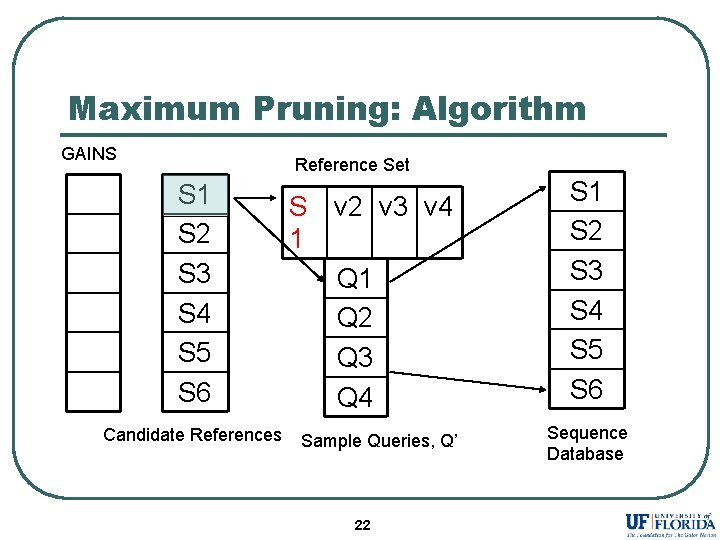
Maximum Pruning: Algorithm GAINS Reference Set S 1 S 2 S 3 S 4 S 5 S 6 Candidate References S 1 v 2 v 3 v 4 Q 1 Q 2 Q 3 Q 4 Sample Queries, Q’ 22 S 1 S 2 S 3 S 4 S 5 S 6 Sequence Database
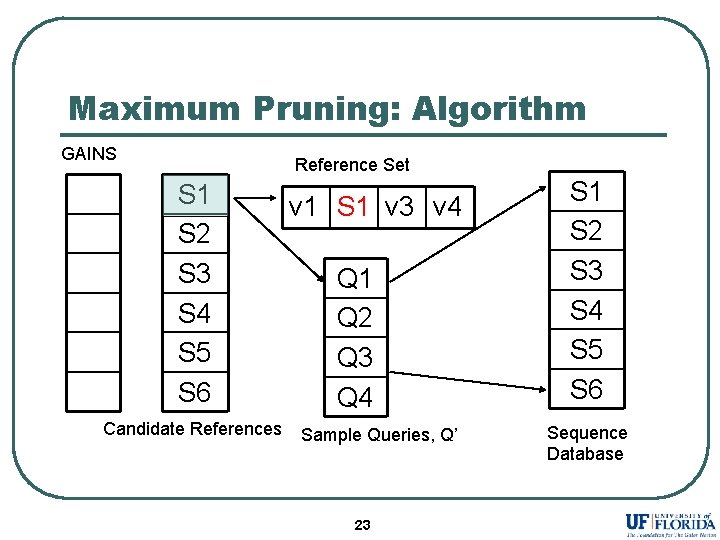
Maximum Pruning: Algorithm GAINS Reference Set S 1 S 2 S 3 S 4 S 5 S 6 Candidate References v 1 S 1 v 3 v 4 Q 1 Q 2 Q 3 Q 4 Sample Queries, Q’ 23 S 1 S 2 S 3 S 4 S 5 S 6 Sequence Database
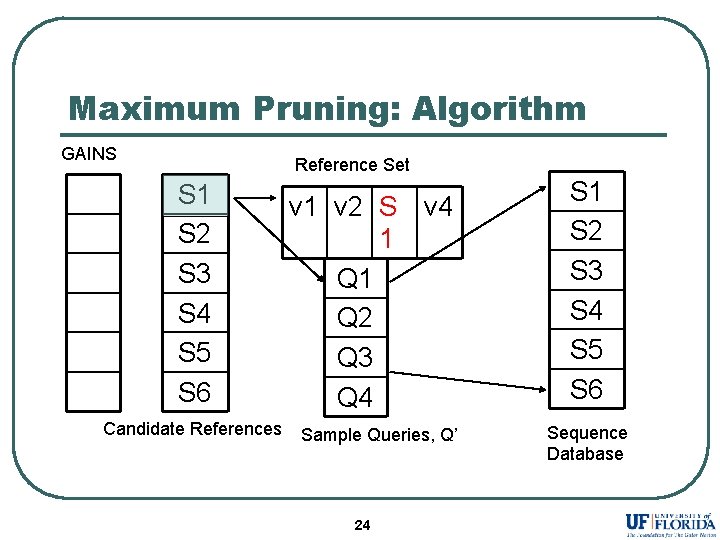
Maximum Pruning: Algorithm GAINS Reference Set S 1 S 2 S 3 S 4 S 5 S 6 Candidate References v 1 v 2 S 1 Q 2 Q 3 Q 4 v 4 Sample Queries, Q’ 24 S 1 S 2 S 3 S 4 S 5 S 6 Sequence Database
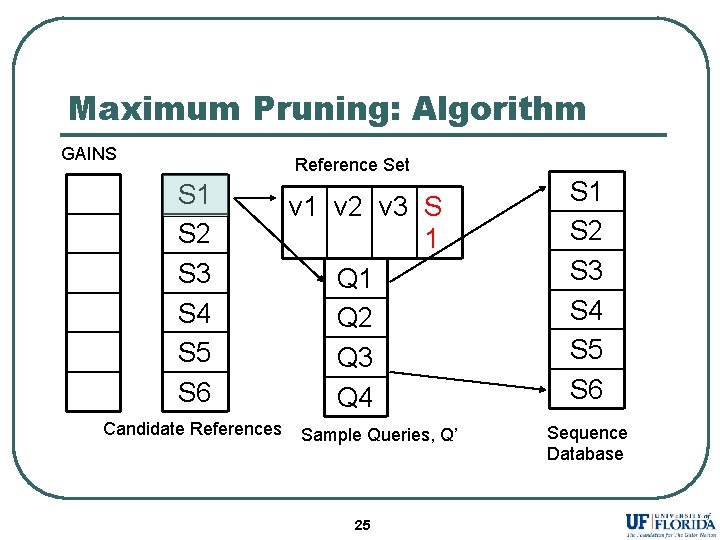
Maximum Pruning: Algorithm GAINS Reference Set S 1 S 2 S 3 S 4 S 5 S 6 Candidate References v 1 v 2 v 3 S 1 Q 2 Q 3 Q 4 Sample Queries, Q’ 25 S 1 S 2 S 3 S 4 S 5 S 6 Sequence Database
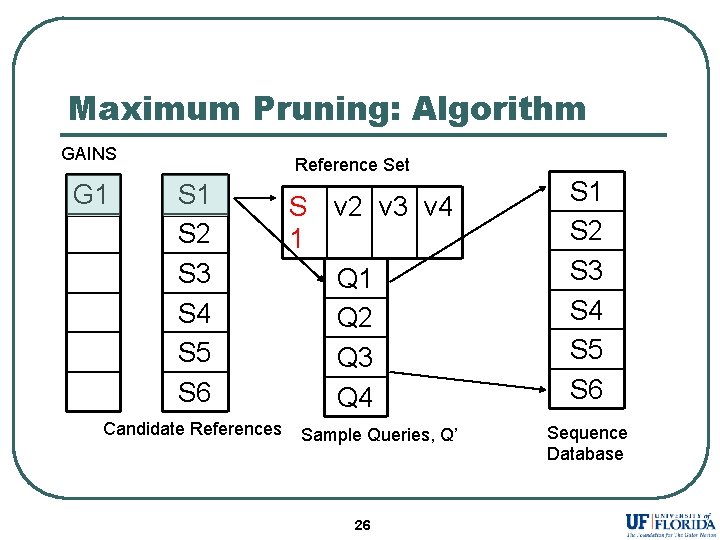
Maximum Pruning: Algorithm GAINS G 1 Reference Set S 1 S 2 S 3 S 4 S 5 S 6 Candidate References S 1 v 2 v 3 v 4 Q 1 Q 2 Q 3 Q 4 Sample Queries, Q’ 26 S 1 S 2 S 3 S 4 S 5 S 6 Sequence Database
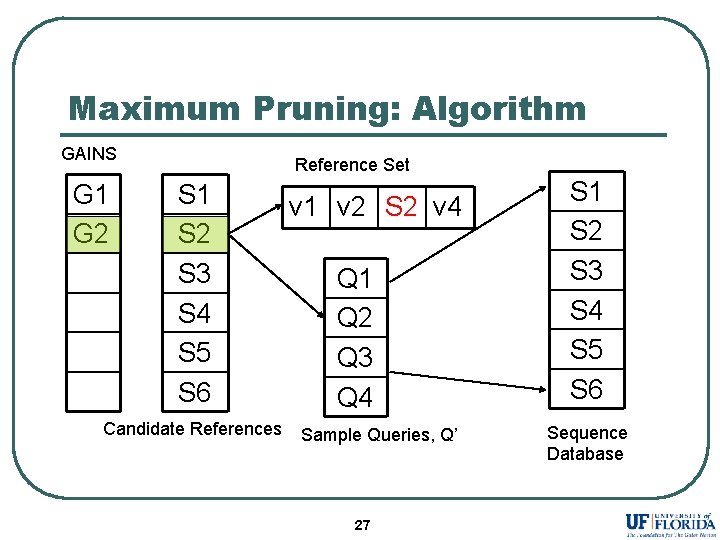
Maximum Pruning: Algorithm GAINS G 1 G 2 Reference Set S 1 S 2 S 3 S 4 S 5 S 6 Candidate References v 1 v 2 S 2 v 4 Q 1 Q 2 Q 3 Q 4 Sample Queries, Q’ 27 S 1 S 2 S 3 S 4 S 5 S 6 Sequence Database
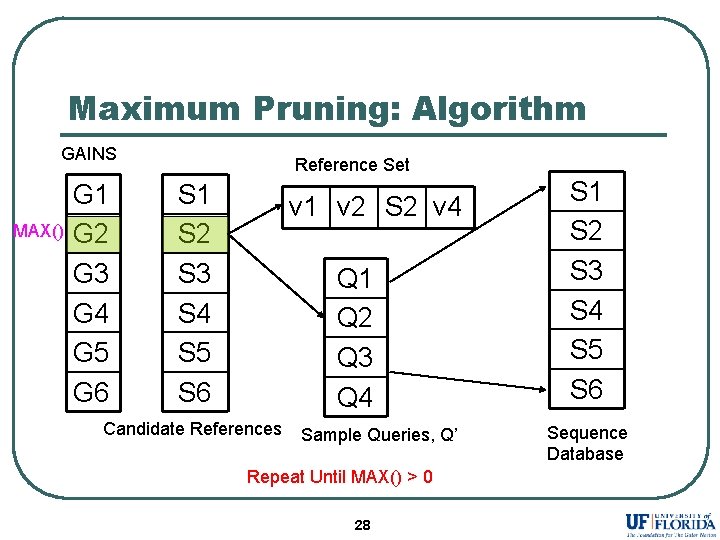
Maximum Pruning: Algorithm GAINS MAX() G 1 G 2 G 3 G 4 G 5 G 6 Reference Set S 1 S 2 S 3 S 4 S 5 S 6 v 1 v 2 S 2 v 4 Q 1 Q 2 Q 3 Q 4 Candidate References Sample Queries, Q’ Repeat Until MAX() > 0 28 S 1 S 2 S 3 S 4 S 5 S 6 Sequence Database
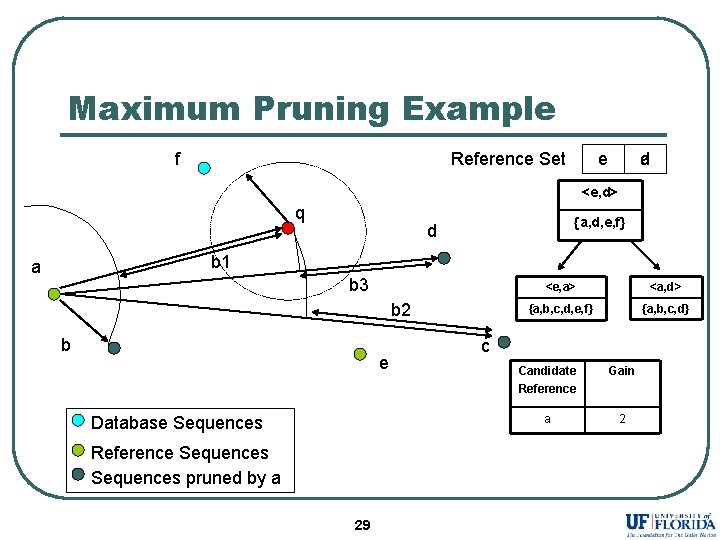
Maximum Pruning Example f Reference Set e d a <e, d> q {a, d, e, f} d b 1 a b 3 b 2 b e Database Sequences Reference Sequences pruned by a 29 <e, a> <a, d> {a, b, c, d, e, f} {a, b, c, d} c Candidate Reference Gain a 2
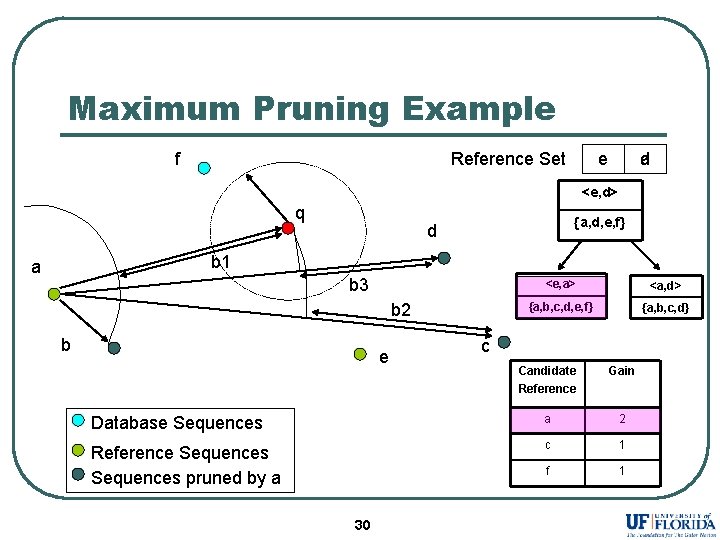
Maximum Pruning Example f Reference Set e d a <e, d> q {a, d, e, f} d b 1 a b 3 b 2 b e <e, a> <a, d> {a, b, c, d, e, f} {a, b, c, d} c Candidate Reference Gain Database Sequences a 2 Reference Sequences pruned by a c 1 f 1 30

Outline l l l Selection of References Assignment of References Search Algorithm Experimental Results Conclusions 31
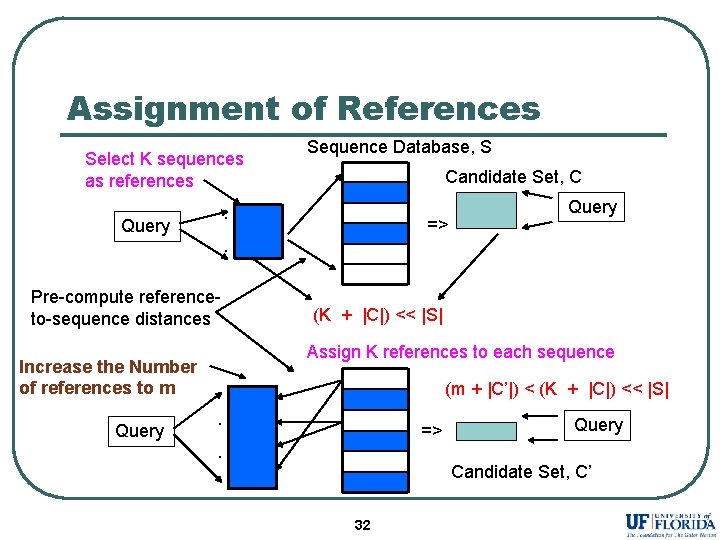
Assignment of References Select K sequences as references Candidate Set, C . Query => Query . Pre-compute referenceto-sequence distances (K + |C|) << |S| Assign K references to each sequence Increase the Number of references to m Query Sequence Database, S (m + |C’|) < (K + |C|) << |S|. => Query . Candidate Set, C’ 32
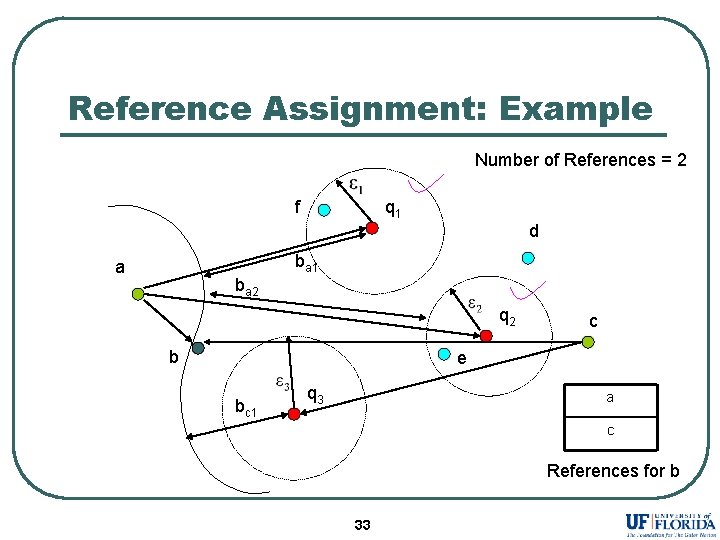
Reference Assignment: Example Number of References = 2 f q 1 d ba 1 a ba 2 q 2 b c e bc 1 q 3 a c References for b 33

Outline l l l Selection of References Reference Assignment Search Algorithm Experimental Results Conclusions 34
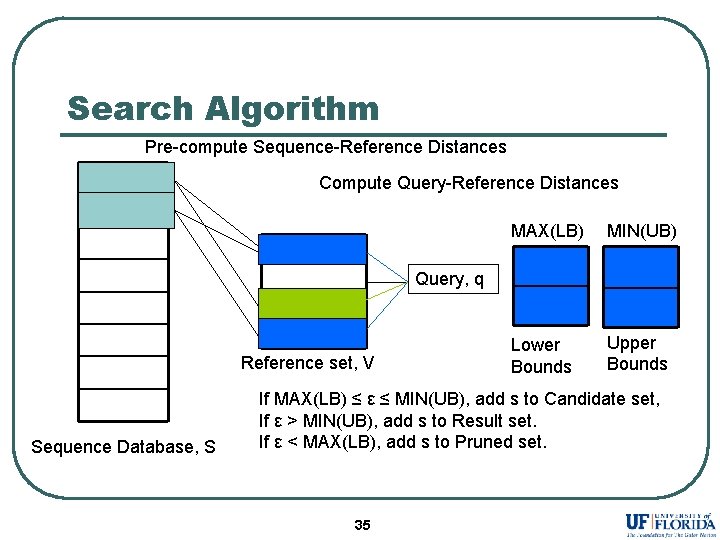
Search Algorithm Pre-compute Sequence-Reference Distances Compute Query-Reference Distances MAX(LB) MIN(UB) Lower Bounds Upper Bounds Query, q Reference set, V Sequence Database, S If MAX(LB) ≤ ε ≤ MIN(UB), add s to Candidate set, If ε > MIN(UB), add s to Result set. If ε < MAX(LB), add s to Pruned set. 35

Outline l l l Selection of References Reference Assignment Search Algorithm Experimental Results Conclusions 36

Experimental Setup l Datasets • • • DNA: Alphabet size of 4 and 20000 sequences. Protein: Alphabet size of 20 and 4000 sequences of up to 500 amino acids. Text: Alphabet size of 36 and 8000 sequences of length 100 each. l Size of Reference Set, m = 200. l Experiments, • Comparison with our methods • Comparison with other methods • • Maximum Variance with same and different reference sets (MV-S and MV-D). Maximum Pruning with same and different reference sets (MP-S and MP-D). • • • Frequency Vector (Kahveci et al. , 2004). Omni (Filho et al. , 2001) Others: M-Tree (Ciaccia et al. , 1007), Slim-Tree (Traina et al. , 2000), DBM-Tree (Vieira et al. , 2004) and DF-Tree (Traina et al. , 2002). 37
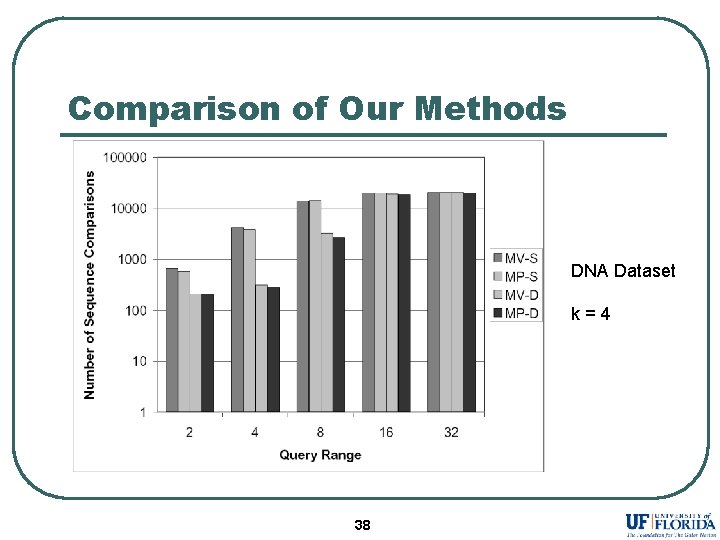
Comparison of Our Methods DNA Dataset k=4 38
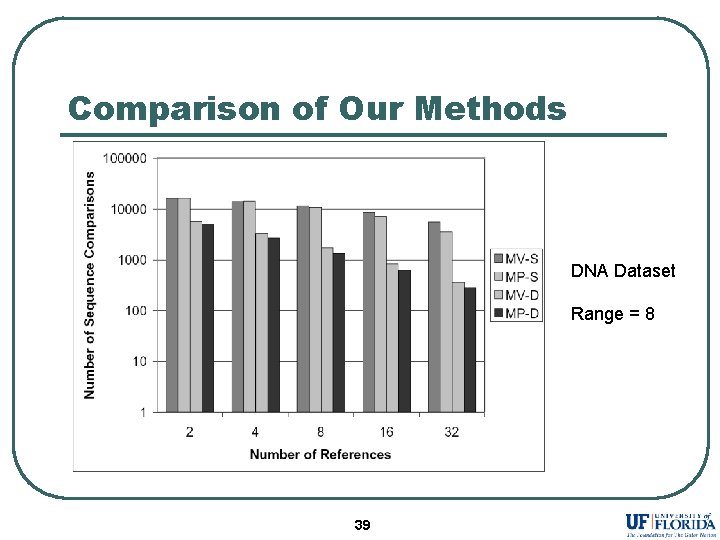
Comparison of Our Methods DNA Dataset Range = 8 39
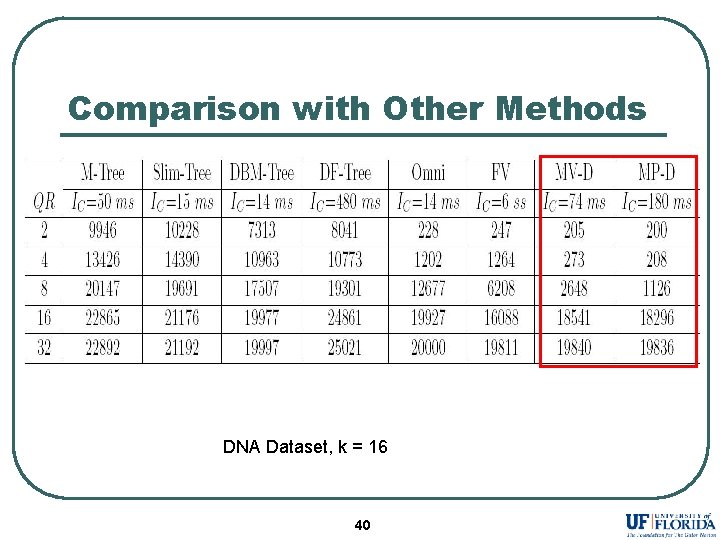
Comparison with Other Methods DNA Dataset, k = 16 40

Conclusion l l l References selected by Maximum Variance and Maximum Pruning eliminates more database sequences as compared to existing selection strategies. Assigning different reference set to each sequence dramatically improves the performance. MP-D outperforms existing methods in almost all the experiments. 41

Thank You Questions ? jgvenkat@cise. ufl. edu
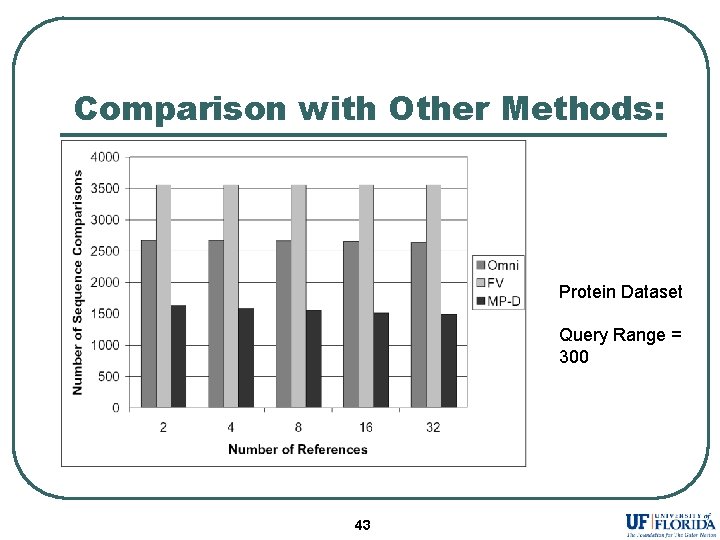
Comparison with Other Methods: Protein Dataset Query Range = 300 43

Assignment of References: Memory Limitations Main memory stores pre-computed reference-tosequence distances along with the references. l For each [s, vi] pair (s S, vi V), store [i, ED(s, vi)] (Takes 8 bytes). l Given the available main memory in bytes, B B = 8 KN + zm N: Number of sequences in the database. K: Number of references per sequence. z: Size of each sequence in bytes. m: Number of references in reference set. l Example: Given B = 1 GB, N = 10 million, z = 100 and m = 1000, then K = 13. l 44
- Slides: 44