RECOORD REcalculated COORdinates Database Jurgen Doreleijers Center for

RECOORD REcalculated COORdinates Database Jurgen Doreleijers Center for Eukaryotic Structural Genomics University of Madison-Wisconsin jurgen@bmrb. wisc. edu Aart Nederveen Bijvoet Center for Biomolecular Research Utrecht University a. j. nederveen@chem. uu. nl Wim Vranken Macromolecular Structure Database European Bioinformatics Institute wim@ebi. ac. uk

Aim • Recalculation of protein structures based on deposited NMR restraints using state of the art methods • Goals: • decrease user- and software-dependent biases • allow a better comparison between structures • comparison between different structure calculation programs • provide a database for the development and assessments of validation tools and calculation protocols
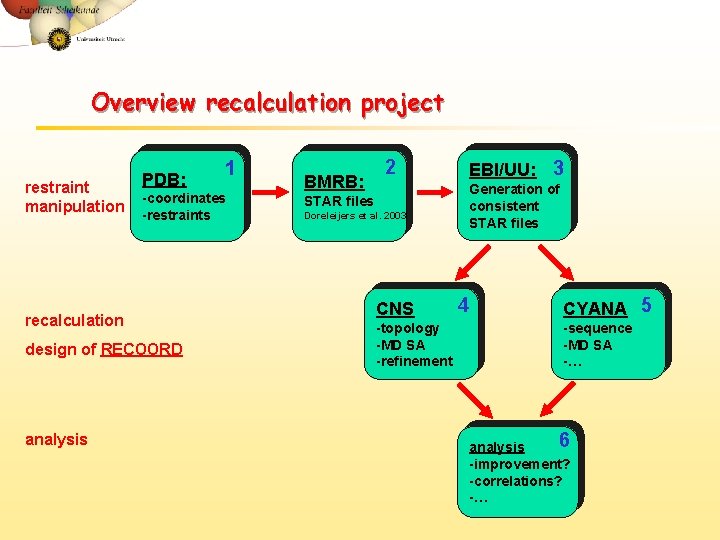
Overview recalculation project restraint manipulation PDB: -coordinates -restraints recalculation design of RECOORD analysis 1 BMRB: 2 EBI/UU: 3 Generation of consistent STAR files Doreleijers et al. 2003 CNS -topology -MD SA -refinement 4 CYANA 5 -sequence -MD SA -… 6 analysis -improvement? -correlations? -…

Databases now publicly available • DOCR/FRED (BMRB) databases containing converted and filtered restraints http: //www. bmrb. wisc. edu/servlets/MRGrid. Servlet • RECOORD (EBI) database containing recalculated coordinates http: //www. ebi. ac. uk/msd/recoord

PDB: Selection -coordinates -restraints • Formats (if distance restraints • CNS/XPLOR • DIANA/DYANA/CYANA • DISCOVER/MSI • PDB entries selected: • only proteins • no HET atoms • multimers allowed (not yet re-calculated) • at least 20 residues Finally 545 monomers were selected • 1 available): BMRB: 2 STAR files Doreleijers et al. 2003

EBI/UU: 3 Conversion issues • Generation of consistent STAR files Data is converted to formats readable by calculation software (e. g. XPLOR/CNS and CYANA) by the Format. Converter available within CCPN software (Wim Vranken, EBI). Problems: • Differences between coordinate and restraint data: • e. g. 1 chain in pdb entry, 2 chains in restraint list • residue numbering can differ in PDB entry and restraint list • restraints for residues not present in PDB entry… • Nomenclature in restraint list
CNS Building topology 4 -topology -MD SA -refinement • • Starting script: generate_easy. inp from CNS • CYANA: sequence based on CNS topology • Add CYSS, HIST+, c. PRO in sequence • Automated generation of disulfide restraints CYANA -sequence -MD SA -… Automated detection in original ensemble of: • Disulfide bridges (<3Å S-S distance in original first models) • CIS peptides (if |w|<25º in original first models) • Protonation state of histidines (use CNS patches HISD, HISE) 5

CNS CONDOR computer cluster CS University Madison -topology -MD SA -refinement • • More than 800 processor used • Example 2 EZM, calculation of 1 model 4 CYANA -sequence -MD SA -… Total CPU time: 31, 169 hours ( 3. 5 years on single workstation) (101 a. a. & 2. 2 GHz P 4 computer) CYANA CNS 31 seconds 340 seconds 5

Evaluation of structure quality • • 6 analysis -improvement? -correlations? -… Agreement with experimental restraints Improvement? Comparison CNS and CYANA Relation NMR data quality and structural quality
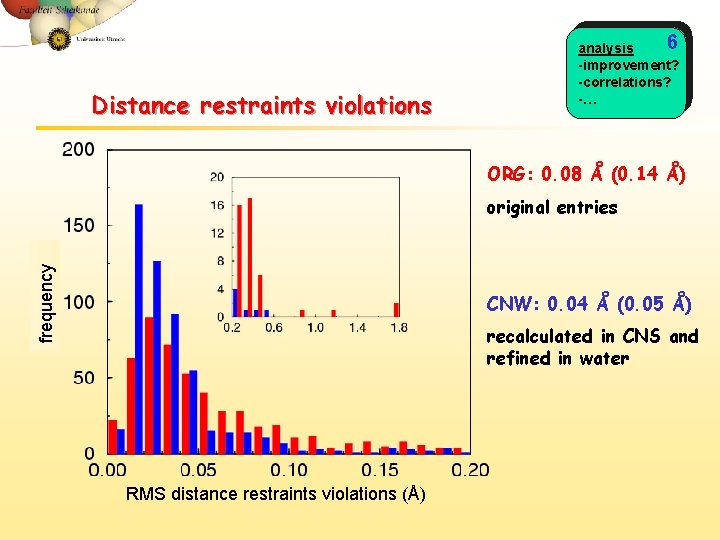
Distance restraints violations 6 analysis -improvement? -correlations? -… ORG: 0. 08 Å (0. 14 Å) frequency original entries CNW: 0. 04 Å (0. 05 Å) recalculated in CNS and refined in water RMS distance restraints violations (Å)
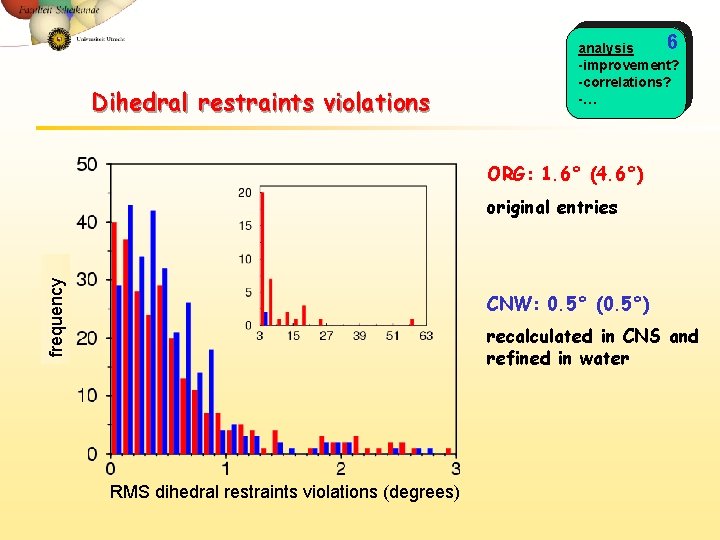
Dihedral restraints violations 6 analysis -improvement? -correlations? -… ORG: 1. 6° (4. 6°) frequency original entries CNW: 0. 5° (0. 5°) recalculated in CNS and refined in water RMS dihedral restraints violations (degrees)
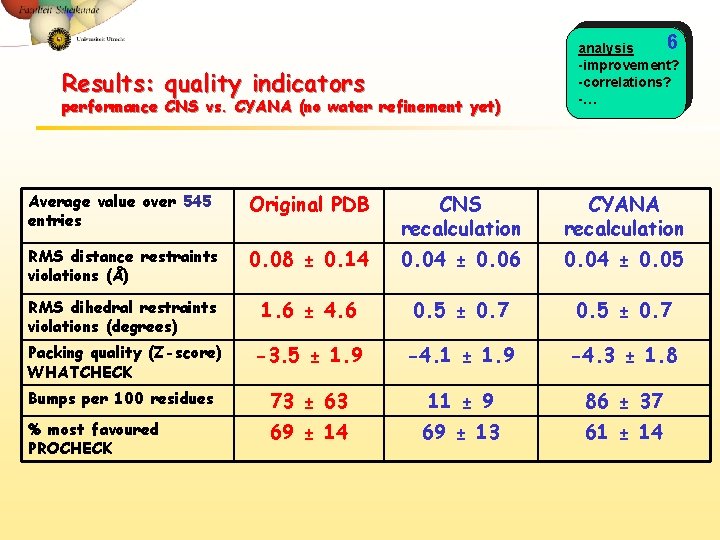
Results: quality indicators performance CNS vs. CYANA (no water refinement yet) 6 analysis -improvement? -correlations? -… Average value over 545 entries Original PDB CNS recalculation CYANA recalculation RMS distance restraints violations (Å) 0. 08 ± 0. 14 0. 04 ± 0. 06 0. 04 ± 0. 05 RMS dihedral restraints violations (degrees) 1. 6 ± 4. 6 0. 5 ± 0. 7 Packing quality (Z-score) WHATCHECK -3. 5 ± 1. 9 -4. 1 ± 1. 9 -4. 3 ± 1. 8 Bumps per 100 residues 73 ± 63 11 ± 9 86 ± 37 % most favoured PROCHECK 69 ± 14 69 ± 13 61 ± 14
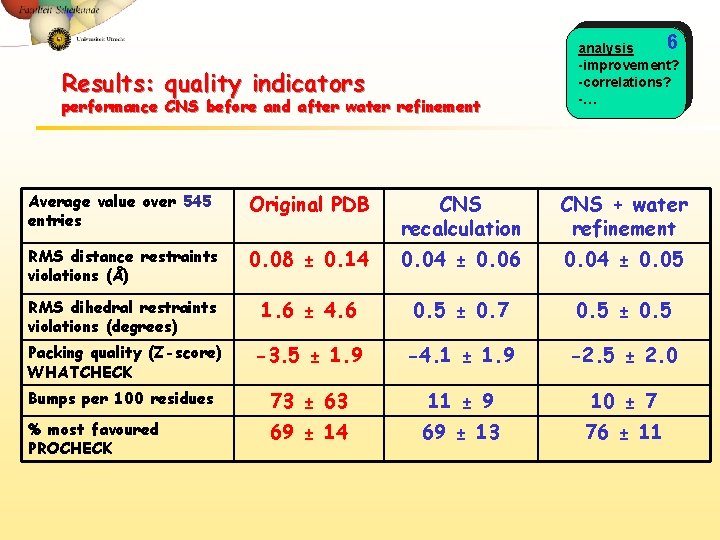
Results: quality indicators performance CNS before and after water refinement 6 analysis -improvement? -correlations? -… Average value over 545 entries Original PDB CNS recalculation CNS + water refinement RMS distance restraints violations (Å) 0. 08 ± 0. 14 0. 04 ± 0. 06 0. 04 ± 0. 05 RMS dihedral restraints violations (degrees) 1. 6 ± 4. 6 0. 5 ± 0. 7 0. 5 ± 0. 5 Packing quality (Z-score) WHATCHECK -3. 5 ± 1. 9 -4. 1 ± 1. 9 -2. 5 ± 2. 0 Bumps per 100 residues 73 ± 63 11 ± 9 10 ± 7 % most favoured PROCHECK 69 ± 14 69 ± 13 76 ± 11
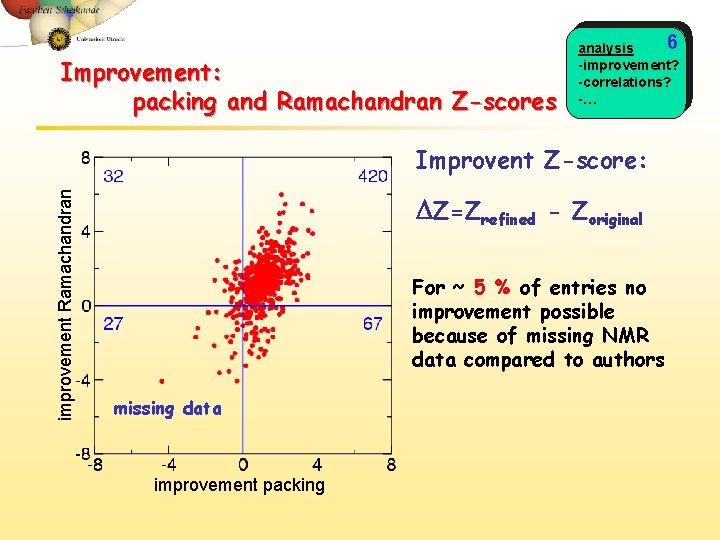
Improvement: packing and Ramachandran Z-scores 6 analysis -improvement? -correlations? -… improvement Ramachandran Improvent Z-score: DZ=Zrefined - Zoriginal For ~ 5 % of entries no improvement possible because of missing NMR data compared to authors missing data improvement packing
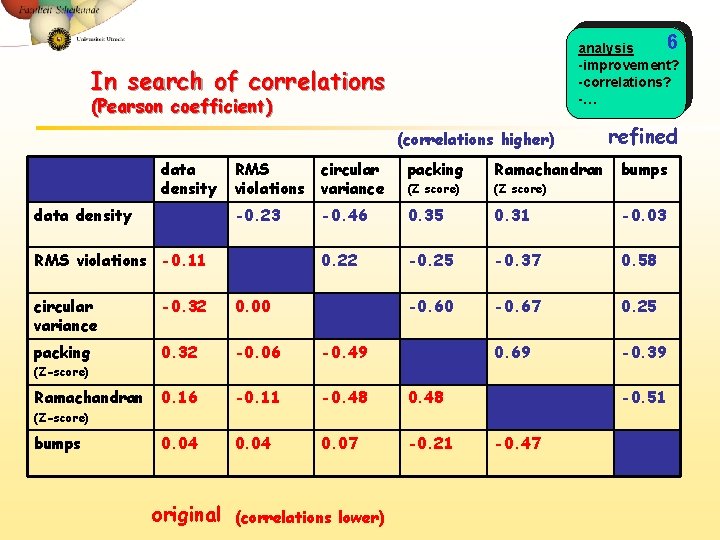
6 analysis -improvement? -correlations? -… In search of correlations (Pearson coefficient) (correlations higher) data density refined RMS violations circular variance packing Ramachandran (Z score) -0. 23 -0. 46 0. 35 0. 31 -0. 03 0. 22 -0. 25 -0. 37 0. 58 -0. 60 -0. 67 0. 25 0. 69 -0. 39 RMS violations -0. 11 circular variance -0. 32 0. 00 packing 0. 32 -0. 06 -0. 49 0. 16 -0. 11 -0. 48 0. 04 0. 07 -0. 21 bumps (Z-score) Ramachandran -0. 51 (Z-score) bumps original (correlations lower) -0. 47
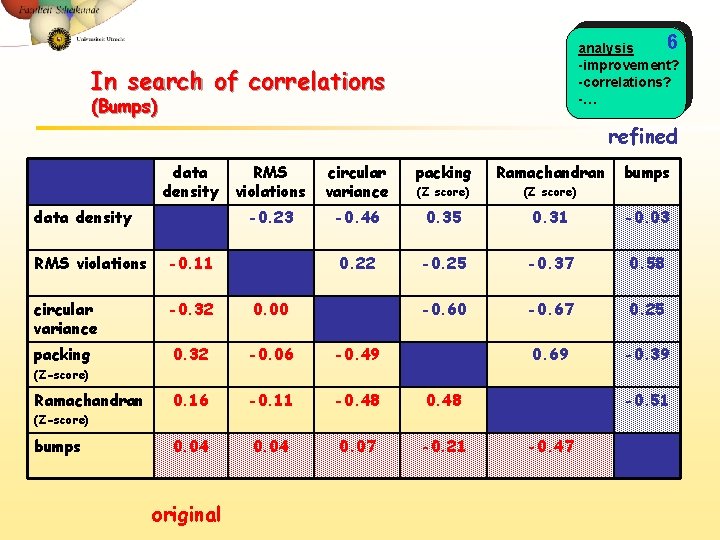
6 analysis -improvement? -correlations? -… In search of correlations (Bumps) refined data density RMS violations circular variance packing Ramachandran (Z score) -0. 23 -0. 46 0. 35 0. 31 -0. 03 0. 22 -0. 25 -0. 37 0. 58 -0. 60 -0. 67 0. 25 0. 69 -0. 39 RMS violations -0. 11 circular variance -0. 32 0. 00 packing 0. 32 -0. 06 -0. 49 0. 16 -0. 11 -0. 48 0. 04 0. 07 -0. 21 bumps (Z-score) Ramachandran -0. 51 (Z-score) bumps original -0. 47
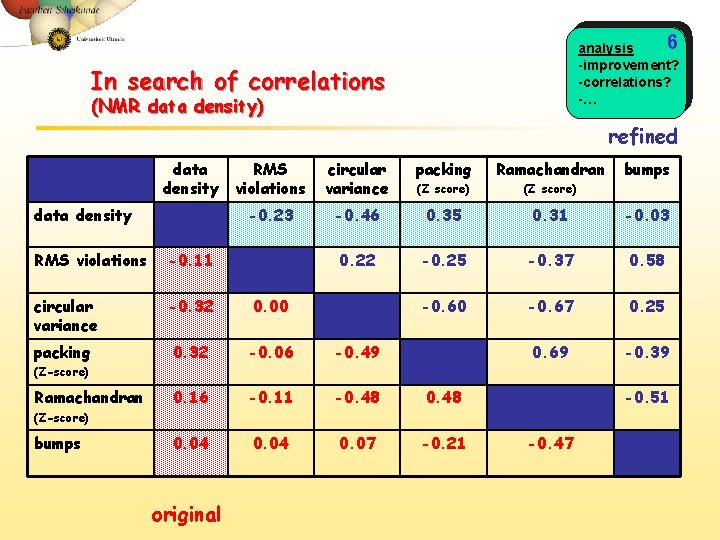
6 analysis -improvement? -correlations? -… In search of correlations (NMR data density) refined data density RMS violations circular variance packing Ramachandran (Z score) -0. 23 -0. 46 0. 35 0. 31 -0. 03 0. 22 -0. 25 -0. 37 0. 58 -0. 60 -0. 67 0. 25 0. 69 -0. 39 RMS violations -0. 11 circular variance -0. 32 0. 00 packing 0. 32 -0. 06 -0. 49 0. 16 -0. 11 -0. 48 0. 04 0. 07 -0. 21 bumps (Z-score) Ramachandran -0. 51 (Z-score) bumps original -0. 47
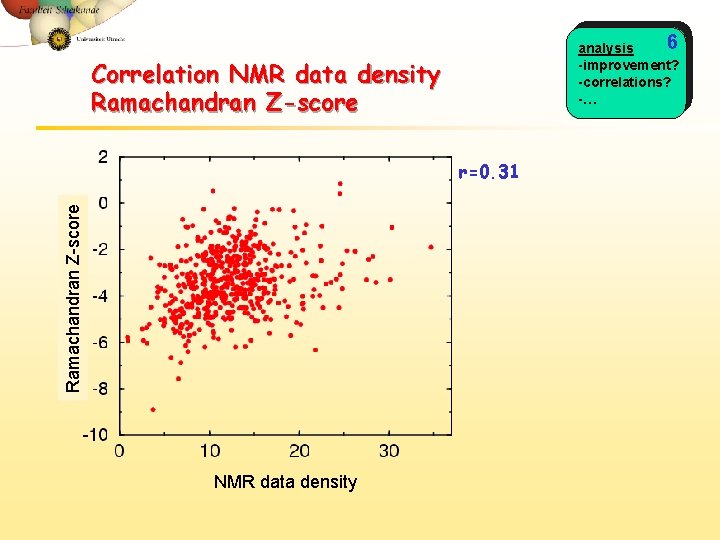
6 analysis -improvement? -correlations? -… Correlation NMR data density Ramachandran Z-score r=0. 31 NMR data density
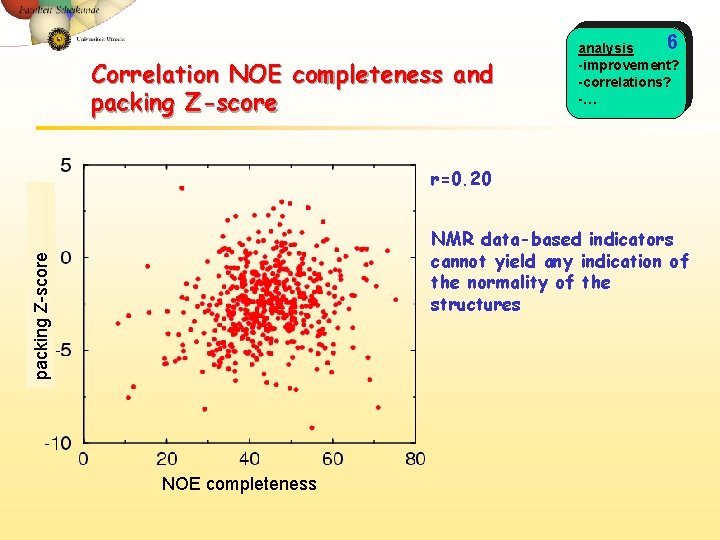
Correlation NOE completeness and packing Z-score 6 analysis -improvement? -correlations? -… r=0. 20 packing Z-score NMR data-based indicators cannot yield any indication of the normality of the structures NOE completeness

6 analysis -improvement? -correlations? -… In search of correlations (Precision) refined data density RMS violations circular variance packing Ramachandran (Z score) -0. 23 -0. 46 0. 35 0. 31 -0. 03 0. 22 -0. 25 -0. 37 0. 58 -0. 60 -0. 67 0. 25 0. 69 -0. 39 RMS violations -0. 11 circular variance -0. 32 0. 00 packing 0. 32 -0. 06 -0. 49 0. 16 -0. 11 -0. 48 0. 04 0. 07 -0. 21 bumps (Z-score) Ramachandran -0. 51 (Z-score) bumps original -0. 47
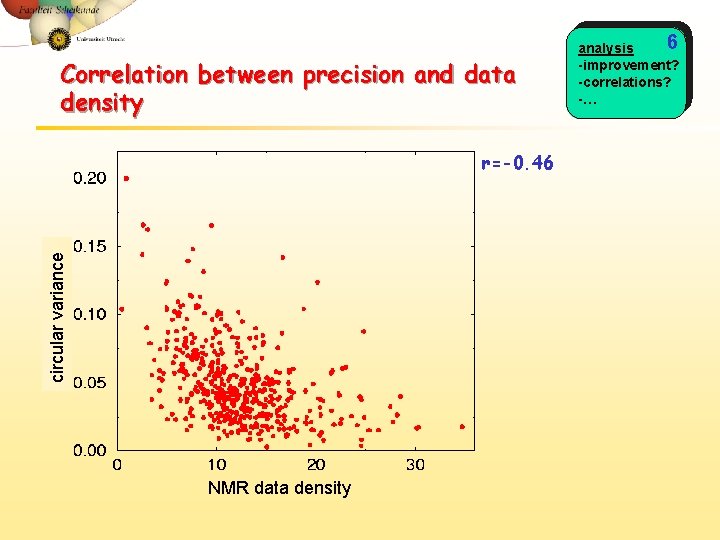
Correlation between precision and data density circular variance r=-0. 46 NMR data density 6 analysis -improvement? -correlations? -…
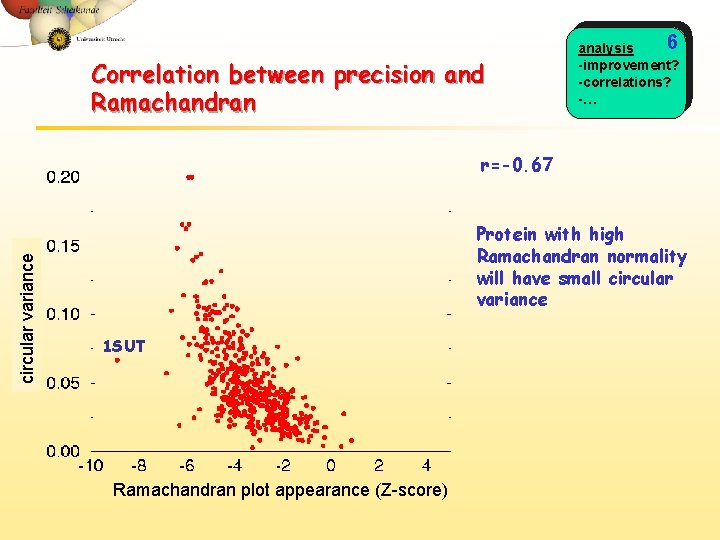
Correlation between precision and Ramachandran 6 analysis -improvement? -correlations? -… circular variance r=-0. 67 Protein with high Ramachandran normality will have small circular variance 1 SUT Ramachandran plot appearance (Z-score)
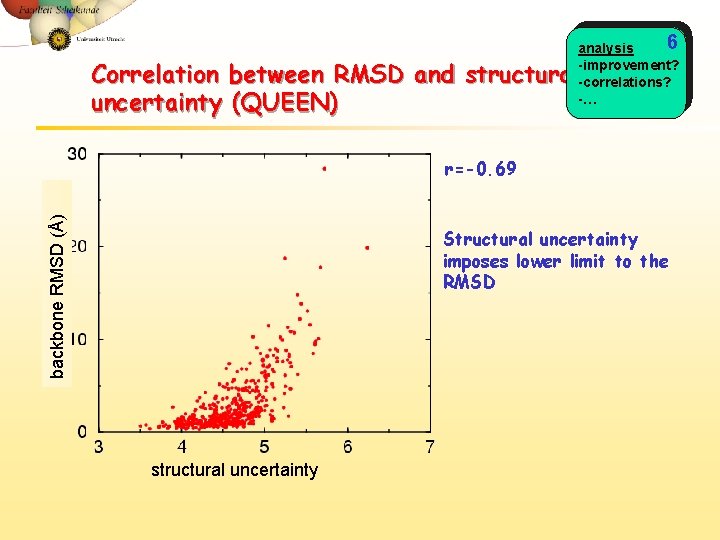
6 analysis -improvement? -correlations? -… Correlation between RMSD and structural uncertainty (QUEEN) backbone RMSD (Å) r=-0. 69 Structural uncertainty imposes lower limit to the RMSD structural uncertainty

Conclusions I • • NMR-STAR files made consistent for 545 out of ± 1700 entries Protocols and scripts available for recalculation in CYANA and CNS Validation database available for testing of new protocols Improvement compared to original data: 1 standard deviation closer to X-ray db • violations in original data do no limit recalculation effort • refinement in water required • 5 % no improvement: data missing

Conclusions II • Correlations higher after recalculation and refinement, though most of them still weak • Highest correlation: precision vs. Ramachandran score & structural uncertainty (QUEEN)

Acknowledgements • • • Utrecht University Alexandre Bonvin Rob Kaptein EBI Cambridge Wim Vranken CESG/BMRB Jurgen Doreleijers Zachary Miller Eldon Ulrich John Markley Radboud University Nijmegen Chris Spronk Sander Nabuurs RIKEN Japan Peter Güntert Institut Pasteur Paris Michael Nilges
- Slides: 26