Recombination and genetic variation models and inference Simon

Recombination and genetic variation – models and inference Simon Myers Department of Statistics, Oxford
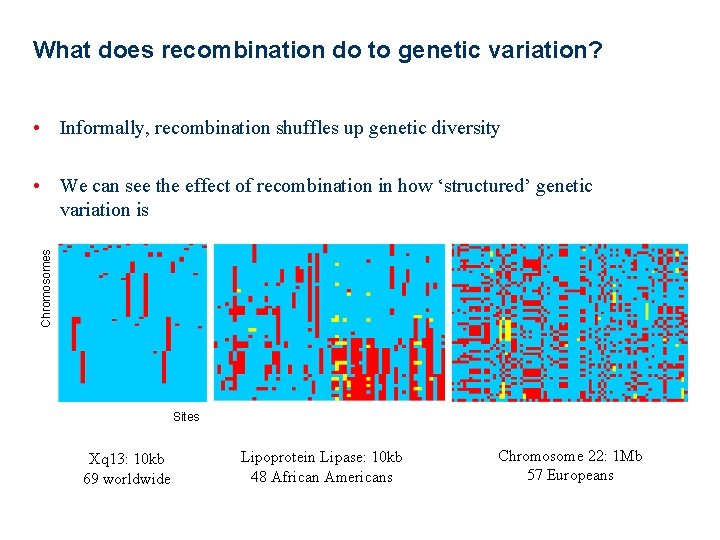
What does recombination do to genetic variation? • Informally, recombination shuffles up genetic diversity Chromosomes • We can see the effect of recombination in how ‘structured’ genetic variation is Sites Xq 13: 10 kb 69 worldwide Lipoprotein Lipase: 10 kb 48 African Americans Chromosome 22: 1 Mb 57 Europeans
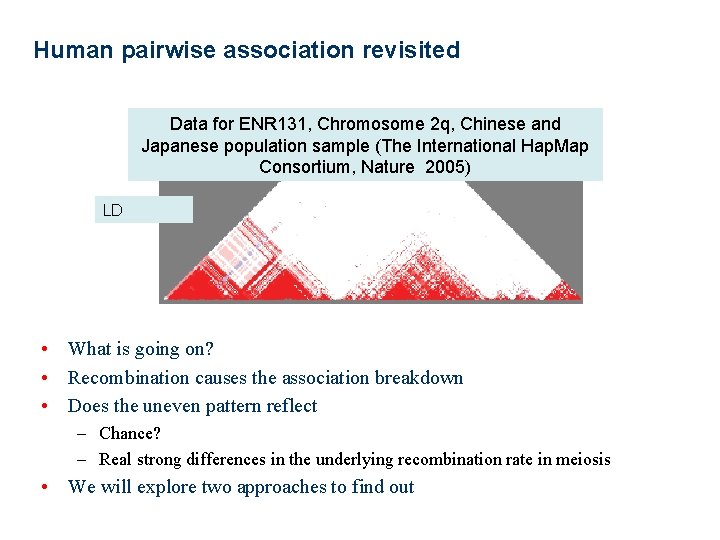
Human pairwise association revisited Data for ENR 131, Chromosome 2 q, Chinese and Japanese population sample (The International Hap. Map Consortium, Nature 2005) LD • What is going on? • Recombination causes the association breakdown • Does the uneven pattern reflect – Chance? – Real strong differences in the underlying recombination rate in meiosis • We will explore two approaches to find out
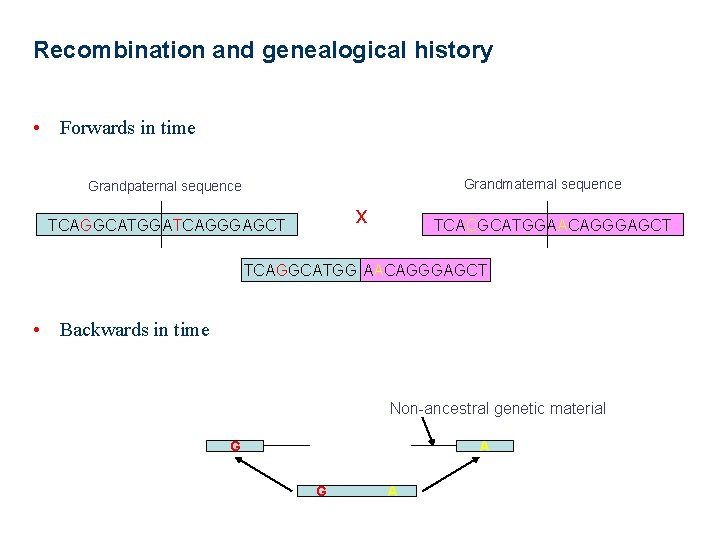
Recombination and genealogical history • Forwards in time Grandmaternal sequence Grandpaternal sequence x TCAGGCATGGATCAGGGAGCT TCACGCATGGAACAGGGAGCT TCAGGCATGG AACAGGGAGCT • Backwards in time Non-ancestral genetic material G A
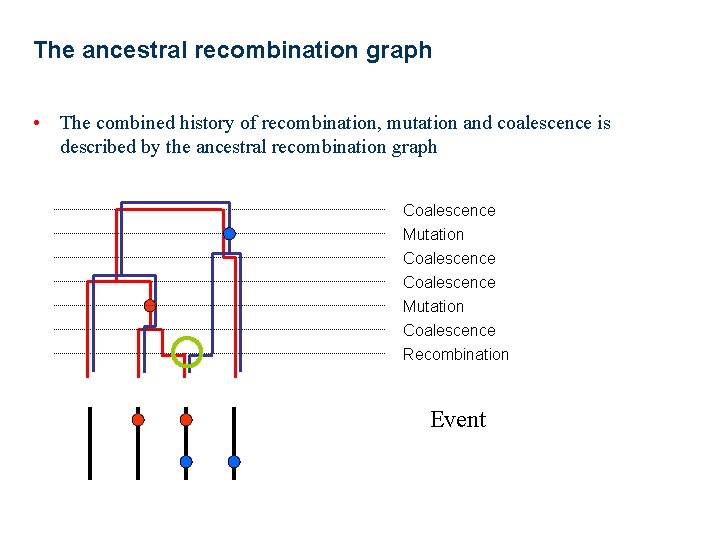
The ancestral recombination graph • The combined history of recombination, mutation and coalescence is described by the ancestral recombination graph Coalescence Mutation Coalescence Recombination Event
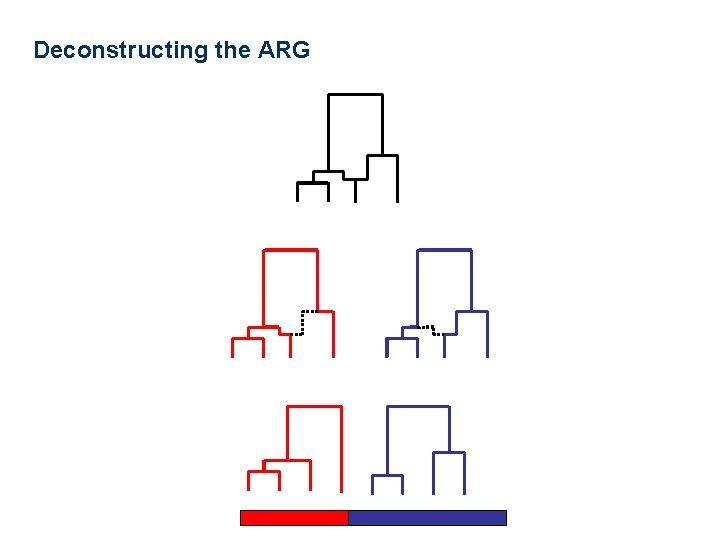
Deconstructing the ARG
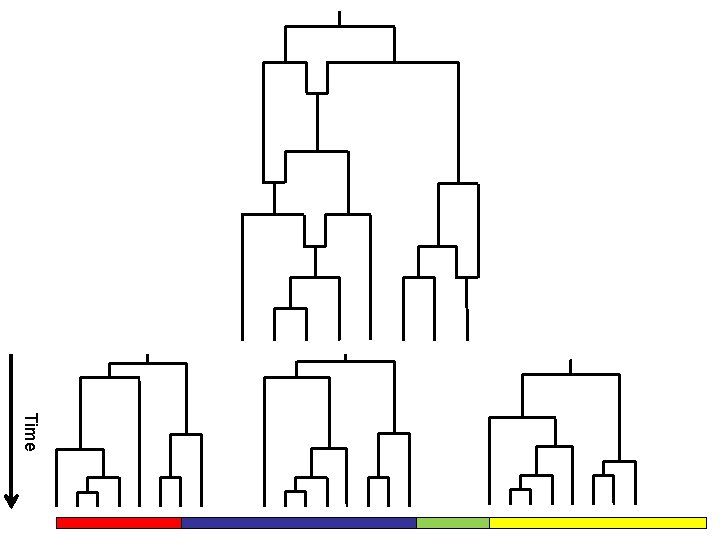
Time

Learning about recombination • Just like there is a true genealogy underlying a sample of sequences without recombination, there is a true ARG underlying samples of sequences with recombination • We can consider nonparametric and parametric ways of learning about recombination • There are several useful nonparametric ways of learning about recombination which we will consider first – These really only apply to species, such as humans, where we can be fairly sure that most SNPs are the result of a single ancestral mutation event – This is formally called the infinite sites model

Why use a non-parametric approach? • Non-parametric approaches require few assumptions about evolution • The infinite sites model, and that’s it! • We can attempt to learn features of the history of a sample based only on this assumption – Robust inference – Identify – “detect” the recombination events that shaped our sample – Clustering of multiple events in a region could signal a high underlying rate • Some drawbacks to this approach
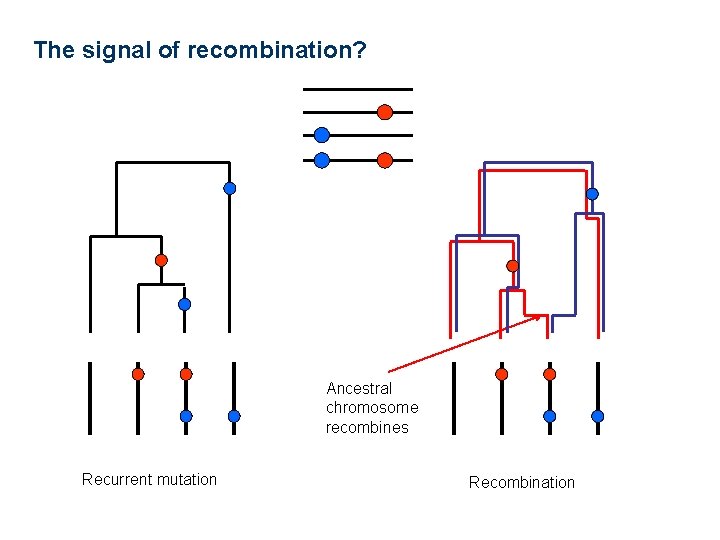
The signal of recombination? Ancestral chromosome recombines Recurrent mutation Recombination
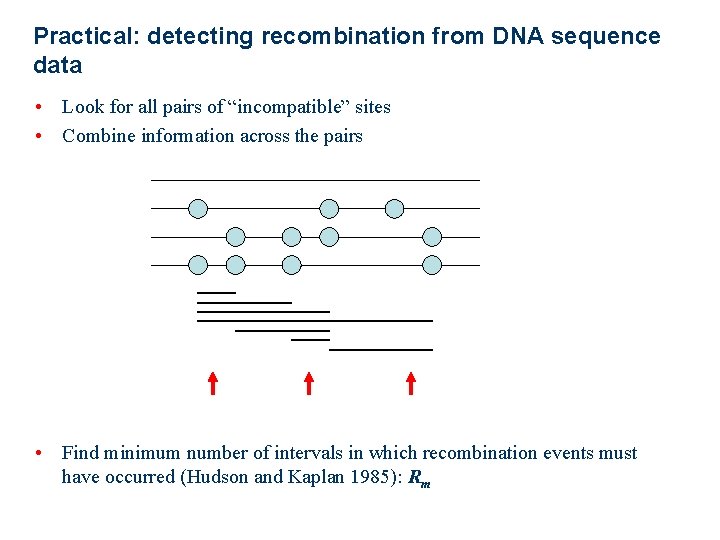
Practical: detecting recombination from DNA sequence data • Look for all pairs of “incompatible” sites • Combine information across the pairs • Find minimum number of intervals in which recombination events must have occurred (Hudson and Kaplan 1985): Rm

Recombination and genetic variation – models and inference, part II Simon Myers Department of Statistics, Oxford
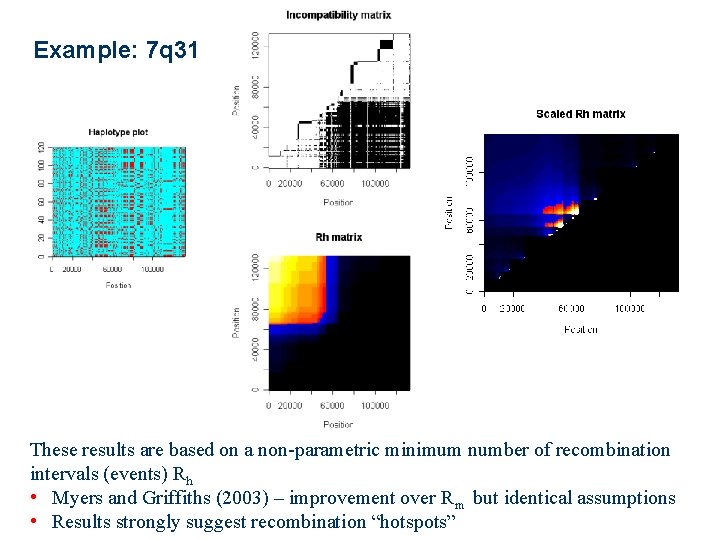
Example: 7 q 31 These results are based on a non-parametric minimum number of recombination intervals (events) Rh • Myers and Griffiths (2003) – improvement over Rm but identical assumptions • Results strongly suggest recombination “hotspots”
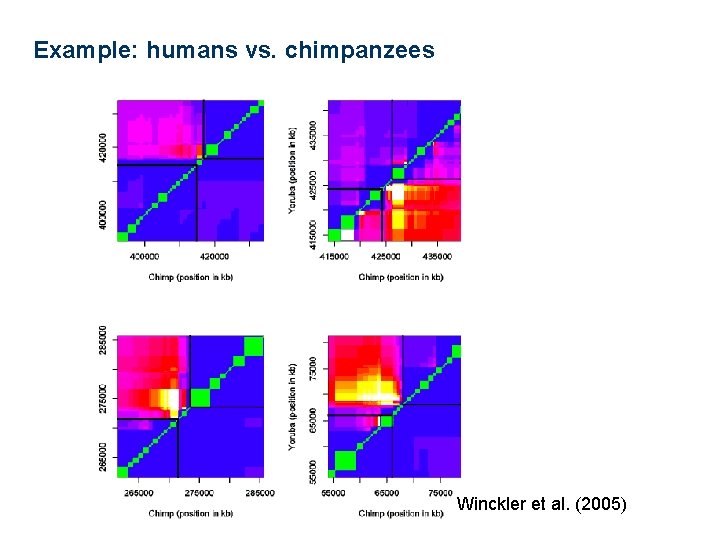
Example: humans vs. chimpanzees Winckler et al. (2005)

Why use parametric approaches? • The infinite-sites model is not applicable to all species HIV Subtype B (2 kb segment) HIV Subtype C (2 kb segment) • There are many more recombination events in the history of the sample than the non-parametric methods can ever detect – Lack of mutations in the right places – Recombination events completely undetectable

Modelling recombination • Model-based approaches to learning about recombination allow us to ask more detailed questions than nonparametric approaches – What is the rate of recombination (as opposed to just the number of events) – Is the rate of recombination across a region constant? – Does gene A have a higher recombination rate than gene B? – What patterns of genetic diversity might I expect to see in other samples from the same (or different) population? • We need a model!

Adding recombination to the coalescent • Each generation, the probability of recombination between two loci is r, working in scaled time, this means that recombination occurs at rate r/2 per sequence where r = 4 Ner • Recombination, mutation and coalescence occur independently: – Coalescence occurs as a Poisson process with rate n(n-1)/2 – Recombination occurs as a Poisson process with rate nr/2 – Mutations on edges added as a Poisson process with rate nq/2 • The time until the next recombination or coalescence event is also a Poisson process with rate nr /2+ n(n-1)/2, and the probability that this next event is a recombination is
Recombination in non-ancestral material • Once a region has recombined, further recombination can occur in both ancestral lineages • However, recombination in non-ancestral DNA cannot in anyway influence patterns of diversity (under a neutral model) • We usually ignore such recombination events in the coalescent X X
Simulating histories with recombination • www. coalescent. dk
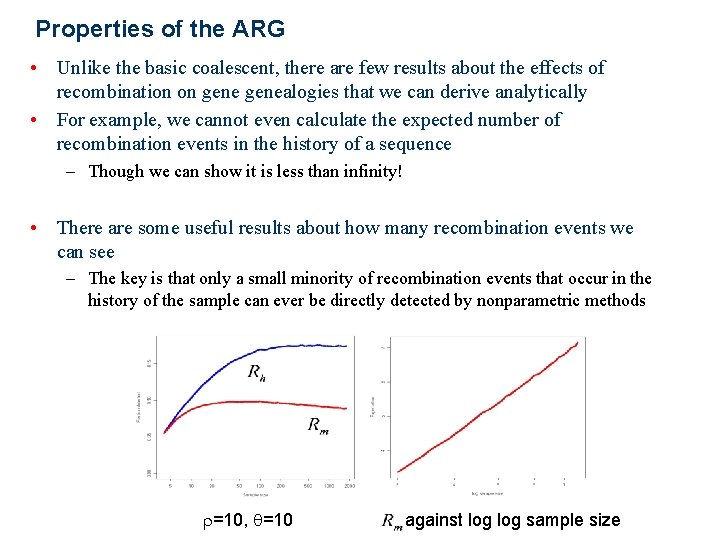
Properties of the ARG • Unlike the basic coalescent, there are few results about the effects of recombination on genealogies that we can derive analytically • For example, we cannot even calculate the expected number of recombination events in the history of a sequence – Though we can show it is less than infinity! • There are some useful results about how many recombination events we can see – The key is that only a small minority of recombination events that occur in the history of the sample can ever be directly detected by nonparametric methods r=10, q=10 against log sample size

Estimating the population recombination rate • The ideal inference procedure would calculate the likelihood of the data – Need to allow recombination rate to vary • …. but full-likelihood inference is effectively impossible for anything but the simplest data sets (and models) • We need alternatives – Calculate the probability of some summary of the data (like ABC) – Approximate the coalescent model – Approximate the likelihood • The composite likelihood of Hudson (2001) approximates the likelihood of the full data by the product of the likelihoods for pairs of sites – Not the real likelihood! – Fast to calculate – Allows a variable recombination rate
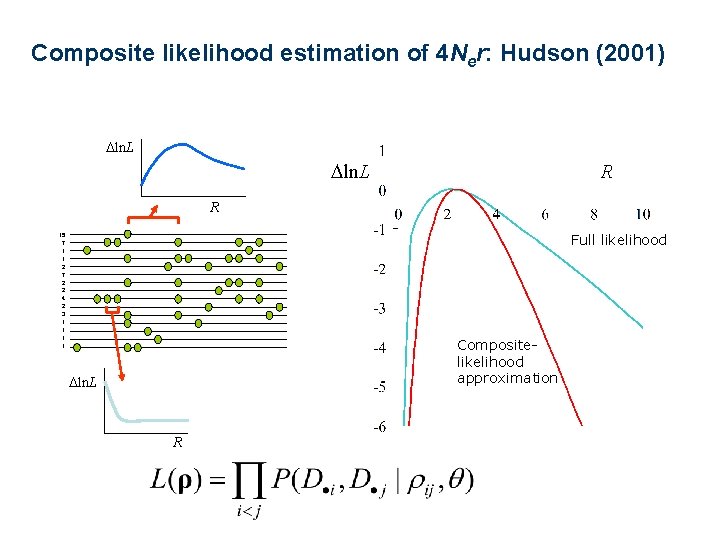
Composite likelihood estimation of 4 Ner: Hudson (2001) Dln. L R R Full likelihood 15 7 1 1 2 7 2 2 4 2 3 1 1 Compositelikelihood approximation Dln. L R
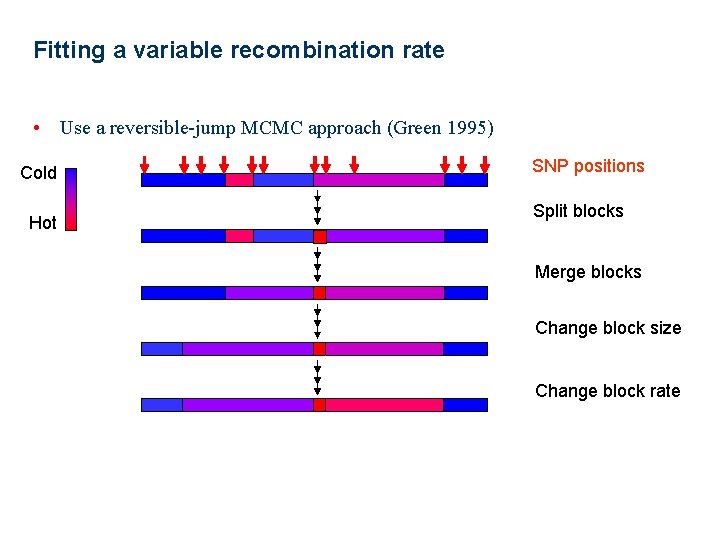
Fitting a variable recombination rate • Use a reversible-jump MCMC approach (Green 1995) Cold Hot SNP positions Split blocks Merge blocks Change block size Change block rate

Acceptance rates Composite likelihood ratio Hastings ratio Ratio of priors Jacobian of partial derivatives relating changes in parameters to sampled random numbers • Include a prior on the number of change points that encourages smoothing
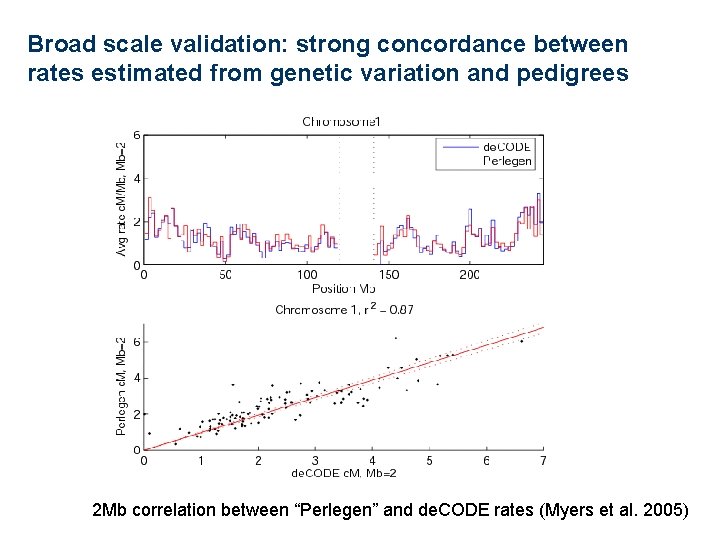
Broad scale validation: strong concordance between rates estimated from genetic variation and pedigrees 2 Mb correlation between “Perlegen” and de. CODE rates (Myers et al. 2005)
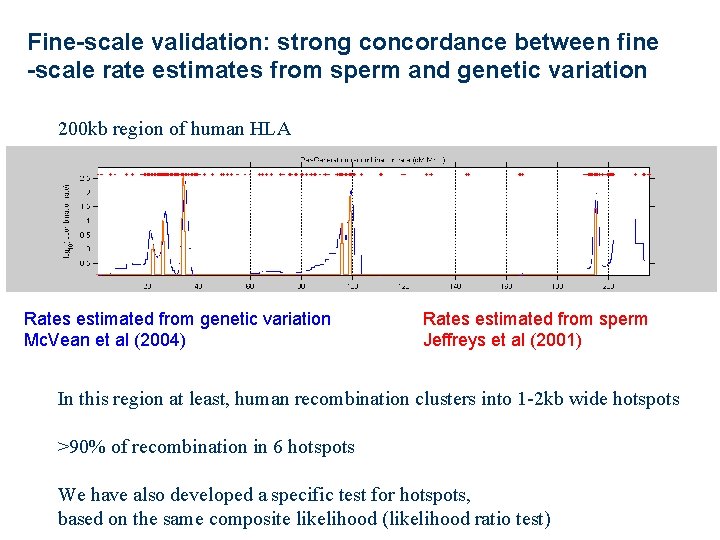
Fine-scale validation: strong concordance between fine -scale rate estimates from sperm and genetic variation 200 kb region of human HLA Rates estimated from genetic variation Mc. Vean et al (2004) Rates estimated from sperm Jeffreys et al (2001) In this region at least, human recombination clusters into 1 -2 kb wide hotspots >90% of recombination in 6 hotspots We have also developed a specific test for hotspots, based on the same composite likelihood (likelihood ratio test)
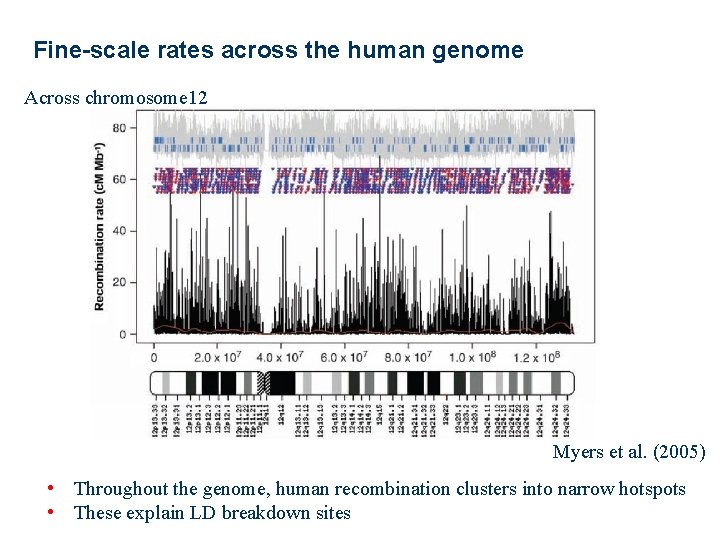
Fine-scale rates across the human genome Across chromosome 12 Myers et al. (2005) • Throughout the genome, human recombination clusters into narrow hotspots • These explain LD breakdown sites
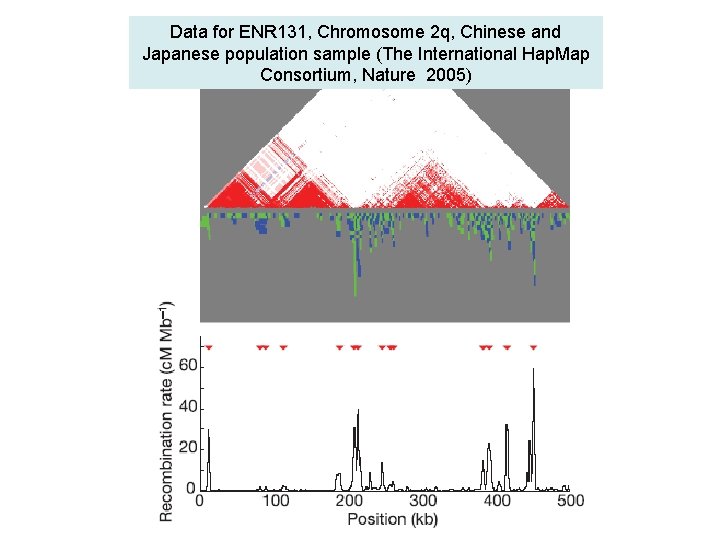
Data for ENR 131, Chromosome 2 q, Chinese and Japanese population sample (The International Hap. Map Consortium, Nature 2005)

Summary • Both non-parametric and model based approaches allow us to ask detailed questions about recombination from population genetic data • Recombination can be incorporated within the coalescent framework • The population recombination rate, r=4 Ner, is the key quantity in determining the effect of recombination on genetic variation • Efficiently estimating recombination rates within a coalescent framework is difficult, but approximate methods have proved a powerful approach • Such methods have allowed us to successfully learn about recombination rates in humans and other species, and reveal “hotspots” across genomes
- Slides: 29