Recombinant protein expression in E coli Bio 4600

Recombinant protein expression in E. coli Bio 4600 2003 Vigdis Lauvrak
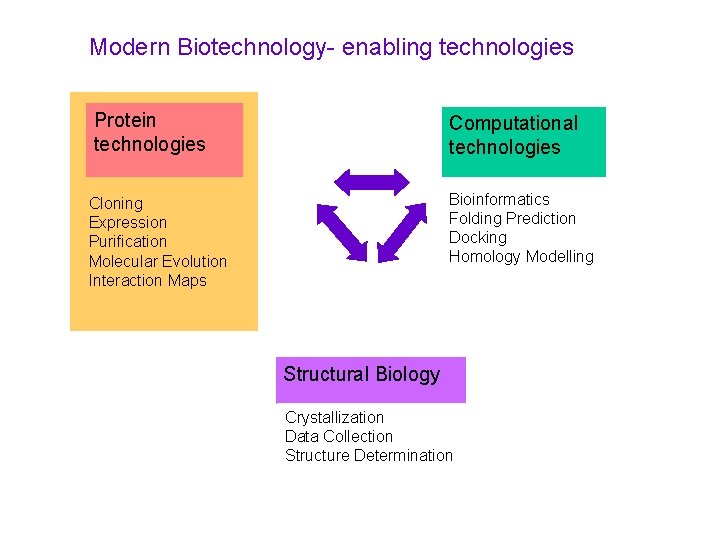
Modern Biotechnology- enabling technologies Protein technologies Computational technologies Bioinformatics Folding Prediction Docking Homology Modelling Cloning Expression Purification Molecular Evolution Interaction Maps Structural Biology Crystallization Data Collection Structure Determination
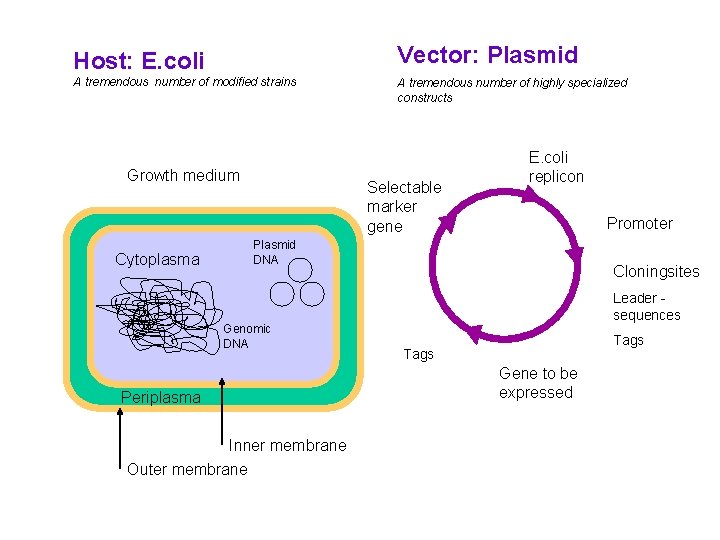
Host: E. coli Vector: Plasmid A tremendous number of modified strains A tremendous number of highly specialized constructs Growth medium Cytoplasma Selectable marker gene E. coli replicon Promoter Plasmid DNA Genomic DNA Periplasma Inner membrane Outer membrane Cloningsites Leader sequences Tags Gene to be expressed

Major options to be considered: • Gene dosage (copy number) • Level of expression • Which compartment to harvest from • Tags for purification, improvement of stability and solubility • Codon usage E. coli: recombinant protein • Purpose of expression: Large scale industrial/or analytical levels?

A replicom is a genetic unit consisting of an origin of DNA replication and its associated elements. origin p. BR 322 col. E 1 p. UC p. MOB 45 p. ACYC p. SC 101 Replicon p. MB 1 col. E 1 mod p. MB 1 p. KN 402 p 15 A p. SC 101 copy number 15 -20 500 -700 15 -118 18 -22 5

Gene dosage Medium to high copy number plasmids • Relaxed replication • Random distribution • Relatively low loss: Continously growth and toxic genes/gene products will lead to plasmid loss. Increased plasmid stability: Selectable markers • Genes for antibiotic resistance • Complementation: An essential chromosomal gene is deleted or mutated an intact copy or a supressor is suplied in trans. • Genes or repressors that lead to cell death upon plasmid loss. Duplication of genomic inserts Increased gene dosage in E-coli genome: • Rec. A dupllication of insert (Olson et al. 1998) : 15 --40 copies (may be unstable without a selectable marker). • Tn 1545 site specific recombination (Peredelchuck and Bennett 1997) - time consuming

Control of expression level Desired: High expression level (10 -30% or more of produced protein) Observed: Many proteins may are toxic at high doses. Solution: Regulation of expression
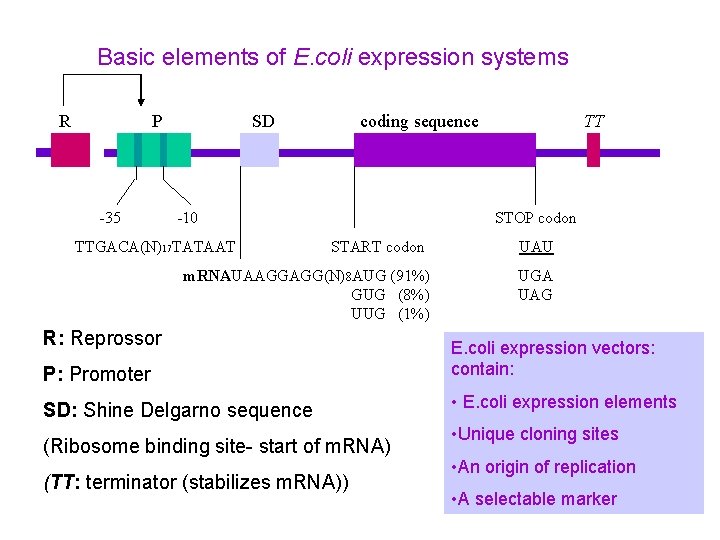
Basic elements of E. coli expression systems R P -35 SD coding sequence -10 TTGACA(N)17 TATAAT TT STOP codon START codon m. RNAUAAGGAGG(N)8 AUG (91%) GUG (8%) UUG (1%) R: Reprossor P: Promoter SD: Shine Delgarno sequence (Ribosome binding site- start of m. RNA) (TT: terminator (stabilizes m. RNA)) UAU UGA UAG E. coli expression vectors: contain: • E. coli expression elements • Unique cloning sites • An origin of replication • A selectable marker

Level of regulation depends on the promoter • The lac operon- the paradigm of protein regulation in E. coli: lactose/ IPTG-induction (derepression) • lac. UV 5 (leaky): IPTG • tac and trc synthetic versions of lac (tighter): IPTG • T 7 -late promoter : Depends on T 7 polymerase • PL promoter- Lambda CI regulated, tight regulation • csp. A: Cold chock induction • pho. A, trp and ara. BAD (PBAD): Nutritional inducible • tet: Tetracycline inducible • Signal dependent promoters: p. H, oxygen conc. , osmolarity etc. (Inexpensive large scale production)
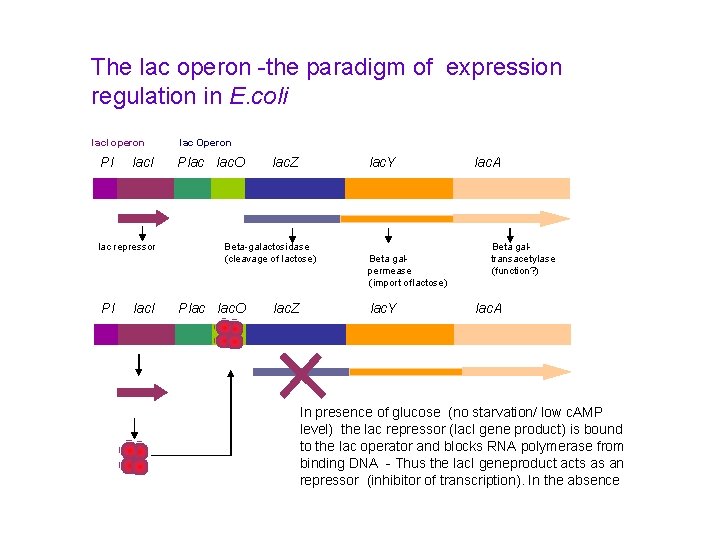
The lac operon -the paradigm of expression regulation in E. coli lac. I operon Pl lac. I lac repressor Pl lac. I lac Operon Plac lac. O lac. Z lac. Y Beta-galactosidase (cleavage of lactose) Plac lac. O lac. Z Beta galpermease (import of lactose) lac. Y lac. A Beta galtransacetylase (function? ) lac. A In presence of glucose (no starvation/ low c. AMP level) the lac repressor (lac. I gene product) is bound to the lac operator and blocks RNA polymerase from binding DNA - Thus the lac. I geneproduct acts as an repressor (inhibitor of transcription). In the absence
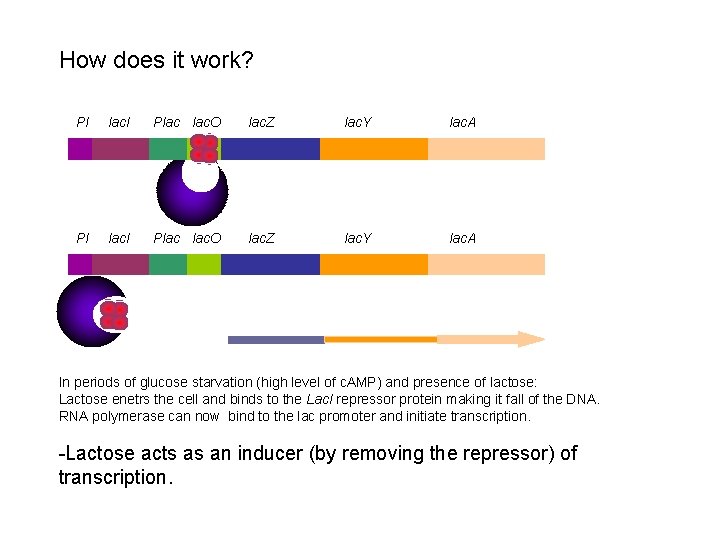
How does it work? Pl lac. I Plac lac. O lac. Z lac. Y lac. A In periods of glucose starvation (high level of c. AMP) and presence of lactose: Lactose enetrs the cell and binds to the Lac. I repressor protein making it fall of the DNA. RNA polymerase can now bind to the lac promoter and initiate transcription. -Lactose acts as an inducer (by removing the repressor) of transcription.

The lac promoter of E. coli expression vectors: • Induction is performed with IPTG which acts as a synthetic lactose analogue that binds the lac. I gene product. • Presence of glucose further prevents transcription from the lac promoter. IPTG Pl lac. I PX lac. O The CI binding site (lac operator) can be combinde with various other promoter sequences to give improved regulation.
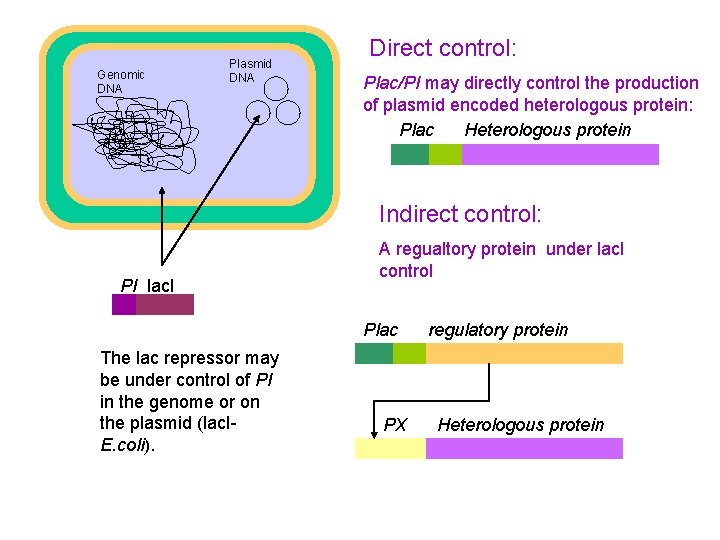
Genomic DNA Plasmid DNA Direct control: Plac/PI may directly control the production of plasmid encoded heterologous protein: Plac Heterologous protein Indirect control: PI lac. I A regualtory protein under lac. I control Plac The lac repressor may be under control of PI in the genome or on the plasmid (lac. IE. coli). PX regulatory protein Heterologous protein
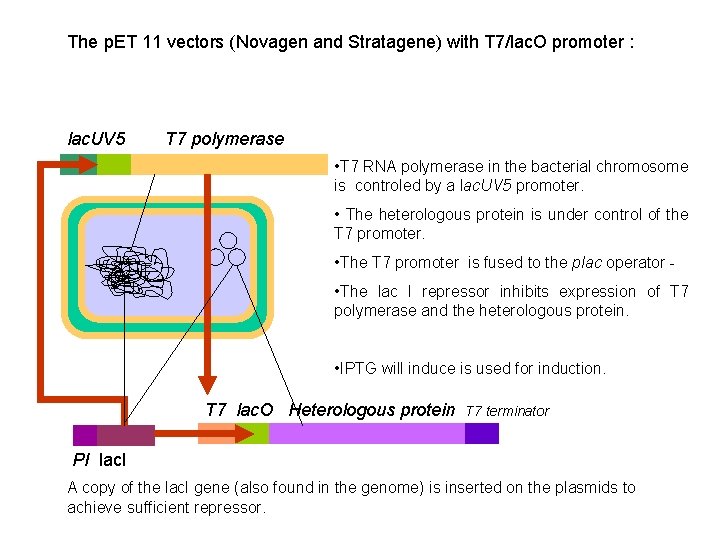
The p. ET 11 vectors (Novagen and Stratagene) with T 7/lac. O promoter : lac. UV 5 T 7 polymerase • T 7 RNA polymerase in the bacterial chromosome is controled by a lac. UV 5 promoter. • The heterologous protein is under control of the T 7 promoter. • The T 7 promoter is fused to the plac operator • The lac I repressor inhibits expression of T 7 polymerase and the heterologous protein. • IPTG will induce is used for induction. T 7 lac. O Heterologous protein T 7 terminator PI lac. I A copy of the lac. I gene (also found in the genome) is inserted on the plasmids to achieve sufficient repressor.
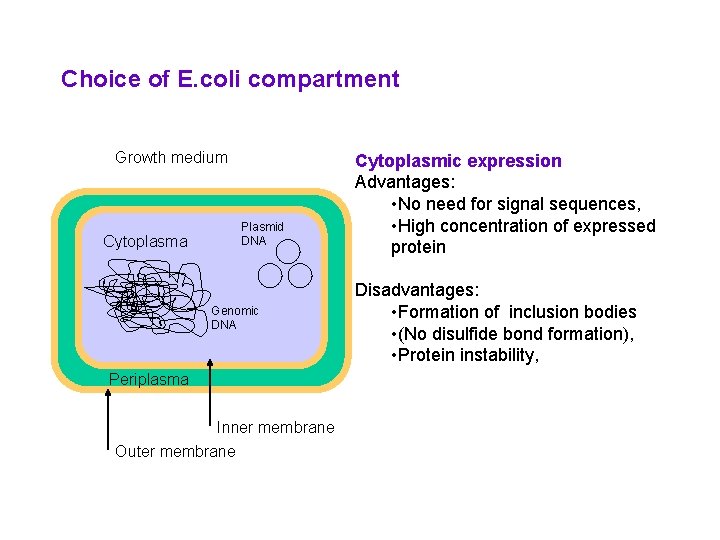
Choice of E. coli compartment Growth medium Cytoplasma Plasmid DNA Genomic DNA Periplasma Inner membrane Outer membrane Cytoplasmic expression Advantages: • No need for signal sequences, • High concentration of expressed protein Disadvantages: • Formation of inclusion bodies • (No disulfide bond formation), • Protein instability,

Periplasm Advantages • Improved folding (no inclusion body formation) • Disulfide bridge formation (may be enhanced by the presence of Dsb. A and Dsb. B proteins) • Fewer proteins and possible leakage to growth medium may facilitate purification. • Less protein degradation. Disadvantages. • Low protein concentration due to inefficient transport and small compartment Solution • Thight regulation of expression • Molecular chaperones (protein specific) • Temperature down shift after induction- less formation of inclusion bodies). Growth media No efficient system for direct transport to growth media. Leakage from periplasm is often used.

Common problems encountered with E. coli expression system: The desired protein may be: Unstable, toxic, insoluble, form inclusion bodies, uncorect folded, depend on disulfide bridges, and active only with postranslational modifications : glycosylation, phosphorylation and amidation. Solutions: Choice of a suitable E. coli strain, tags, fusions and leader sequences can solve many problems including disulfide bridge formation, but proteins that need correct postranslational modifications as underlined above have to be produced in Eucaryotic systems.

Solutions: • Thight regulation of expression • Coexpression of molecular chaperones (protein specific) • Reduction of rate of protein synthesis (lower growth rate by temperature down shift after induction) • Fusion moiteties may increase folding, solubility and resistance to proteolysis. • Use of protease deficient E. coli strains • Use of thioredoxin reductase (trx. B) og glutatione reductase (gor) double mutants may give disulfide bridge formation in cytosol • Periplasmic expression




Characteristics of suitable induction sensitive promotors • High strength • Tight regulation • Simple and cost effective induction: Basic research: IPTG (lactose analogue (toxic)) Tetracycline Thermal Industrial production of theraeutics: Thermal Chemical Nutrional
- Slides: 22