RECOMBINANT DNA TECHNOLOGY DNA SEQUENCING Dr I Viji

RECOMBINANT DNA TECHNOLOGY, DNA SEQUENCING Dr. I. Viji Margaret Assistant Professor, PG Dept. of Zoology, Sarah Tucker College, Tirunelveli

� Introduction � Recombinant DNA technology, joining together of DNA molecules from two different species that are inserted into a host organism to produce new genetic combinations that are of value to science, medicine, agriculture, and industry. Since the focus of all genetics is the gene, the fundamental goal of laboratory geneticists is to isolate, characterize, and manipulate genes. Although it is relatively easy to isolate a sample of DNA from a collection of cells, finding a specific gene within this DNA sample can be compared to finding a needle in a haystack. Consider the fact that each human cell contains approximately 2 metres (6 feet) of DNA. Therefore, a small tissue sample will contain many kilometres of DNA. However, recombinant DNA technology has made it possible to isolate one gene or any other segment of DNA, enabling researchers to determine its nucleotide sequence, study its transcripts, mutate it in highly specific ways, and reinsert the modified sequence into a living organism.

� Steps involved in recombinant DNA technology � Stage 1. Isolation of the Genetic Material (DNA) � Stage 2. Cutting of DNA at Specific Locations � Stage 3. Isolation of Desired DNA Fragment � Stage # 4. Amplification of Gene of Interest using PCR � Stage 5. Ligation of DNA Fragment into a Vector � Stage 6. Insertion of Recombinant DNA into the Host Cell/Organisms

� Stage 1. Isolation of the Genetic Material (DNA): � Nucleic acid is the genetic material, which is present in all living organisms. In majority of organisms, this is present in the form of deoxyribonucleic acid (DNA). DNA must be present into pure form, i. e. , free from other macro-molecules (like proteins, RNA, enzymes, etc. ) in order to cut the DNA with restrictor enzymes. � Isolation of genetic material (DNA) is carried out in the following steps: � a) Since the DNA is enclosed within the membranes, so, in order to release DNA along with other macro-molecules such as proteins, polysaccharides and lipids, bacterial cells/plant or animal tissues are treated with the enzyme lysozyme (bacteria), cellulose (plant cells), chitinase (fungus), respectively. � (b) RNA can be removed by the treatment with ribonuclease, whereas proteins can be removed by the treatment with protease. � (c) Other molecules can be removed by appropriate treatments and ultimately purified DNA will precipitate out, after the addition of chilled ethanol. This can be seen as collection of fine threads in the suspension.

� Stage 2. Cutting of DNA at Specific Locations: � Restriction enzyme digestions are performed by incubating purified DNA molecules with the restriction enzyme. This is done at the optimal conditions for that specific enzyme. � Stage 3. Isolation of Desired DNA Fragment: � Using agarose gel electrophoresis, the activity of the restriction enzymes can be checked. Since, the DNA is negatively charged, it moves towards the positive electrode or anode and DNA tends to separates out in this process. After that the desired DNA fragment is eluted out.
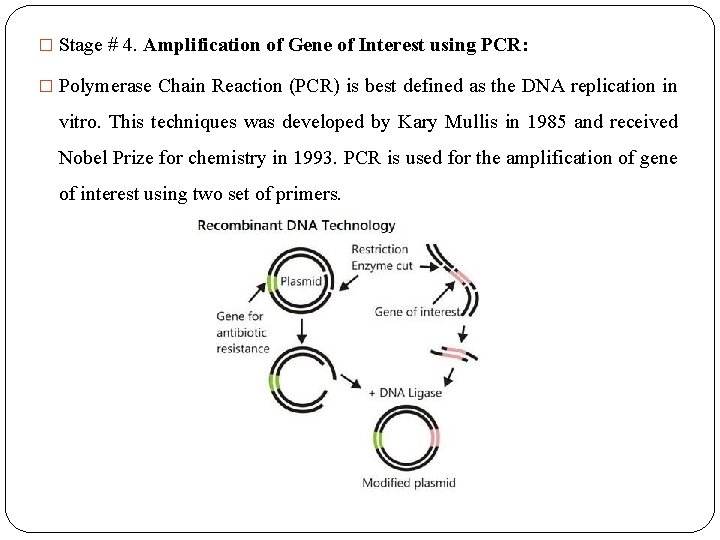
� Stage # 4. Amplification of Gene of Interest using PCR: � Polymerase Chain Reaction (PCR) is best defined as the DNA replication in vitro. This techniques was developed by Kary Mullis in 1985 and received Nobel Prize for chemistry in 1993. PCR is used for the amplification of gene of interest using two set of primers.

� Stage 5. Ligation of DNA Fragment into a Vector: � This process requires a vector DNA and a source DNA. In order to obtain sticky ends, both of these should cut with the same endonuclease. After which both are ligated by mixing vector DNA, gene of interest and enzyme DNA ligase to form the recombinant DNA/hybrid DNA. � Stage 6. Insertion of Recombinant DNA into the Host Cell/Organisms: � This can occur by several methods, before which the recipient cells are made competent to receive the DNA. If a recombinant DNA bearing gene for resistance to an antibiotic (e. g. , ampicillin) is transferred into E. coli cells, the host cells become transformed into ampicillin resistant cells. � The ampicillin resistance gene in this case is called a selectable marker. When transformed cells are grown on agar plates containing ampicillin, only transformants will grow and others will die.

� Stage 7. Obtaining or Culturing the Foreign Gene Product: � When you insert a piece of alien DNA into a cloning vector and transfer it into a bacterial cell, the alien DNA gets multiplied. The ultimate aim is to produce a desirable protein expression. The expressions of the foreign gene or genes in host cells involve understanding of many technical details. � If the protein encoded gene is expressed in the heterologous host, it is called recombinant protein. The cells harbouring cloned genes of interest are grown on small scale in the laboratory. These cell cultures are used for extracting the desired protein using various separation techniques.

� DNA sequencing � Definition � DNA sequencing is the process of determining the sequence of nucleotides (As, Ts, Cs, and Gs) in a piece of DNA. � Nucleotides are not the only determinants of phenotypes, but are essential to their formation. Each individual and organism has a specific nucleotide base sequence.

� Sanger sequencing, the target DNA is copied many times, making fragments of different lengths. Fluorescent “chain terminator” nucleotides mark the ends of the fragments and allow the sequence to be determined. � Sanger sequencing: The chain termination method � Regions of DNA up to about 900900 base pairs in length are routinely sequenced using a method called Sanger sequencing or the chain termination method. Sanger sequencing was developed by the British biochemist Fred Sanger and his colleagues in 1977. � In the Human Genome Project, Sanger sequencing was used to determine the sequences of many relatively small fragments of human DNA. (These fragments weren't necessarily 900900 bp or less, but researchers were able to "walk" along each fragment using multiple rounds of Sanger sequencing. ) The fragments were aligned based on overlapping portions to assemble the sequences of larger regions of DNA and, eventually, entire chromosomes.

� Although genomes are now typically sequenced using other methods that are faster and less expensive, Sanger sequencing is still in wide use for the sequencing of individual pieces of DNA, such as fragments used in DNA cloning or generated through polymerase chain reaction (PCR).

� Method of Sanger sequencing � The DNA sample to be sequenced is combined in a tube with primer, DNA polymerase, and DNA nucleotides (d. ATP, d. TTP, d. GTP, and d. CTP). The four dyelabeled, chain-terminating dideoxy nucleotides are added as well, but in much smaller amounts than the ordinary nucleotides. � The mixture is first heated to denature the template DNA (separate the strands), then cooled so that the primer can bind to the single-stranded template. Once the primer has bound, the temperature is raised again, allowing DNA polymerase to synthesize new DNA starting from the primer. DNA polymerase will continue adding nucleotides to the chain until it happens to add a dideoxy nucleotide instead of a normal one. At that point, no further nucleotides can be added, so the strand will end with the dideoxy nucleotide.

� This process is repeated in a number of cycles. By the time the cycling is complete, it’s virtually guaranteed that a dideoxy nucleotide will have been incorporated at every single position of the target DNA in at least one reaction. That is, the tube will contain fragments of different lengths, ending at each of the nucleotide positions in the original DNA (see figure below). The ends of the fragments will be labeled with dyes that indicate their final nucleotide.

� After the reaction is done, the fragments are run through a long, thin tube containing a gel matrix in a process called capillary gel electrophoresis. Short fragments move quickly through the pores of the gel, while long fragments move more slowly. As each fragment crosses the “finish line” at the end of the tube, it’s illuminated by a laser, allowing the attached dye to be detected. � The smallest fragment (ending just one nucleotide after the primer) crosses the finish line first, followed by the next-smallest fragment (ending two nucleotides after the primer), and so forth. Thus, from the colors of dyes registered one after another on the detector, the sequence of the original piece of DNA can be built up one nucleotide at a time. The data recorded by the detector consist of a series of peaks in fluorescence intensity, as shown in the chromatogram above. The DNA sequence is read from the peaks in the chromatogram.

�Uses and limitations �Sanger sequencing gives high-quality sequence for relatively long stretches of DNA (up to about 900900 base pairs). It's typically used to sequence individual pieces of DNA, such as bacterial plasmids or DNA copied in PCR. �However, Sanger sequencing is expensive and inefficient for larger-scale projects, such as the sequencing of an entire genome or metagenome (the “collective genome” of a microbial community). For tasks such as these, new, large-scale sequencing techniques are faster and less expensive.

THANK YOU
- Slides: 16