Reaction discovery enabled by DNAtemplated synthesis and in

Reaction discovery enabled by DNA-templated synthesis and in vitro selection Matthew W. Kanan, Mary M. Rozenman, Kaori Sakurai, Thomas M. Snyder & David R. Liu Nature, vol. 431, 2004 Presented by Seok, Ho-Sik Bio. Intelligence Lab

What they did? n A reaction discovery approach that uses DNA-templated organic synthesis and in vitro selection to simultaneously evaluate many combination of different substrates for bondforming reactions in a single solution Bio. Intelligence Lab
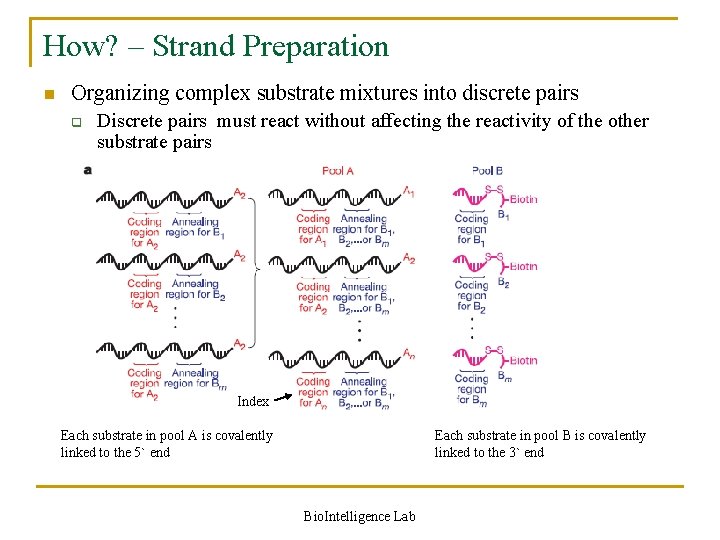
How? – Strand Preparation n Organizing complex substrate mixtures into discrete pairs q Discrete pairs must react without affecting the reactivity of the other substrate pairs Index Each substrate in pool A is covalently linked to the 5` end Each substrate in pool B is covalently linked to the 3` end Bio. Intelligence Lab

Strand Preparation in detail n n Recent developments in DNA-templated organic synthesis indicate that DNA annealing can organize many substrates in a single solution into DNA sequence-programmed pairs Two pools of DNA-linked substrates, with n substrates in pool A and m substrates in pool B q q Each substrate in pool A is covalently linked to the 5` end of a set of DNA oligonucleotides containing one ‘coding region’ (uniquely identifying that substrate) and one of m different ‘annealing regions’ Each of the m substrates in pool B is attached to the 3` end of an oligonucleotide containing a coding region that uniquely identifies the substrate and complements one of the m annealing regions in pool A Bio. Intelligence Lab
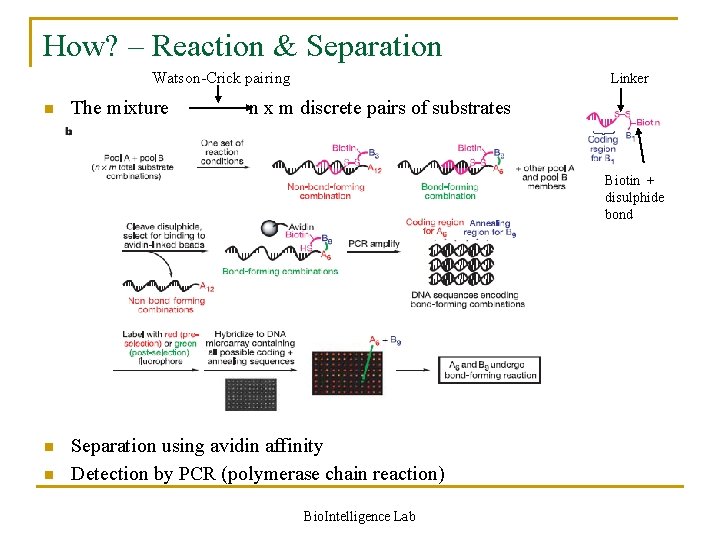
How? – Reaction & Separation Watson-Crick pairing n The mixture Linker n x m discrete pairs of substrates Biotin + disulphide bond n n Separation using avidin affinity Detection by PCR (polymerase chain reaction) Bio. Intelligence Lab
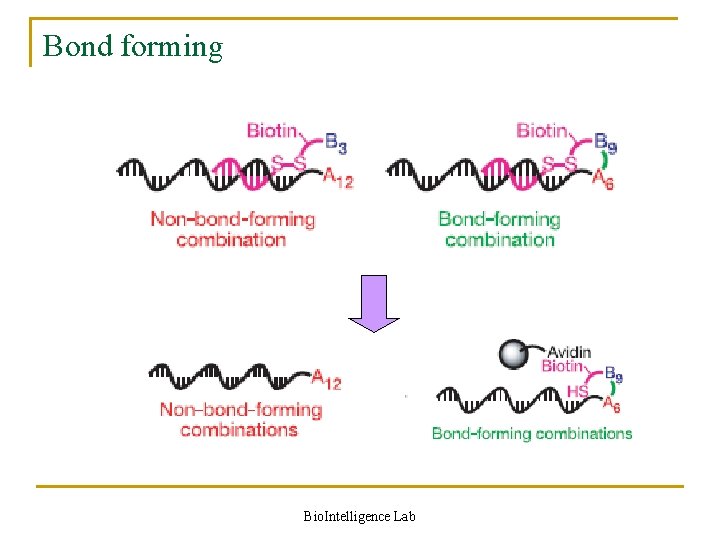
Bond forming Bio. Intelligence Lab

Reaction in detail n Role of Watson-Crick base pairing q When pools A and B are combined in a single aqueous solution, Watson –Crick base pairing organizes the mixture into n x m discrete pairs of substrates attached to complementary sequences q Only substrates linked to complementary oligonucleotides experience effective molarities in the millimolar range n Possibility of interference by the DNA structure q Minimized by using long and flexible substrate–DNA linkers Bio. Intelligence Lab
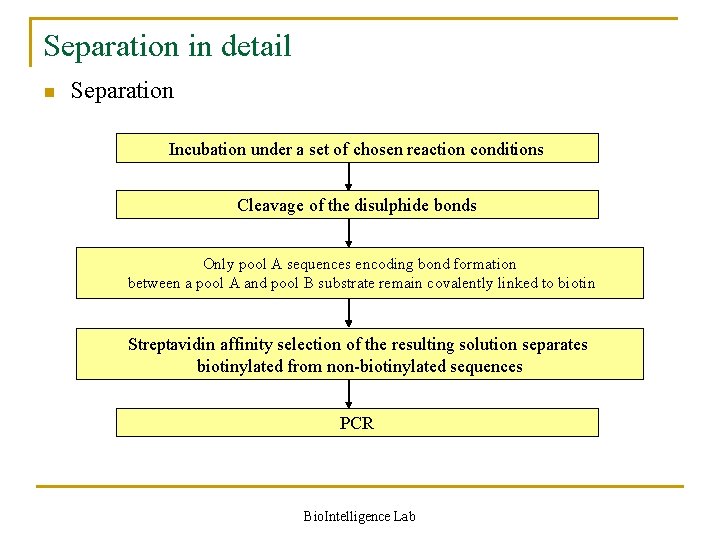
Separation in detail n Separation Incubation under a set of chosen reaction conditions Cleavage of the disulphide bonds Only pool A sequences encoding bond formation between a pool A and pool B substrate remain covalently linked to biotin Streptavidin affinity selection of the resulting solution separates biotinylated from non-biotinylated sequences PCR Bio. Intelligence Lab
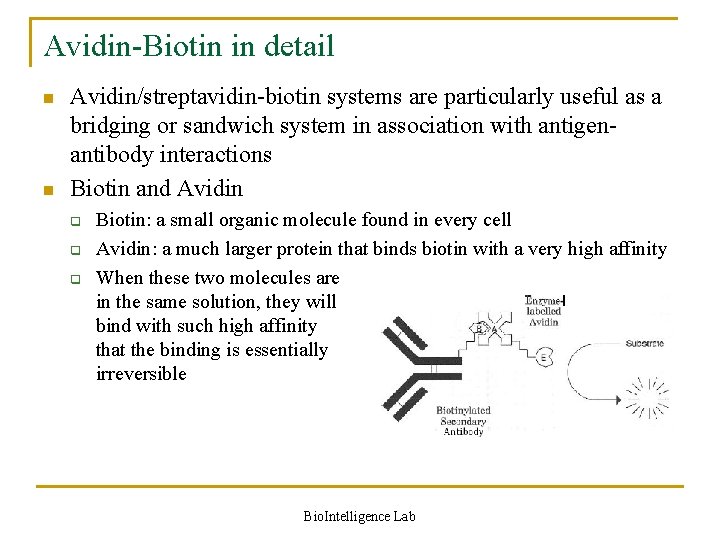
Avidin-Biotin in detail n n Avidin/streptavidin-biotin systems are particularly useful as a bridging or sandwich system in association with antigenantibody interactions Biotin and Avidin q q q Biotin: a small organic molecule found in every cell Avidin: a much larger protein that binds biotin with a very high affinity When these two molecules are in the same solution, they will bind with such high affinity that the binding is essentially irreversible Bio. Intelligence Lab
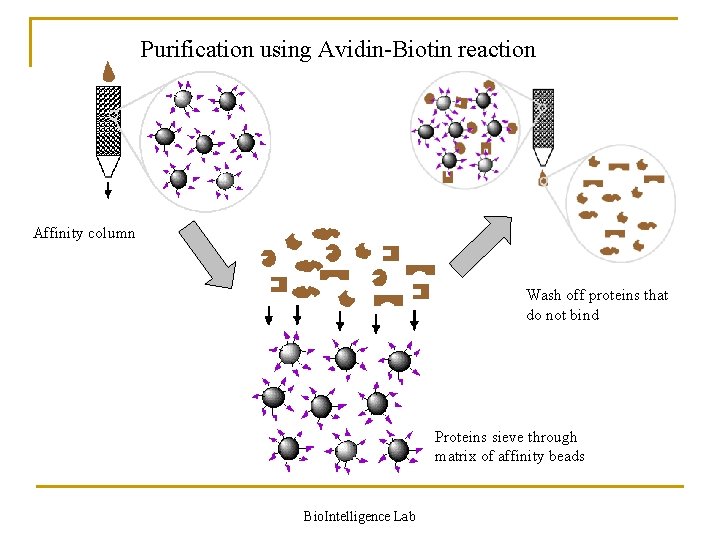
Purification using Avidin-Biotin reaction Affinity column Wash off proteins that do not bind Proteins sieve through matrix of affinity beads Bio. Intelligence Lab
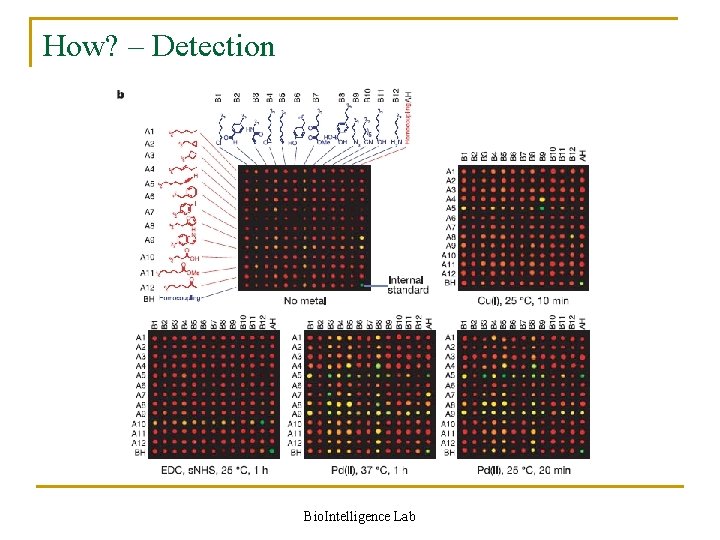
How? – Detection Bio. Intelligence Lab

Detection in detail n Capturing q n Amplification q n Sequences encoding bond-forming substrate pairs were amplified by PCR with a DNA primer labeled with the cyanine fluorophore Cy 3 Comparison q n Sequences encoding bond-forming substrate pairs were captured with streptavidin-linked magnetic particles Aliquot of the pool A sequences before selection was amplified by PCR with a Cy 5 -labelled primer Scoring q The ratios of Cy 3 (green) to Cy 5 (red) fluorescence for all array locations were calculated and ordered by rank, and spots with green/red fluorescence ratios significantly higher than the majority of spots (in the experiments below, ratios above 1. 5) were considered to be positive Bio. Intelligence Lab
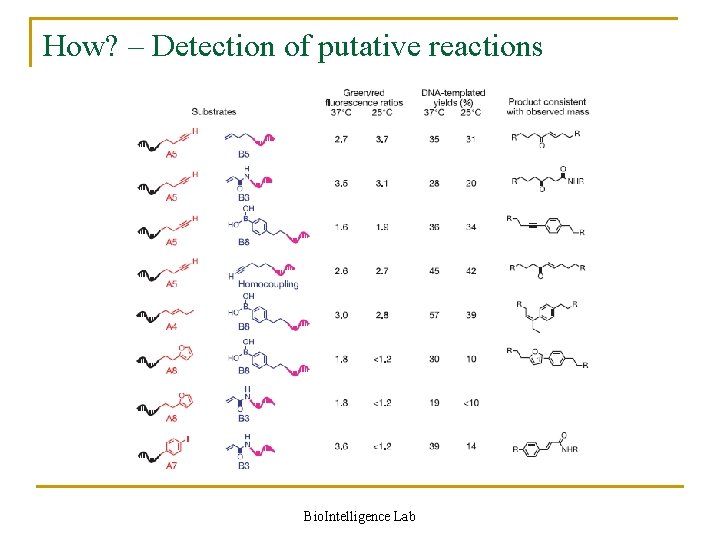
How? – Detection of putative reactions Bio. Intelligence Lab

Detection of putative reactions in detail Denaturing Polya. Arylamide Gel Electrophoresis (PAGE) analysis Matrix Assisted Laser Desorption Ionization–Time-of-Flight (MALDI–TOF) mass spectrometry n n PAGE q Comparison of strand positions MALDI-TOF q Once inside the ionisation source the sample molecules are ionised, because ions are easier to manipulate than neutral molecules q These ions are extracted into the analyser region of the mass spectrometer where they are separated according to their mass-to-charge ratios Bio. Intelligence Lab

DNA computing and the reaction discovery n Another method for providing strands n Another method for selecting strands q n Expensive but precise way of detecting q n In addition to affinity, we can use cleavage of bonds MALDI–TOF mass spectrometry Possibility of advanced DNA computing q Practical limitation of 10 k q Possibility of DNA as a just template or catalyst q Using product of DNA-templated reaction for computing Bio. Intelligence Lab
- Slides: 15