Rational vaccine design using reverse vaccinology database data

Rational vaccine design using reverse vaccinology, database data analysis, and literature mining Yongqun “Oliver” He Unit for Laboratory Animal Medicine Department of Microbiology and Immunology Center for Computational Medicine and Bioinformatics Comprehensive Cancer Center University of Michigan Medical School Ann Arbor, MI 48109

Outline I. Introduction II. Vaxign: Rational vaccine design using reverse vaccinology III. Rational vaccine design based on database data analysis IV. Rational vaccine design by literature mining V. Summary and Discussion

Infectious diseases resulted in 9. 2 million deaths in 2013 (about 17% of all deaths) ― Global Burden of Disease (GBD) Study 2013 “We must make this the decade of vaccines. ” -- Bill Gates (2010)

How to rationally design vaccines? Can bioinformatics help?

Outline I. Introduction Vaxign: Rational vaccine design using reverse vaccinology III. Rational vaccine design based on database data analysis IV. Rational vaccine design by literature mining V. Summary and Discussion

Reverse Vaccinology (RV) Vaccine development starts with bioinformatics genome sequence analysis Dr. Rino Rappuoli Pioneer of Reverse Vaccinology Neisseria meningitidis serogroup B meningococcal B (Men. B) disease Pizza M, et al. Science. 2000 10; 287(5459): 1816 -20. Milestone: Novartis’s vaccine Bexsero approved by Europe & USA - Johri et al. Nat Rev Microbiol. 2006 Dec; 4(12): 932– 42. - Rappuoli R. Curr Opin Microbiol. 2000 Oct; 3(5): 445 -50.

Vaxign: Vaccine Design System • The 1 st web-based reverse vaccinology system • Predicted criteria by Vaxign pipeline: o o o Antigen subcellular localization (original criterion) Adhesin (critical for host invasion) Transmembrane domains Epitope binding to MHC class I and class II Pathogen gene ortholog analysis Sequence similarities to human, mouse and/or pig proteins • Freely available: http: //www. violinet. org/vaxign Reference: He Y, Xiang Z, Mobley HLT. Vaxign: the first web-based vaccine design program for reverse vaccinology and an application for vaccine development. Journal of Biomedicine and Biotechnology. Volume 2010 (2010), Article ID 297505, 15 pages. [PMID: 20671958]

Two Forms of Vaxign Usage Pre-computed data query • • • Pre-computed results > 200 genomes in database Easy to query Dynamic analysis • • Runtime execution Can analyze up to 500 proteins at one time
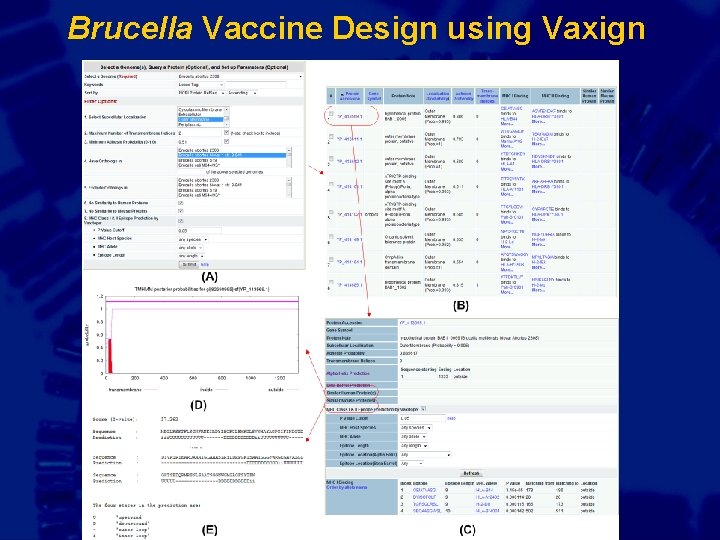
Brucella Vaccine Design using Vaxign Gr

Vaxign case study: Brucellosis is a zoonotic disease caused by bacterium Brucella Brucellosis in livestock Human brucellosis Global Livestock Production and Health Atlas, Animal Production and Health Division, 2006, FAO Ref: Pappas G, et al. Lancet Infect Dis. 2006 Feb; 6(2): 91 -9. >500, 000 new human cases worldwide annually

Brucella Genomes for Vaccine Design Species Host B. abortus cattle, human B. canis dogs, foxes, coyotes, human B. melitensis sheep, goats, human B. neotomae desert wood rats B. ovis rams B. suis pigs, human 4 new Brucella spp. : B. cetacean B. pinnipedia seal B. microti voles B. inopinata unknown • 10 Brucella spp. • 4 pathogenic to humans • 10 genomes in NCBI Ref. Seq database • > 20 new genomes were sequenced • Question: Can we use Vaxign to predict Brucella vaccine candidates?

Vaxign Prediction of Brucella vaccine targets *, represents five genomes from virulent B. abortus strain 9 -941, B. melitensis strains 16 M and ATCC 23457, and B. suis strains 1330 and ATCC 23445. **, represents the genome from B. ovis strain ATCC 25840; The indicated two proteins are Omp 2 b (YP_414102. 1) and Omp 31 -1 (YP_414995. 1). Ref. He Y, Xiang Z. Bioinformatics analysis of Brucella vaccines and vaccine targets using VIOLIN. Immunome Research. 2010 Sep 27; 6 Suppl 1: S 5.

Experimental Verification of Vaxignpredicted Brucella vaccine candidates • Experiments by Dr. Thomas Ficht’s lab in Texas A&M: o 5 Vaxign-predicted proteins expressed & tested o 2 proteins with the highest predicted adhesin scores conferred an invasive phenotype to the non-invasive BL -21 E. coli strain in alveolar epithelial cells o All proteins elicited Ig. G antibody production in Brucellaexposed goats, mouse and humans. o 3 proteins elicited protection in mice against challenges • This is a great demonstration of Vaxign usefulness Reference: Gomez et al. Immunogenicity and protective efficacy of Brucella outer membrane proteins identified via a reverse vaccinology approach. 2012 Annual Brucellosis Conference, Chicago, IL, USA.
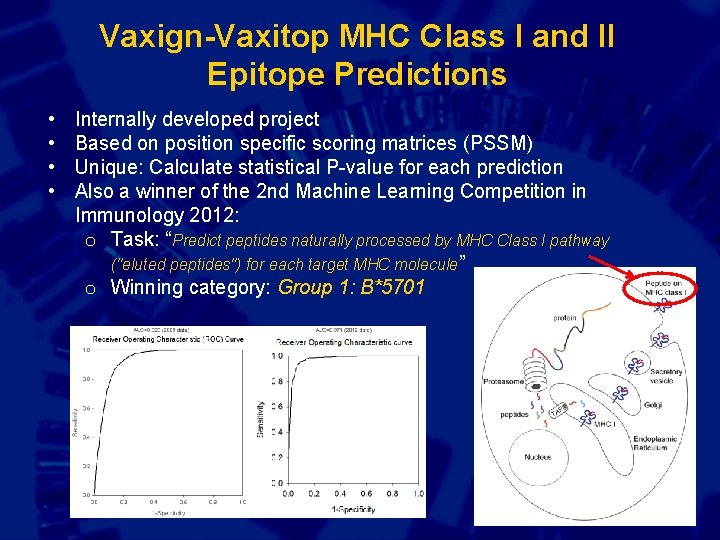
Vaxign-Vaxitop MHC Class I and II Epitope Predictions • • Internally developed project Based on position specific scoring matrices (PSSM) Unique: Calculate statistical P-value for each prediction Also a winner of the 2 nd Machine Learning Competition in Immunology 2012: o Task: “Predict peptides naturally processed by MHC Class I pathway ("eluted peptides") for each target MHC molecule” o Winning category: Group 1: B*5701

Other Vaxign Applications • Uropathogenic E. coli (UPEC) vaccine prediction o Collaboration with Dr. Harry Mobley here in UM • African Swine Fever Virus vaccine development o Collaboration with Mangkey Bounpheng et al. o All predicted & expressed proteins were found immunogenic o More experiments under way • Human herpes simplex virus vaccine design • Francisella tularensis vaccine design • Vaxign can also be used for virulence factor prediction (e. g. , Campylobacter fetus, Corynebacterium diphtheriae) • ~50 citations of Vaxign in Google Scholar References: • He Y, Xiang Z, Mobley HLT. Vaxign: the first web-based vaccine design program for reverse vaccinology and an application for vaccine development. J Biomed Biotechnol. 2010; 2010: 297505. • Xiang Z, He Y. Genome-wide prediction of vaccine targets for human herpes simplex viruses using Vaxign reverse vaccinology. BMC Bioinformatics. In press.

What’s More about Vaccine Design Vaxign is to predict protective antigens, which can be improved using other data, including database and literature data In addition Need to design more than protective antigens, e. g. , DNA vaccine plasmid, vaccine vector, adjuvant, gene for deletion, …

Outline I. Introduction II. Vaxign: Rational vaccine design using reverse vaccinology III. Rational vaccine design based on database data analysis IV. Rational vaccine design by literature mining V. Summary and Discussion

VIOLIN: Vaccine Investigation and Online Information Network Comprehensive vaccine R & D database & data analysis system http: //www. violinet. org • • • >3400 vaccines (licensed, clinical trial, or lab-verified) 200 pathogens or non-inf. diseases (e. g. , cancer) ~1200 pathogen genes: o o Protegen: Protective antigens Virmugens: Genes mutated to make live attenuated vaccines >1, 000 host genes Databases of different vaccine components o o Vaxjo: Vaccine adjuvants Vaxvec: Vaccine vectors DNAVax. DB: DNA vaccines and DNA vaccine plasmids Vaxvec: Vaccine vectors > 2, 000 papers manually annotated

Protegen: Protective Antigen Database • Protective antigens (Ags): pathogen Ags tested in experimentally verified vaccines • 857 protective Ags collected in Protegen • Enriched patterns among protective Ags found – Secreted and outer membrane proteins favorable, depending on pathogens – Many conserved domains were identified, for example, • Protegen data have been used by many as “Gold Standard” for testing Reference: Yang B, Sayers S, Xiang Z, He Y. Protegen: a web-based protective antigen database and analysis system. Nucleic Acids Research. 2010. 39: D 1073 -8.

Can we predict protective antigens using Protegen database data? • Protegen data can be used to identify enriched patterns, which can guide rational vaccine design • E. g. , from analysis of 270 bacterial protective antigens o o Extracellular or cell wall proteins: 64% for Gram+ Extracellular or outer membrane: 48% for Gram. Adhesins or adhesin-like: 54% G+; 40% GEnriched conserved domains, e. g. , Ton. B • A Support Vector Machine (SVM) classification: 92% true positive rate of sequence-based protection Reference: He Y, Xiang Z. Bioinformatics analysis of bacterial protective antigens in manually curated Protegen database. Procedia in Vaccinology. 2012. Vol 6. Pages 3 -9.

“Virmugen”: virulence factors whose mutants are live attenuated vaccines • • Virmugen. DB: database that collects and analyzes virmugens URL: http: //www. violinet. org/virmugendb Statistics: 225 virmugens from 57 pathogens; 196 vaccines 16 “aro” genes: for aromatic amino acid synthesis • Enriched functions: Nutrient metabolism & Cell membrane formation • Support rational live attenuated vaccine design Reference: Racz R, Chung M, Xiang Z, and He Y. Systematic annotation and analysis of “virmugens” - virulence factors whose mutants can be used as live attenuated vaccines. Vaccine. 2013 Jan 21; 31(5): 797 -805.

Vaxjo: Vaccine adjuvants http: //www. violinet. org/vaxjo • 93 vaccine adjuvants • Used in 379 vaccines Question: Does the information help adjuvant selection in vaccine design?

Vaxvec: Recombinant Vaccine Vectors and recombinant vector vaccines • 59 recombinant vectors • 187 recombinant vector vaccines • Bacterial antigens in Vaxvec o Adhesins: 19% o Compare DNAVax. DB: 30 -39% Microbes used as recombinant vaccine vectors: Question: Does the information help recombinant vector vaccine design?
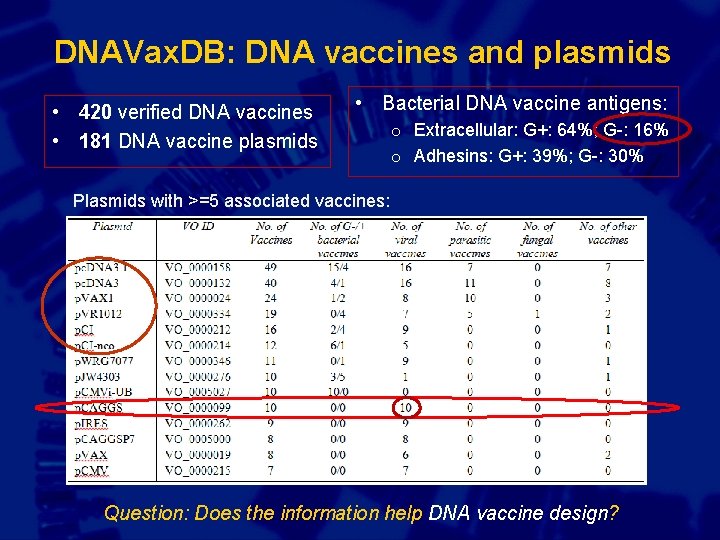
DNAVax. DB: DNA vaccines and plasmids • 420 verified DNA vaccines • 181 DNA vaccine plasmids • Bacterial DNA vaccine antigens: o Extracellular: G+: 64%; G-: 16% o Adhesins: G+: 39%; G-: 30% Plasmids with >=5 associated vaccines: Question: Does the information help DNA vaccine design?

Ex: Develop DNA Vaccines for M. bovis • Mycobacterium bovis: – Aerobic bacterium that can cause tuberculosis, primarily in cattle, also in humans – part of M. tuberculosis complex – Not well studies: in VIOLIN, only BCG and p. CI-Ag 85 B • May use M. tuberculosis or other vaccines information • Plasmid selection: – Commonly used plasmids in M. tuberculosis: p. JW 4303, pc. DNA 3. 1. – Some plasmids are used more in Gram +/- bacteria and viruses • Protective antigens: – Used in M. tuberculosis DNA vaccines: Ag 85 A, Kat. G, MPT 64 find orthologs in M. bovis. – Prediction using Vaxign

Outline I. Introduction II. Vaxign: Rational vaccine design using reverse vaccinology III. Rational vaccine design based on database data analysis IV. Rational vaccine design by literature mining V. Summary and Discussion
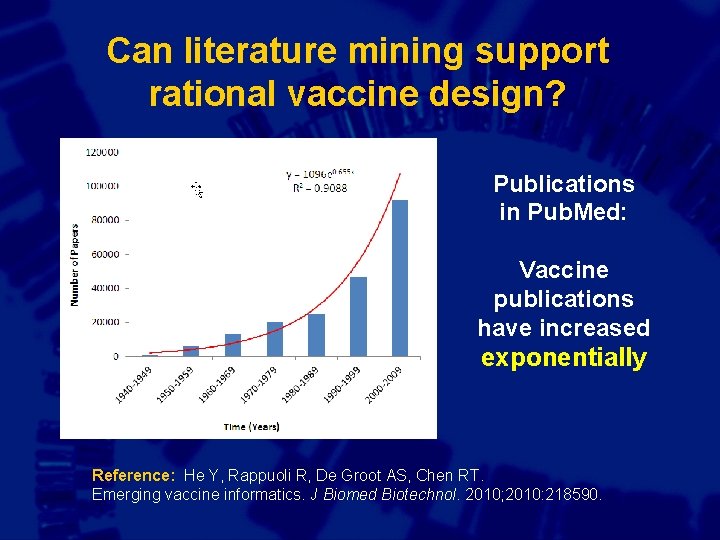
Can literature mining support rational vaccine design? Publications in Pub. Med: Vaccine publications have increased exponentially Reference: He Y, Rappuoli R, De Groot AS, Chen RT. Emerging vaccine informatics. J Biomed Biotechnol. 2010; 2010: 218590.

Ex. Search genes associated with “live attenuated Brucella vaccine” in Pub. Med Method: Default Me. SH-based Pub. Med search

Vaccine Ontology (VO) defines more live attenuated vaccines § VO: Open source ontology for vaccines Inferred hierarchy Asserted hierarchy
Ex. Search genes associated with “live attenuated Brucella vaccine” in Pub. Med Method: VO-based Sci. Miner Pub. Med search

Ex. Search for genes associated with Brucella vaccine in Pub. Med • • • Method: VO-Sci. Miner VO includes and classifies 67 Brucella vaccine ~15, 000 Pub. Med abstracts searched 140 Brucella vaccine related genes identified Functions of these genes studied: ü protective antigens ü virulence factors ü other genes closely related to Brucella. • Some genes useful for vaccine development Reference: Hur J, Xiang Z, Feldman EL, He Y. Ontology-based Brucella vaccine literature indexing and systematic analysis of gene-vaccine association network. BMC Immunology. 12(1): 49 2011 Aug 26. PMID: 21871085.

Summary & Discussion • Vaxign: Vaccine design by predicting protective antigens through bioinformatics analysis of genome sequences (i. e. , reverse vaccinology) • VIOLIN vaccine component databases can support design of vaccine components • Literature mining also supports rational vaccine design • Discussion: What’s more about vaccine design? o Using OMICS data? o More or better software?

Acknowledgements He Lab at UM: U. of Michigan (UM): VO Collaborators: • • • • Junguk Hur • Arzucan Ozgur • Dragomir R. Radev • Brian Athey • Harry Mobley • Barry Smith (U Baffalo) • Lindsay Cowell (UT SW) • Bjoern Peters (LIAI) • Alan Rutternberg (Baffalo) • Melanie Courtot (Canada) • Richard Scheuermann (JCVI) Zuoshuang Xiang Asiyah Yu Lin Andrew Hodges Rebecca L. Racz Samantha Sayers Monica Chung Omar Tibi Mukti A Patel Shelley Zhang Desikan Jagannathan Joseph Alan Ostrow Erica Laura Marcos G. Bill Jourdian Funding: • NIH-NIAID Grant 1 R 01 AI 081062 • NIH/NIAID 1 R 21 AI 057875 • UM CCMB pilot grant • NIH 1 U 54 DA 021519
- Slides: 33