Rama Balakrishnan Ami GO Saccharomyces Genome Database SGD

Rama Balakrishnan Ami. GO Saccharomyces Genome Database (SGD) Stanford University

What is Ami. GO? • Web application that reads from the GO Database (my. SQL) • Allows to – browse the ontologies – view annotations from various species – compare sequences (GOst) • Ontologies are loaded into the database from the gene_ontology. obo file • Annotations are loaded from the gene_association files submitted by the various annotating groups – Only ‘Non-IEA’ annotations are loaded

Ami. GO http: //www. godatabase. org Node has children, can be clicked to view children

Some basics Node has children, can be clicked to view children Node has been opened, can be clicked to close Leaf node or no children Is_a relationship Part_of relationship pie chart summary of the numbers of gene products associated to any immediate descendants of this term in the tree .

Searching the Ontologies
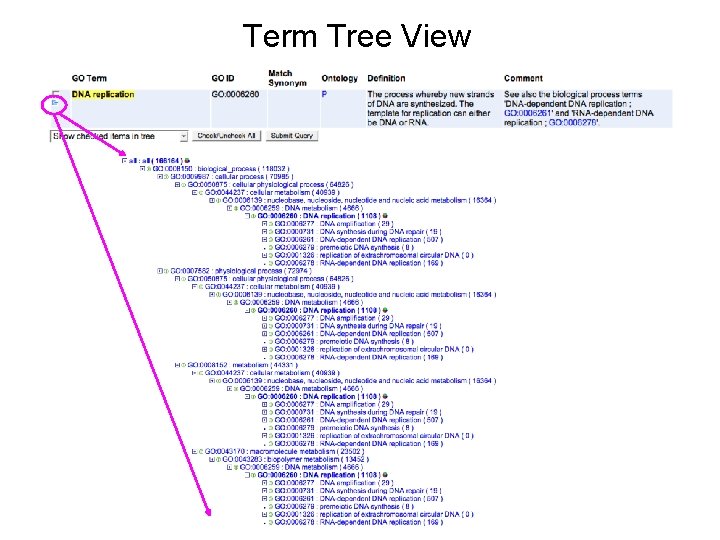
Term Tree View
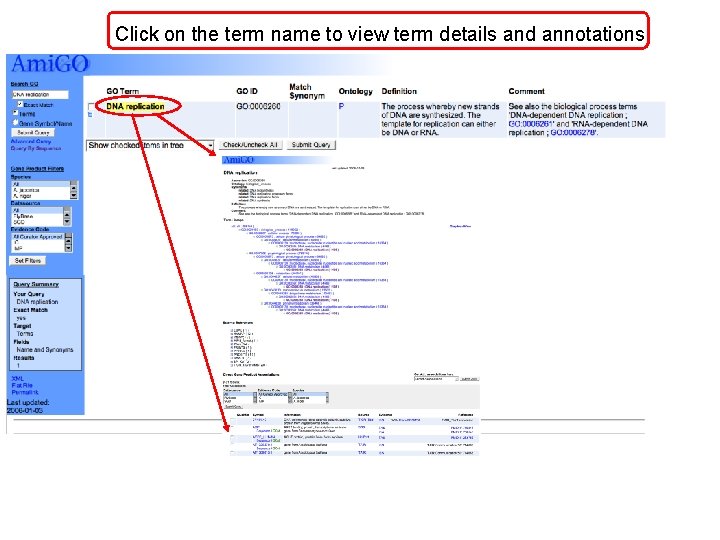
Click on the term name to view term details and annotations
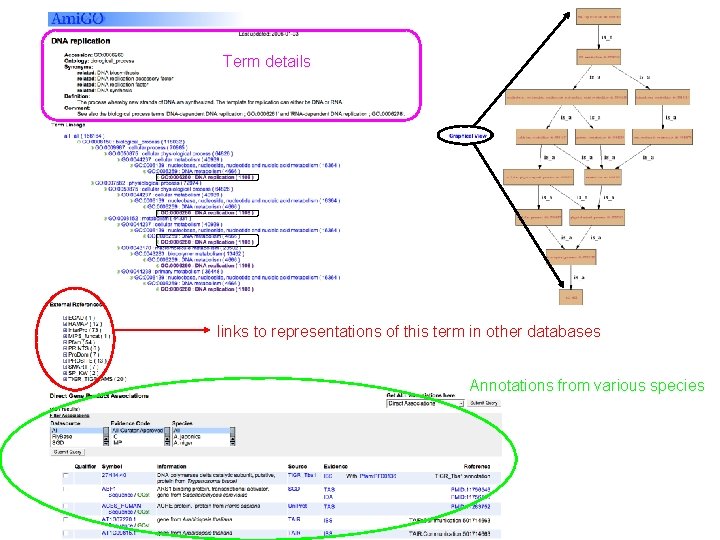
Term details links to representations of this term in other databases Annotations from various species
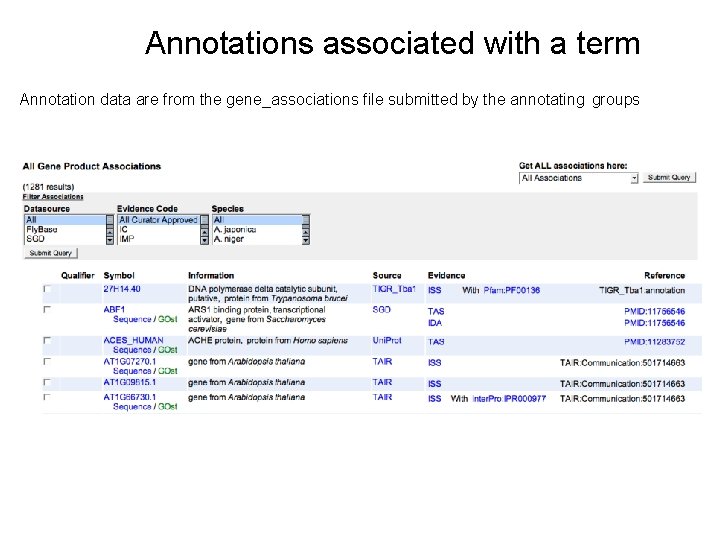
Annotations associated with a term Annotation data are from the gene_associations file submitted by the annotating groups
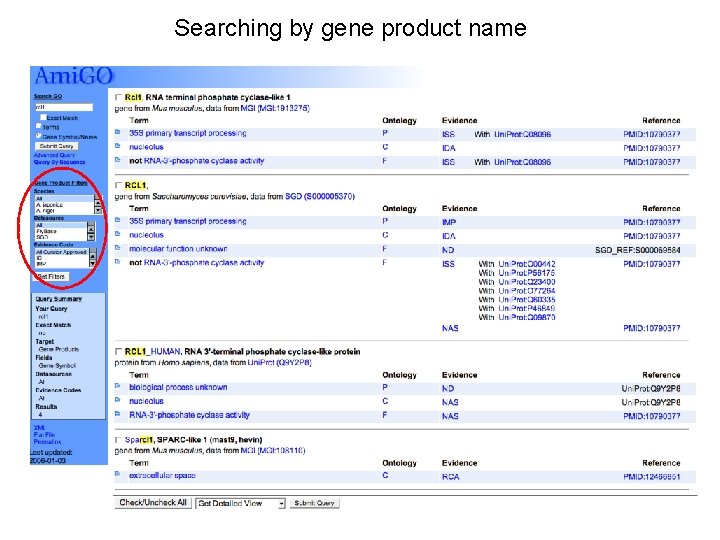
Searching by gene product name
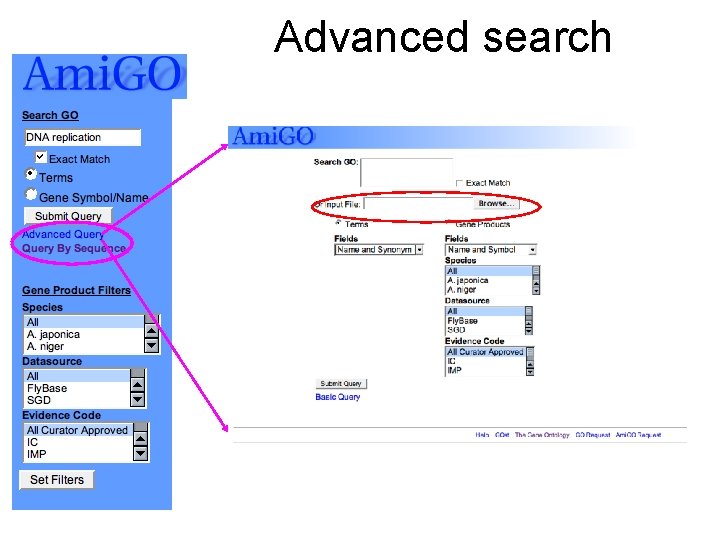
Advanced search

GOST-Gene Ontology bla. ST • • Blast a protein sequence against all gene products that have a GO annotation Can be accessed from the Ami. GO entry page (front page)
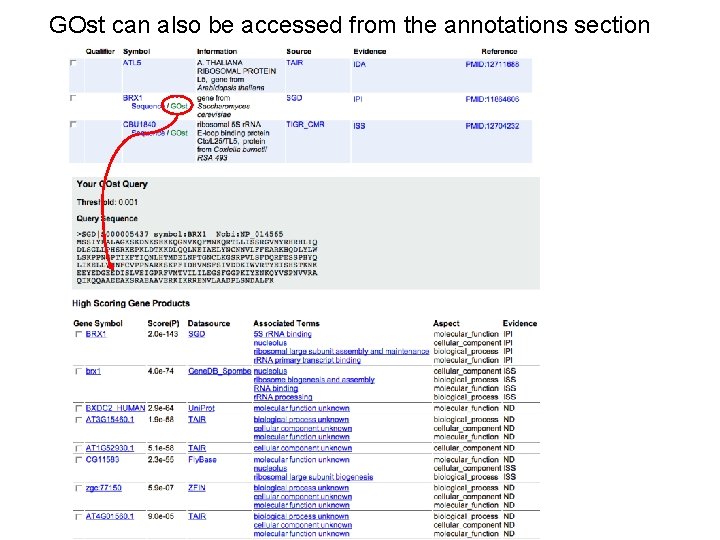
GOst can also be accessed from the annotations section

We welcome your Input • You can make your suggestions on the Source. Forge site • URL: https: //sourceforge. net/projects/geneontology/ • gohelp@geneontology. org
- Slides: 14