Radiation Targets 1 DNA Chromosome and Chromatid Damage












































- Slides: 71

Radiation Targets 1: DNA, Chromosome and Chromatid Damage and Repair Bill Mc. Bride Dept. Radiation Oncology David Geffen School Medicine UCLA, Los Angeles, Ca. wmcbride@mednet. ucla. edu WMc. B 2008 www. radbiol. ucla. edu

Objectives: • Know the limitations of different assays for different types of DNA strand breaks • Know the different types of DNA and chromosome radiation damage • Understand that multiple DNA repair mechanisms exist and why • Be able to discuss repair of base, single strand double strand DNA breaks • Know the molecules involved in homologous recombination and non-homologous end joining and how these initiate DNA damage response pathways • Understand how DNA repair activates the DNA damage response pathway • Recognize the role of DNA repair mutations in carcinogenesis WMc. B 2008 www. radbiol. ucla. edu

• DNA repair enzymes continuously monitor chromosomes to correct damaged nucleotides – Endogenous mutagens - ROS from cellular respiration, hydrolysis, metabolites that act as alkylating agents – Exogenous mutagens - U. V. , cigarette smoke, dietary factors • Apurinic/Apyrimidinic sites are the most common form of naturally occurring DNA damage – 10 -20, 000 apurinic, 500 apyrimidinic, and 170 8 oxyguanines sites produced per day per cell under physiologic conditions • The number of DSB/cell/day in vivo are not well known but 510% of dividing mammalian cells in culture have at least 1 chromosome break or chromatid gap • Each time a cell divides it forms 10 DSBs, and 50, 000 SS! WMc. B 2008 www. radbiol. ucla. edu

• Failure to repair leads to block in DNA replication, permanent cell cycle arrest, senescence, or death • These are the barriers that prevent development of genomic instability and carcinogenesis WMc. B 2008 www. radbiol. ucla. edu

Relevant Properties of DNA when Measuring Damage • A very long double-stranded helix with basestacking • Complementary strands are hybridized to each other via H-bonding and unwind under alkaline conditions • Negatively-charged at physiological p. H • Yield of radiation-induced damage is affected by macromolecular organization of DNA From: Watson et al. “Mol. Biol. of the Cell” www. radbiol. ucla. edu WMc. B 2008

Assays Popular and classic DNA damage assays • Neutral and alkaline elution through filters or separation on sucrose gradients - a classic assay for DSB and SSB+DSB, respectively • Comet assay - sensitive assay for SSB that can be used for single cells; less sensitive (10 Gy) for DSBs • H 2 AX focus formation at DNA DSB - sensitive, currently favored DSB assay Research DNA damage assays • DNA unwinding assay - a research assay • Pulsed field gel electrophoresis - research assay that needs a high radiation dose • Quantification of damaged bases - a very specific assay, mainly for oxidative stress • PCR-based assays - new range of research assays that still require validation Chromosome/chromatid assays • Micronucleus formation - classic assay especially in occupational exposure • Chromosome/ chromatid aberration - classic exposure dosimetric assay – Conventional staining – Banding – FISH www. radbiol. ucla. edu WMc. B 2008

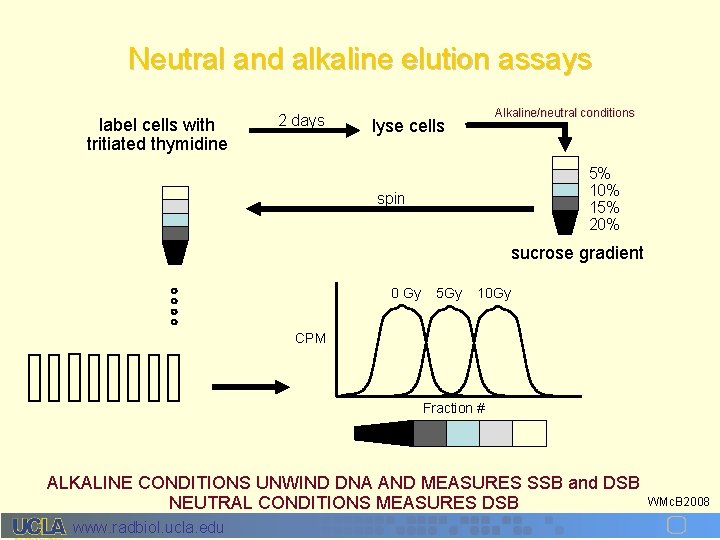
Neutral and alkaline elution assays label cells with tritiated thymidine 2 days Alkaline/neutral conditions lyse cells 5% 10% 15% 20% spin sucrose gradient 0 Gy 5 Gy 10 Gy CPM Fraction # ALKALINE CONDITIONS UNWIND DNA AND MEASURES SSB and DSB NEUTRAL CONDITIONS MEASURES DSB www. radbiol. ucla. edu WMc. B 2008

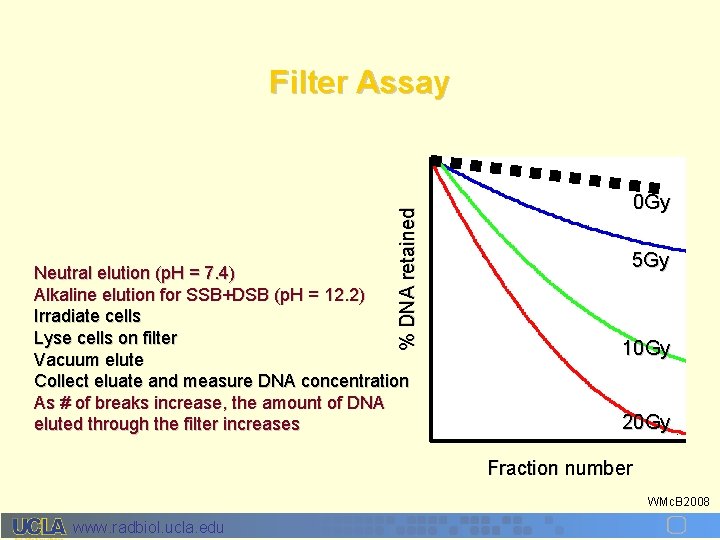
% DNA retained Filter Assay Neutral elution (p. H = 7. 4) Alkaline elution for SSB+DSB (p. H = 12. 2) Irradiate cells Lyse cells on filter Vacuum elute Collect eluate and measure DNA concentration As # of breaks increase, the amount of DNA eluted through the filter increases 0 Gy 5 Gy 10 Gy 20 Gy Fraction number WMc. B 2008 www. radbiol. ucla. edu

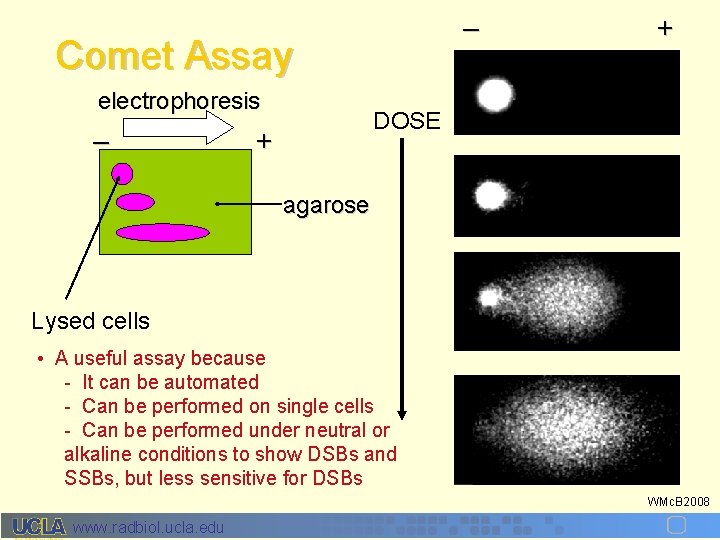
– Comet Assay electrophoresis – + DOSE + agarose Lysed cells • A useful assay because - It can be automated - Can be performed on single cells - Can be performed under neutral or alkaline conditions to show DSBs and SSBs, but less sensitive for DSBs WMc. B 2008 www. radbiol. ucla. edu

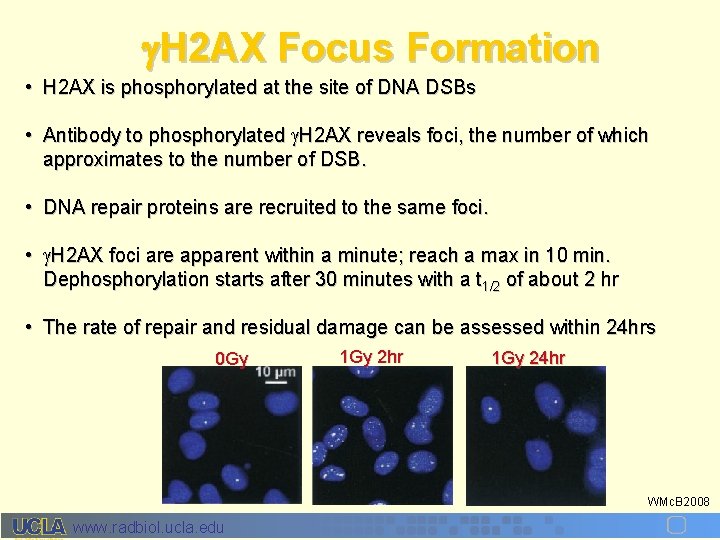
H 2 AX Focus Formation • H 2 AX is phosphorylated at the site of DNA DSBs • Antibody to phosphorylated H 2 AX reveals foci, the number of which approximates to the number of DSB. • DNA repair proteins are recruited to the same foci. • H 2 AX foci are apparent within a minute; reach a max in 10 min. Dephosphorylation starts after 30 minutes with a t 1/2 of about 2 hr • The rate of repair and residual damage can be assessed within 24 hrs 0 Gy 1 Gy 2 hr 1 Gy 24 hr WMc. B 2008 www. radbiol. ucla. edu

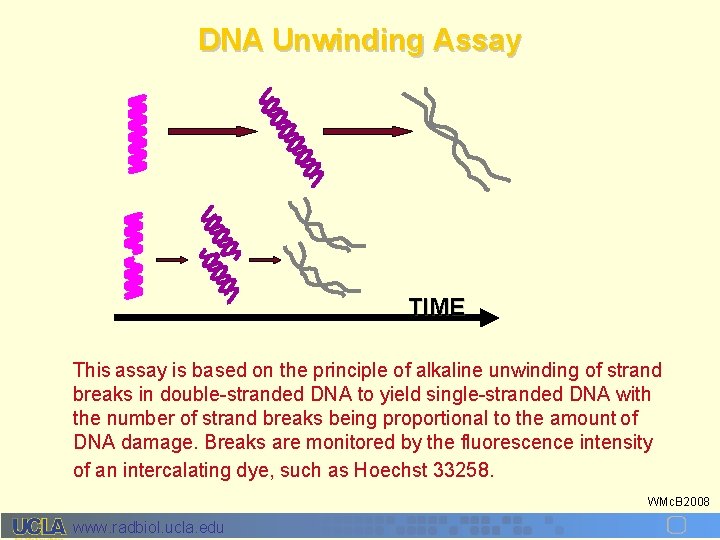
DNA Unwinding Assay TIME This assay is based on the principle of alkaline unwinding of strand breaks in double-stranded DNA to yield single-stranded DNA with the number of strand breaks being proportional to the amount of DNA damage. Breaks are monitored by the fluorescence intensity of an intercalating dye, such as Hoechst 33258. WMc. B 2008 www. radbiol. ucla. edu

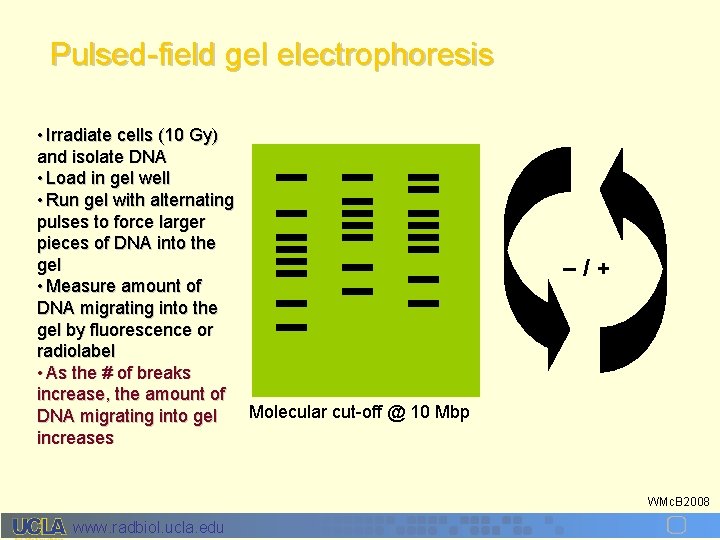
Pulsed-field gel electrophoresis • Irradiate cells (10 Gy) and isolate DNA • Load in gel well • Run gel with alternating pulses to force larger pieces of DNA into the gel • Measure amount of DNA migrating into the gel by fluorescence or radiolabel • As the # of breaks increase, the amount of Molecular cut-off @ 10 Mbp DNA migrating into gel increases –/+ WMc. B 2008 www. radbiol. ucla. edu

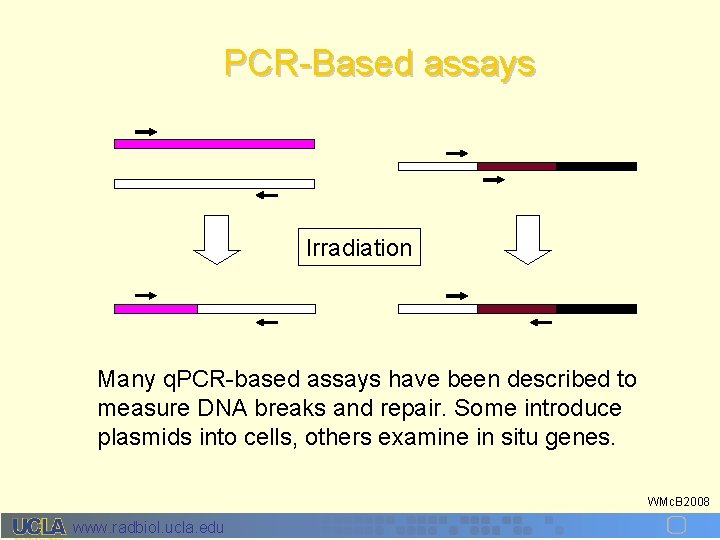
PCR-Based assays Irradiation Many q. PCR-based assays have been described to measure DNA breaks and repair. Some introduce plasmids into cells, others examine in situ genes. WMc. B 2008 www. radbiol. ucla. edu

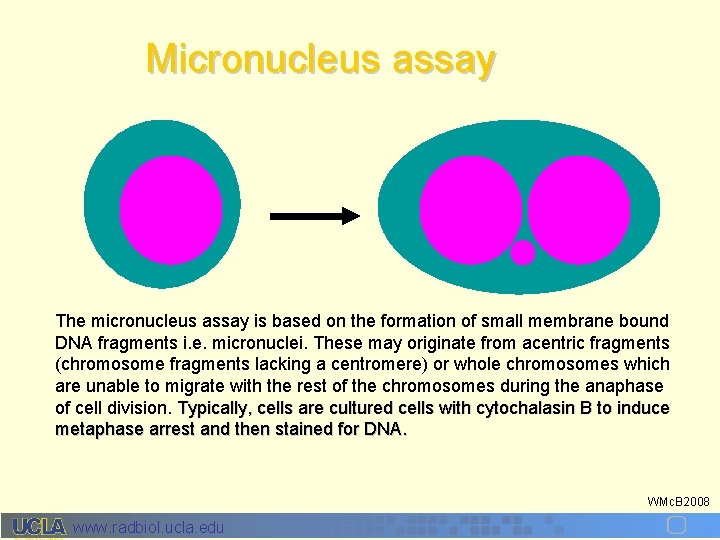
Micronucleus assay The micronucleus assay is based on the formation of small membrane bound DNA fragments i. e. micronuclei. These may originate from acentric fragments (chromosome fragments lacking a centromere) or whole chromosomes which are unable to migrate with the rest of the chromosomes during the anaphase of cell division. Typically, cells are cultured cells with cytochalasin B to induce metaphase arrest and then stained for DNA. WMc. B 2008 www. radbiol. ucla. edu

Chromosome/Chromatid Aberrations Cytogenetic damage is normally assessed at first metaphase after irradiation. The type of cytogenetic damage depends upon where in the cell cycle the cell is when it is irradiated Mitosis Chromosome aberrations • G 1 irradiation • Both sister chromatids involved Chromatid aberrations • S or G 2 irradiation • Usually only 1 chromatid involved There are 2 basic types of lesion • Deletion-type • Exchange-type G 2 G 1 S phase WMc. B 2008 www. radbiol. ucla. edu

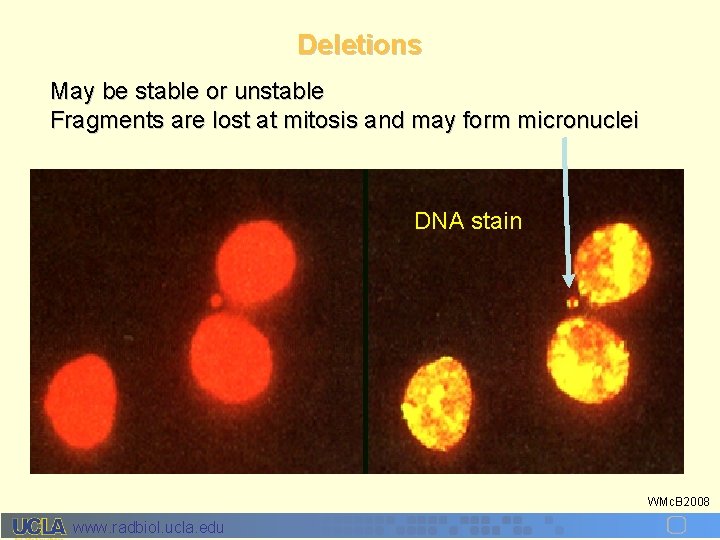
Deletions May be stable or unstable Fragments are lost at mitosis and may form micronuclei DNA stain WMc. B 2008 www. radbiol. ucla. edu

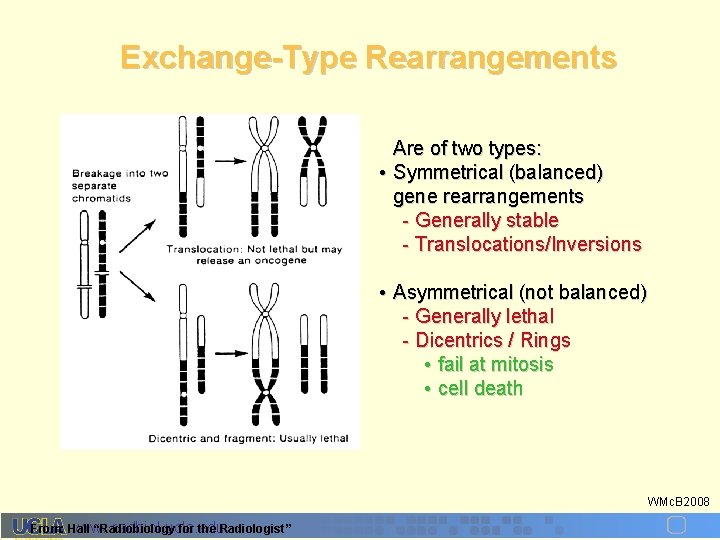
Exchange-Type Rearrangements Are of two types: • Symmetrical (balanced) gene rearrangements - Generally stable - Translocations/Inversions • Asymmetrical (not balanced) - Generally lethal - Dicentrics / Rings • fail at mitosis • cell death WMc. B 2008 www. r adbiol. ucla. edu From: Hall “Radiobiology for the Radiologist ”

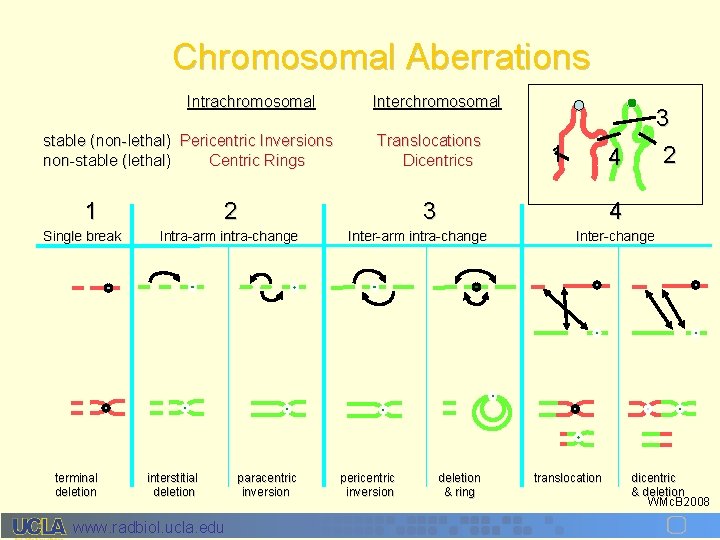
Chromosomal Aberrations Intrachromosomal stable (non-lethal) Pericentric Inversions non-stable (lethal) Centric Rings 1 Single break terminal deletion Interchromosomal Translocations Dicentrics 2 Intra-arm intra-change interstitial deletion www. radbiol. ucla. edu paracentric inversion 1 4 3 4 Inter-arm intra-change pericentric inversion 3 2 deletion & ring Inter-change translocation dicentric & deletion WMc. B 2008

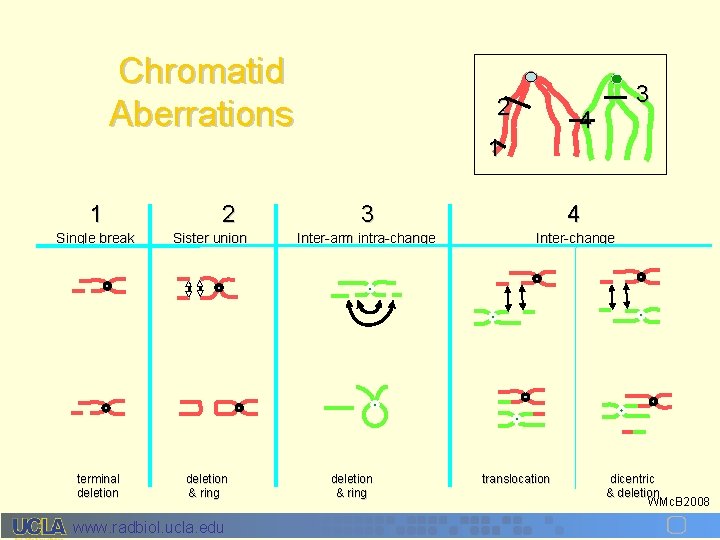
Chromatid Aberrations 1 2 Single break Sister union terminal deletion & ring www. radbiol. ucla. edu 2 3 4 1 3 4 Inter-arm intra-change Inter-change deletion & ring translocation dicentric & deletion WMc. B 2008

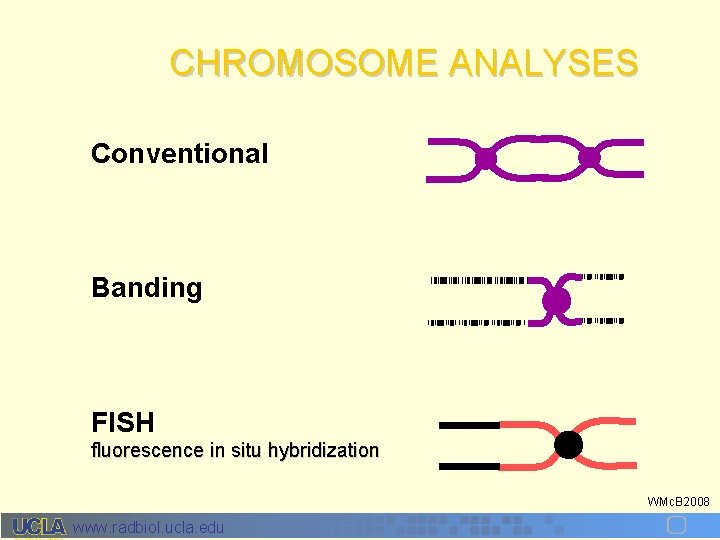
CHROMOSOME ANALYSES Conventional Banding FISH fluorescence in situ hybridization WMc. B 2008 www. radbiol. ucla. edu

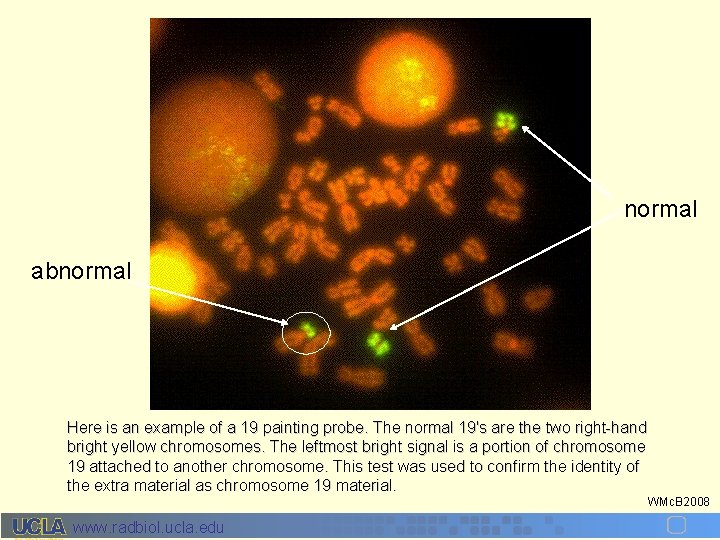
normal abnormal Here is an example of a 19 painting probe. The normal 19's are the two right-hand bright yellow chromosomes. The leftmost bright signal is a portion of chromosome 19 attached to another chromosome. This test was used to confirm the identity of the extra material as chromosome 19 material. www. radbiol. ucla. edu WMc. B 2008

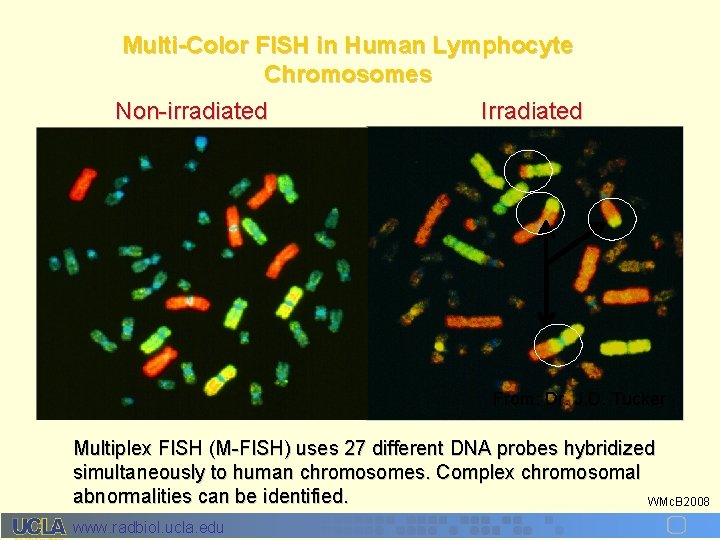
Multi-Color FISH in Human Lymphocyte Chromosomes Non-irradiated Irradiated From: Dr. J. D. Tucker Multiplex FISH (M-FISH) uses 27 different DNA probes hybridized simultaneously to human chromosomes. Complex chromosomal abnormalities can be identified. WMc. B 2008 www. radbiol. ucla. edu

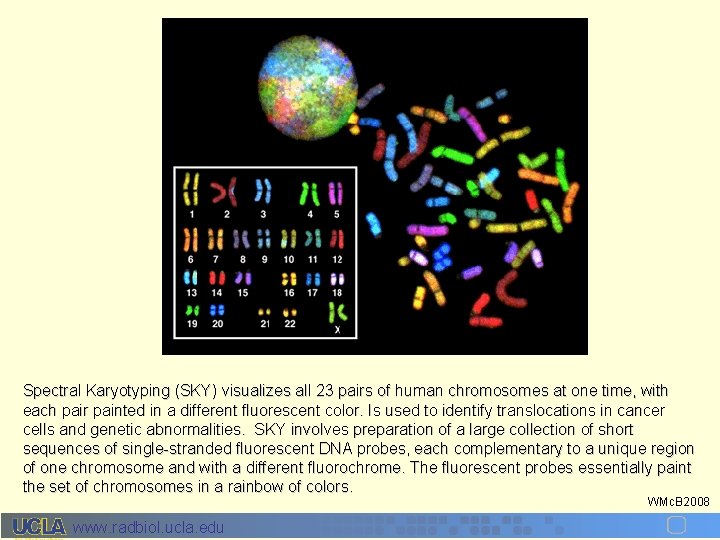
Spectral Karyotyping (SKY) visualizes all 23 pairs of human chromosomes at one time, with each pair painted in a different fluorescent color. Is used to identify translocations in cancer cells and genetic abnormalities. SKY involves preparation of a large collection of short sequences of single-stranded fluorescent DNA probes, each complementary to a unique region of one chromosome and with a different fluorochrome. The fluorescent probes essentially paint the set of chromosomes in a rainbow of colors. WMc. B 2008 www. radbiol. ucla. edu

Yield of radiation-induced chromosome damage Deletions Terminal deletion = 1 hit Chromatid deletion = 1 hit Interstitial deletion = 2 hits Yield (Y) ~ linear Y = k + D k = background = proportionality DOSE (Gy) Cornford and Bedford Rad Res 111: 385, 1987 Fate: Deletions lost at mitosis WMc. B 2008 www. radbiol. ucla. edu

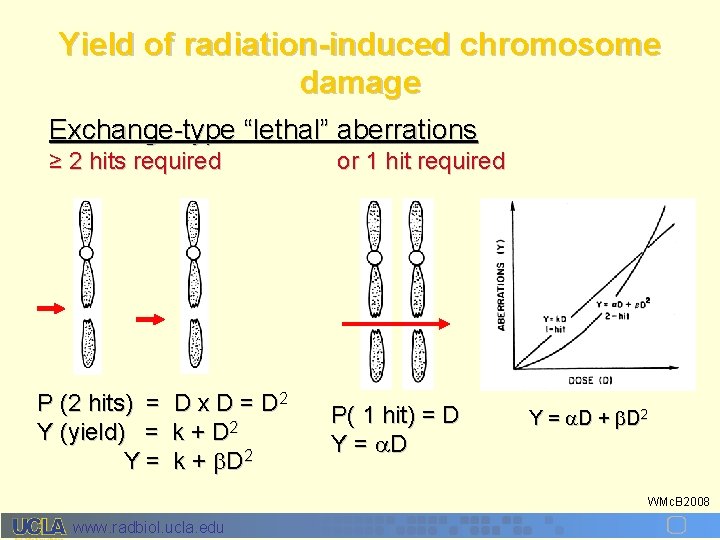
Yield of radiation-induced chromosome damage Exchange-type “lethal” aberrations ≥ 2 hits required P (2 hits) = Y (yield) = Y= D x D = D 2 k + b D 2 or 1 hit required P( 1 hit) = D Y = D + b D 2 WMc. B 2008 www. radbiol. ucla. edu

A plot of # “lethal” aberrations vs natural log S. F. showed that an average of 1 lethal lesion decreased survival by e. In other words, S. F. = e –( D + b. D 2) WMc. B 2008 www. radbiol. ucla. edu

DNA Repair • • • Classically, there are 2 types Sub-Lethal and Potentially Lethal Damage These are operationally-defined terms that differ in the experimental set up in which they are demonstrated – PLDR - single dose – SLDR - split (fractionated) doses • The molecular mechanisms may be similar, but this is not clear WMc. B 2008 www. radbiol. ucla. edu

Potentially Lethal Damage • Potentially lethal damage is defined as damage that could cause death, but is modified by post-irradiation conditions WMc. B 2008 www. radbiol. ucla. edu

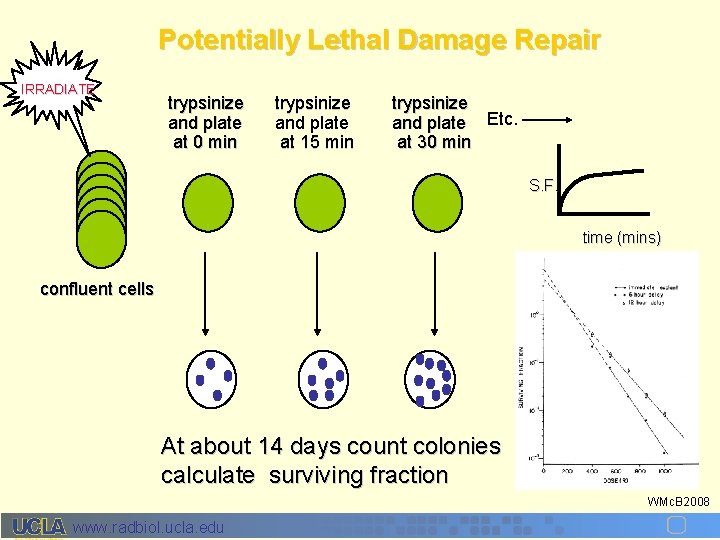
Potentially Lethal Damage Repair IRRADIATE trypsinize and plate at 0 min trypsinize and plate at 15 min trypsinize and plate Etc. at 30 min S. F. time (mins) confluent cells At about 14 days count colonies calculate surviving fraction WMc. B 2008 www. radbiol. ucla. edu

Sub-lethal (or accumulated) damage results from accumulation of events that individually are incapable of killing a cell but that together can be lethal 4 nm Repairable Sublethal Damage 2 nm Unrepairable Multiply Damaged Site WMc. B 2008 www. radbiol. ucla. edu

• To account for the time gap between the production of 2 sublethal lesions (dose rate), Lea and Catcheside (J Genetics 44: 216, 1942) introduced the factor q • S. F. = e –( D + qb. D 2) WMc. B 2008 www. radbiol. ucla. edu

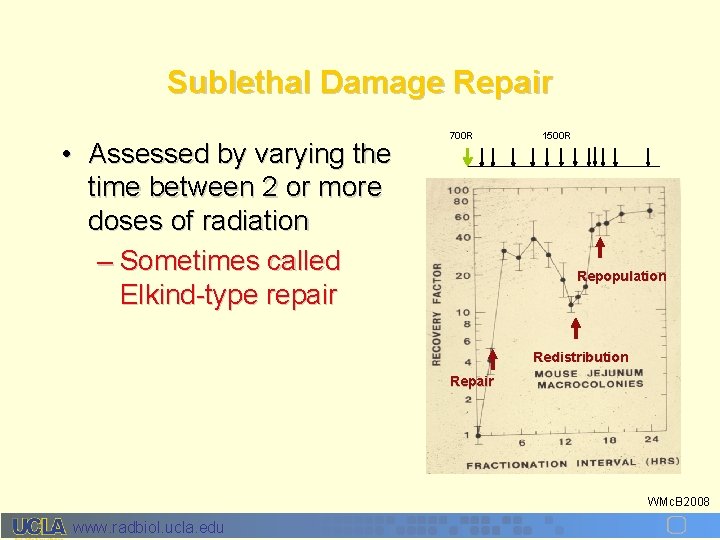
Sublethal Damage Repair • Assessed by varying the time between 2 or more doses of radiation – Sometimes called Elkind-type repair 700 R 1500 R Repopulation Redistribution Repair WMc. B 2008 www. radbiol. ucla. edu

Some Molecular Forms of DNA Repair • Base Excision Repair • • • – Repairs most of the 10 -20, 000 apurinic and 500 apyrimidinic sites/cell/day that form spontaneously – Important for repair of most SSB and base damage after IR. – Persistence leads to a block in DNA replication, cytotoxic mutations, genetic instability. – apurinic/apyrimidinic (AP) endonuclease removes 1 -3 nucleotides – T 1/2 <5 mins. Active genes repaired faster than inactive Nucleotide Excision Repair – Repairs U. V. photodamage, chemical adducts, crosslinks by removing pyrimidine dimers and other helix distorting lesions. Of minor importance for IR. – Involved in Global Genome repair and Transcription-Coupled repair – About 30 nucleotides are excised DNA Mismatch Repair – Corrects base-base mismatches and small loops – Important in removing replication errors. Of minor importance for IR. – Important in connection with hereditary colorectal cancer (h. MSH 2, h. MLH 1, h. PMS 2) and microsatellite instability Double Strand Break Repair WMc. B 2008 www. radbiol. ucla. edu

• Enzymes exist that reverse rather than excise DNA damage exist – • The use of repair molecules and processes depends on a lot of factors – • eg. MGMT (O 6 -methylguanine DNA methyltransferase) removes methyl and other alkyl groups • “Patients with glioblastoma containing a methylated MGMT promoter benefited from temozolomide, whereas those who did not have a methylated MGMT promoter did not have such a benefit. ” Hegi et al NEJM 352: 997 -1003, 2005 eg. Repair of DNA-DNA cross-links after XRT uses NER There about 130 DNA repair genes. Luckily, there are 3 major molecular processes in common – – – Nucleases remove damaged DNA Polymerases lay down the new structures Ligases restore the phosphodiester backbone WMc. B 2008 www. radbiol. ucla. edu

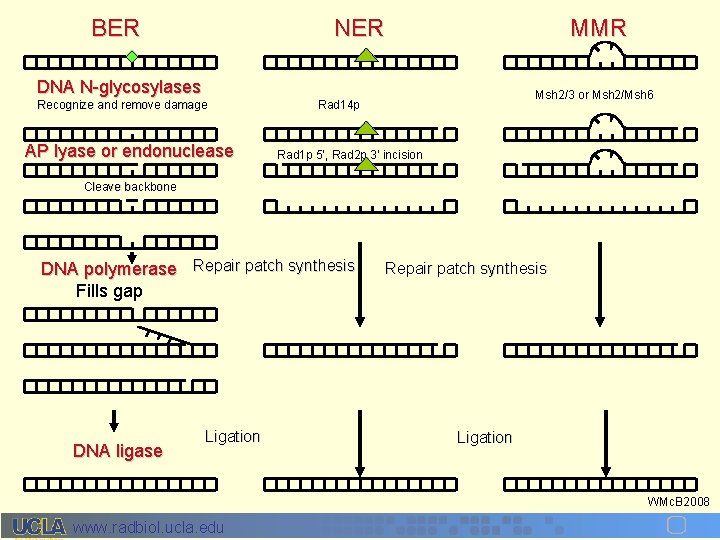
BER NER MMR DNA N-glycosylases Recognize and remove damage AP lyase or endonuclease Msh 2/3 or Msh 2/Msh 6 Rad 14 p Rad 1 p 5’, Rad 2 p 3’ incision Cleave backbone DNA polymerase Repair patch synthesis Fills gap DNA ligase Ligation Repair patch synthesis Ligation WMc. B 2008 www. radbiol. ucla. edu

DSBs • DSBs can be formed physiologically or pathologically • Physiological – During VDJ recombination to form Ab or T cell receptors – Class switch breaks to make different Ab isotypes – Mutations to increase Ab affinity – During meiosis • Pathological – Ionizing radiation – ROS during cellular respiration – DNA replication across a nick – Enzymic action especially at fragile DNA sites – Topoisomerase failures WMc. B 2008 www. radbiol. ucla. edu

DSB Repair Recombination Models of DSB Repair #1 • Homologous Recombination – Uses a sister chromatid (in S and G 2) or a second chromosome (in M) as a template – Does not occur in G 1 – Is relatively error free – Mutants defective in HR have increased chromosomal aberrations but can generally repair DSBs (inefficiently) – The major molecular players are: • MRN complex, Rad 51/Rad 52/XRCC 2/ Rad 54/BRCA 1/2 WMc. B 2008 www. radbiol. ucla. edu

DSB Repair • Models of DSB Repair #2 • Non Homologous End Joining uses a non-homologous template with little or no microhomology – Imprecise, makes mistakes (an advantage in the immune system) – Active at any time in cell cycle – Efficient at restoring chromosomal integrity – The major mechanism of DSB repair – Used physiologically in VDJ rejoining of T cell and Ig receptors – Mammalian mutants deficient in NHEJ are deficient in DNA repair and immunity (severe combined immune deficiency scid - in mice and humans) – The major molecular players are: • Ku 70/Ku 80 - Artemis/DNA-PKcs - Cerrunos/XRCC 4/ligase. IV • Microhomology-mediated end joining is an inefficient alternative that is Ku/ligase. IV independent WMc. B 2008 www. radbiol. ucla. edu

Non Homologous End Joining • • • Ku 70/80 (or 86) heterocomplex tethers DNA and recruits DNA-PKcs that promotes binding of various proteins: Nucleases that remove damaged DNA – Artemis/DNA-PKcs bind to form a 5’ to 3’ endonuclease that makes blunt ends – DNA-PK is activated on binding DNA – Autophosphorylation aids binding of other repair proteins Polymerases that lay down the nucleotide structure – Pol X family members and Td. T that have varying degrees of template dependency. pol can add nucleotides randomly to generate microhomology that assists repair Ligases restore the phosphodiester backbone – Cernunnos (XLF)/XRCC 4/DNA ligase IV complex – XRCC 4/DNA ligase IV are flexible being able to ligate just one strand or across gaps Each enzyme has a range of flexibilities, allowing many outcomes WMc. B 2008 www. radbiol. ucla. edu

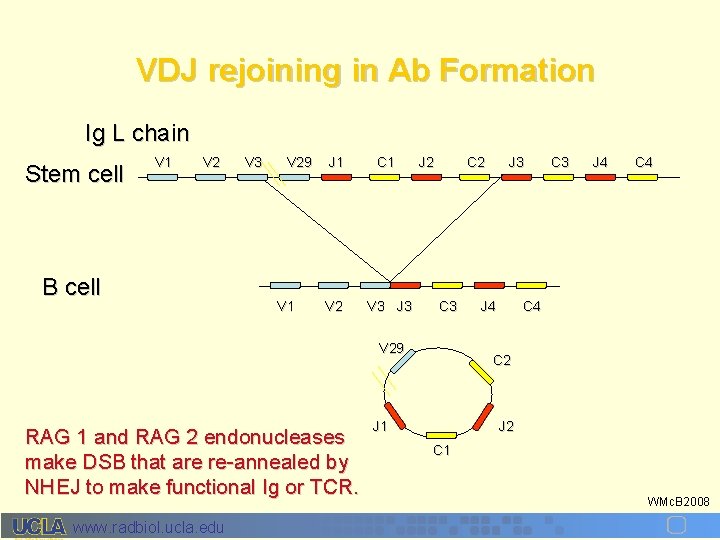
VDJ rejoining in Ab Formation Ig L chain Stem cell V 1 V 2 B cell V 3 V 29 V 1 J 1 C 1 V 2 V 3 J 2 C 3 V 29 RAG 1 and RAG 2 endonucleases make DSB that are re-annealed by NHEJ to make functional Ig or TCR. www. radbiol. ucla. edu J 3 J 4 C 4 C 2 J 1 J 2 C 1 WMc. B 2008

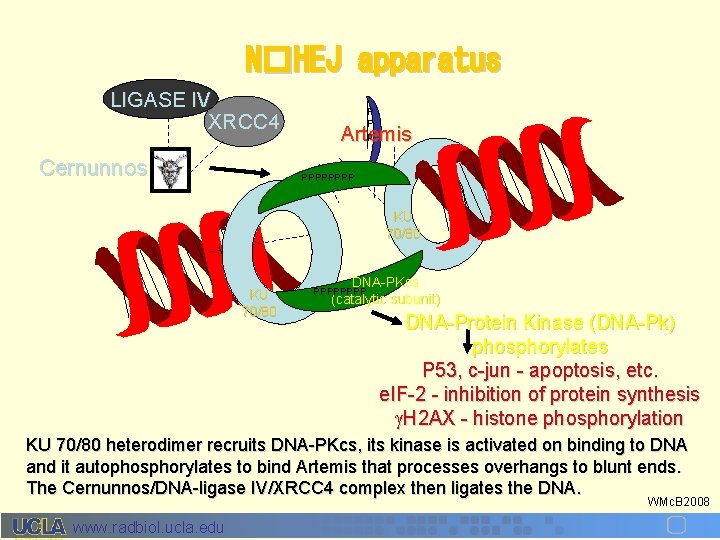
N�HEJ apparatus LIGASE IV XRCC 4 Cernunnos P P p Artemis PPPP KU 70/80 DNA-PKcs (catalytic subunit) PPPP DNA-Protein Kinase (DNA-Pk) phosphorylates P 53, c-jun - apoptosis, etc. e. IF-2 - inhibition of protein synthesis H 2 AX - histone phosphorylation KU 70/80 heterodimer recruits DNA-PKcs, its kinase is activated on binding to DNA and it autophosphorylates to bind Artemis that processes overhangs to blunt ends. The Cernunnos/DNA-ligase IV/XRCC 4 complex then ligates the DNA. WMc. B 2008 www. radbiol. ucla. edu

DNA-PK • Only protein known to be activated by binding DSB • Required for DNA DSB repair and V(D)J rejoining by NHEJ • Composed of DNA-PKcs (p 450), KU 70, Ku 80 • A large molecule - 4127 aa, 470 k. Da, 180 kbp • Is a ser/thr kinase with homology to PI-3 kinase, but has no lipid activity. • Scid mice defective in DNA-PKcs • Most Scid humans are defective in Artemis, which is phosphorylated by DNA-PK and binds to DNA DSB to form an endonuclease Foci are formed that act as an amplification platform WMc. B 2008 www. radbiol. ucla. edu

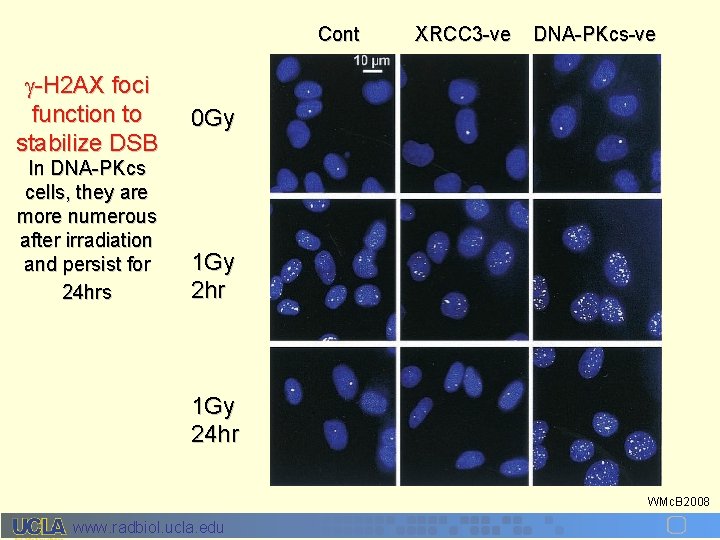
Cont -H 2 AX foci function to stabilize DSB In DNA-PKcs cells, they are more numerous after irradiation and persist for 24 hrs XRCC 3 -ve DNA-PKcs-ve 0 Gy 1 Gy 2 hr 1 Gy 24 hr WMc. B 2008 www. radbiol. ucla. edu

DSB Repair The Mre 11/ Rad 50/ NBS 1 (MRN) Complex is involved as a tether for DSB for HR Mre 11 has nuclease activity - NBS = Nijmegen Breakage Syndrome protein (nibrin) binds ATM - Nijmegen Breakage Syndrome patients are - Radiation sensitive Have microcephaly Immune deficiencies Predisposition to lymphoid malignancies Cells show - defect in DSB repair - cell cycle arrest abnormalities - Including radio-resistant DNA synthesis WMc. B 2008 www. radbiol. ucla. edu

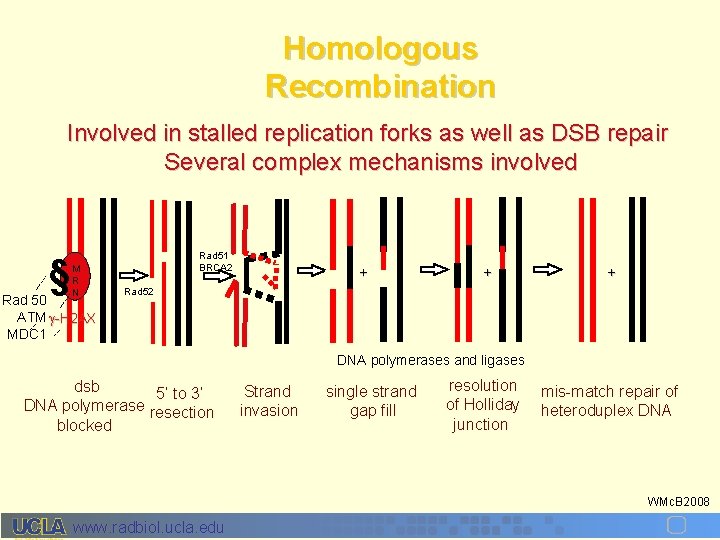
Homologous Recombination Involved in stalled replication forks as well as DSB repair Several complex mechanisms involved § M R N Rad 50 ATM -H 2 AX MDC 1 Rad 51 BRCA 2 + + + Rad 52 DNA polymerases and ligases dsb 5’ to 3’ DNA polymerase resection blocked Strand invasion single strand gap fill resolution of Holliday junction mis-match repair of heteroduplex DNA WMc. B 2008 www. radbiol. ucla. edu

• Chromatin structure decondenses at site of DSB • Histone acetylation and ubiquitination involved in DNA repair WMc. B 2008 www. radbiol. ucla. edu

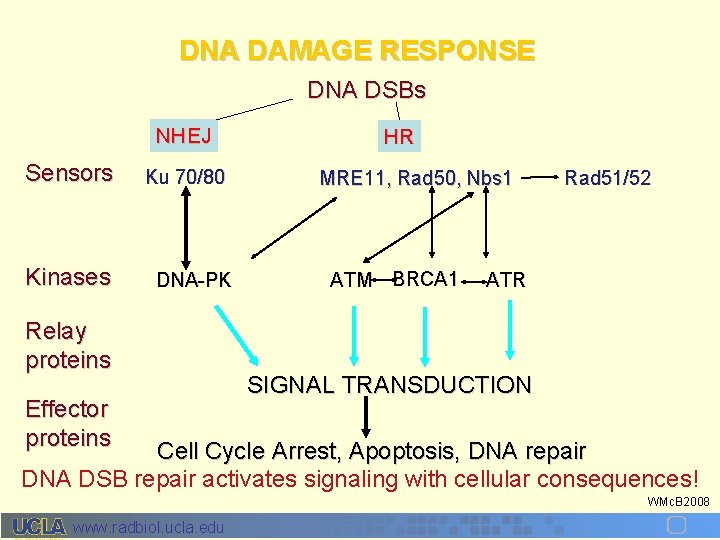
DNA DAMAGE RESPONSE DNA DSBs NHEJ Sensors Kinases Ku 70/80 DNA-PK Relay proteins Effector proteins HR MRE 11, Rad 50, Nbs 1 ATM BRCA 1 Rad 51/52 ATR SIGNAL TRANSDUCTION Cell Cycle Arrest, Apoptosis, DNA repair DNA DSB repair activates signaling with cellular consequences! WMc. B 2008 www. radbiol. ucla. edu

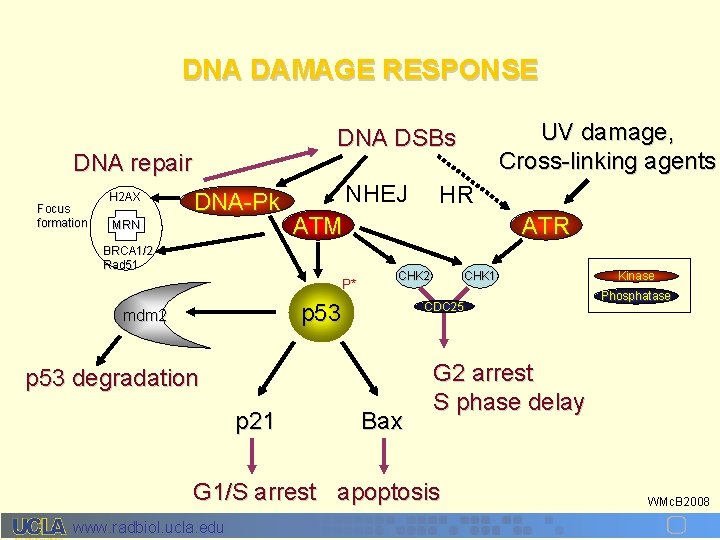
DNA DAMAGE RESPONSE DNA repair Focus formation H 2 AX UV damage, Cross-linking agents DNA DSBs DNA-Pk MRN NHEJ ATM BRCA 1/2 Rad 51 P* ATR CHK 2 p 53 mdm 2 CHK 1 CDC 25 p 53 degradation p 21 HR Bax Phosphatase G 2 arrest S phase delay G 1/S arrest apoptosis www. radbiol. ucla. edu Kinase WMc. B 2008

• DNA repair genes are genomic “caretaker” genes preventing cancer by removing DNA mutations • Defects in DNA repair genes are very common in cancers • Loss of many DNA repair genes is embryonic lethal or results in genomic instability • Individuals who are ‘carriers’ of defective DNA repair genes may be especially sensitive to irradiation and radiation-induced cancers and may be 5 -10% of the population – Epidemiological studies have shown that AT heterozygotes have a predisposition for cancer, especially for breast cancer in women. WMc. B 2008 www. radbiol. ucla. edu

Autosomal Recessive Disorders with Repair Defects • Xeroderma pigmentosum (XP) and related Cockayne’s syndrome –U. V. sensitivity –At least 7 genes (ERCC 1 -6; excision repair cross complementing) –DNA binding and damage recognition, helicase, endonucleases, transcription factors, inability to excise dimers • Fanconi’s anemia –Mutated in 90% aplastic anemias, commonly in leukemias, 20% solid tumors –Sensitivity to X-linking agents (e. g. mitomycin C) - genomic instability – 7 genes cloned (A, C, D 1, D 2, E, F, G); D 1 is BRCA 2 • Bloom’s and Werner’s syndromes –Helicases mutated –Defective recombination and replication –Cancer predisposition • Li Fraumeni syndrome –Rare autosomal dominant –Breast, soft tissue, bone sarcomas with multiple primaries in childhood – 70% have p 53 mutations, others have CHK 2 mutations • Ataxia telangiectasia • Nijmegen-breakage syndrome www. radbiol. ucla. edu WMc. B 2008

Ataxia Telengiectasia • • • Rare autosomal recessive - 1: 20, 000 -1: 1000, described in 1920 s 70 -250 fold excess of leukemia/lymphoma and carcinomas (1960 s) Sensitive to ionizing radiation (1974) Cerebellar degeneration, progressive ataxia, telangiectasia, immune deficiency (T and B) Chromosomal instability, DSB repair defect, initial damage unaltered Signal transduction defect - low, late p 53/ GADD 45/ c-jun induction No G 1 arrest, no S phase delay (RDS), G 2 arrest altered AT gene (1995) homology to phosphoinositol-3 kinase superfamily ATM truncations and missense might give different outcomes Missense found in 8% of breast cancer patients, 20%CLL. No increase in truncations. AT heterozygotes have 1. 3 -2. 9 fold increase in breast cancer risk. No obvious increase in cytotoxic radiosensitivity. WMc. B 2008 www. radbiol. ucla. edu

Nijmegen Breakage Syndrome • • • Nibrin (Nbs) gene on chromosome 8 q 21 Microcephaly, growth and mental retardation High leukemia risk Radiation sensitivity Late, low p 53, lack G 1/S arrest Nibrin binds in MRE 11, RAD 50 (MRN) complex • AT-Like Disorder (ATLD) is an Mre 11 defect WMc. B 2008 www. radbiol. ucla. edu

BRCA 1 and BRCA 2 Tumor Suppressor Genes • • • BRCA 1: – Average 65 % lifetime risk for breast cancer – 40 percent to 60 percent lifetime risk for second breast cancer (not reappearance of first tumor) – Average 39 percent lifetime risk for ovarian cancer – Increased risk for other cancer types, such as prostate cancer – BRCA 1 cancers tend to be “basal-like”, ER-ve – Expressed in proliferating cells at G 1/S – Associate with rad 51 which is involved in DSB repair in HR BRCA 2 is FANC-D 1 – Average 45 % lifetime risk for breast cancer in females, 6% in males – Average 11 percent lifetime risk for ovarian cancer – Increased risk for other cancer types, such as pancreatic, prostate, laryngeal, stomach cancer, and melanoma Cells with mutated BRCA 1/2 are slightly more sensitive to radiation, cisplatin and MMC because of their role in homologous recombination WMc. B 2008 www. radbiol. ucla. edu

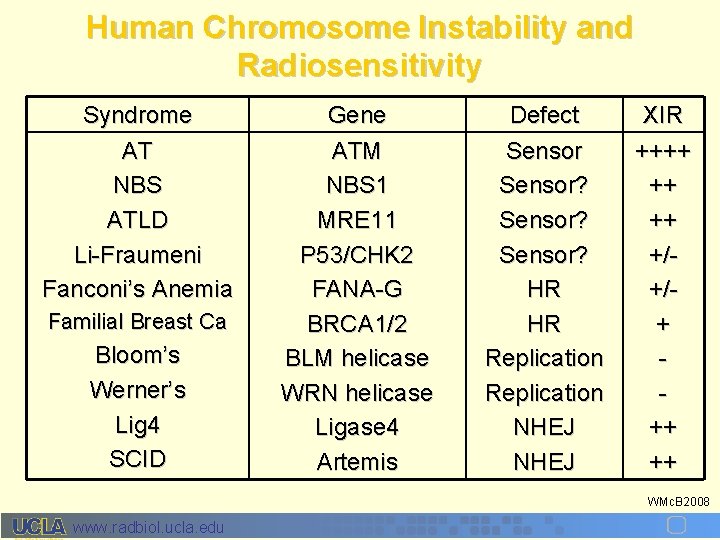
Human Chromosome Instability and Radiosensitivity Syndrome AT NBS ATLD Li-Fraumeni Fanconi’s Anemia Familial Breast Ca Bloom’s Werner’s Lig 4 SCID Gene ATM NBS 1 MRE 11 P 53/CHK 2 FANA-G BRCA 1/2 BLM helicase WRN helicase Ligase 4 Artemis Defect Sensor? HR HR Replication NHEJ XIR ++++ ++ ++ +/+/+ ++ ++ WMc. B 2008 www. radbiol. ucla. edu

Questions on Radiation Targets 1: DNA, Chromosome and Chromatid Damage and Repair WMc. B 2008 www. radbiol. ucla. edu

What is the most common form of DNA damage existing in cells under normal conditions? 1. Double strand breaks 2. Apurinic/apyrimidinic sites 3. Interstrand crosslinks 4. Thymidine dimer formation WMc. B 2008 www. radbiol. ucla. edu

What assay would be the most sensitive to measure radiation-induced DNA double strand breaks? 1. Neutral elution of DNA 2. H 2 AX focus formation 3. Comet assay under neutral conditions 4. Pulsed field electrophoresis of DNA WMc. B 2008 www. radbiol. ucla. edu

What radiation damage is measured by the alkaline elution of DNA technique 1. Single strand breaks 2. Base damage 3. Double strand breaks 4. Single and double strand breaks 5. DNA-protein crosslinks WMc. B 2008 www. radbiol. ucla. edu

Which of the following is true for the H 2 AX focus formation assay? 1. It measures the ability of radiation to transform normal cells towards cancer 2. Most foci are seen at 24 hours 3. The foci that form after about 10 mins approximate to the number of radiation-induced DNA DSBs 4. The foci are dependent of activation of ATM kinase WMc. B 2008 www. radbiol. ucla. edu

The micronucleus assay 1. Is a measure of DNA damage 2. Measures fragments of nuclei that are lost at mitosis 3. Is a measure of histone damage 4. Measure micronuclei formed by chromosome translocations 5. Uses micro. RNA techniques to measure DNA damage WMc. B 2008 www. radbiol. ucla. edu

Radiation-induced chromosome damage that is usually lethal is most likely due to 1. Deletions 2. Translocation 3. Exchange-type aberrations 4. Gene loss 5. Dicentics or rings WMc. B 2008 www. radbiol. ucla. edu

Which of the following is correct about sublethal damage repair. It occurs 1. When cells are held in a non-proliferating state 2. Between fractions of radiation 3. At the G 1/s checkpoint 4. Only at low radiation doses WMc. B 2008 www. radbiol. ucla. edu

Which of the following repair mechanisms is most important after X-ray exposure of cells 1. Mismatch repair 2. Nucleotide excision repair 3. Double strand break repair 4. Base excision repair WMc. B 2008 www. radbiol. ucla. edu

Double strand breaks are least likely to contribute to DNA lesions in which of the following situations 1. Cellular respiration 2. VDJ rejoining to make antibodies 3. Meiosis 4. Antibody class switching WMc. B 2008 www. radbiol. ucla. edu

What sets DNA repair of double strand breaks by homologous recombination apart from non homologous end joining mechanisms? Its involvement in 1. G 1 cell cycle phase only 2. Cell cycle phases other than G 1 3. All cell cycle phases 4. Increasing genomic instability WMc. B 2008 www. radbiol. ucla. edu

What sets DNA repair of double strand breaks by homologous recombination apart from non homologous end joining mechanisms 1. It does not involve the MRN complex 2. It involves ligases 3. It activates the BRCA tumor suppressor protein 4. It is error-prone WMc. B 2008 www. radbiol. ucla. edu

Which of the following is NOT true concerning DNA protein kinase? It is 1. Critical for DNA DSB repair via the nonhomologous end joining pathway 2. Formed from Ku proteins and DNA-PK catalytic subunit 3. Activated on binding DNA DSB 4. Defective in many humans with severe combined immune deficiency (Scid) disease 5. Defective in scid mice WMc. B 2008 www. radbiol. ucla. edu

Which of the following is true about DNA repair genes 1. They are “gatekeeper” genes that directly regulate tumor growth by inhibiting growth or by promoting cell death 2. They are “caretaker” genes that prevent cancercausing mutations 3. There about 20 in human cells 4. Loss of an individual gene is not commonly a problem as the system is highly redundant WMc. B 2008 www. radbiol. ucla. edu

Which of the following in NOT true about Ataxia Telengiectasia lymphoblastoid cells 1. They have a D 0 of about 50 c. Gy 2. They show a higher than normal level of initial DNA double strand breaks after exposure to ionizing irradiation 3. They show no G 1 arrest, but normal S phase arrest after radiation exposure 4. They are slow to elevate p 53 following radiation exposure WMc. B 2008 www. radbiol. ucla. edu

Which of the following sets Nijmegan Breakage Syndrome apart from Ataxia Telengiectasia 1. Cells show less sensitivity to ionizing radiation 2. Patients have no immune deficiency 3. Patients have no neurological problems 4. Cells show normal cell cycle arrest following irradiation 5. It is part of the MRN complex WMc. B 2008 www. radbiol. ucla. edu

Answers 1. 2. 3. 4. 5. 6. 7. 8. 9. 10. 11. 12. 13. 14. 15. 16. NA 2 2 4 3 2 5 2 3 1 2 3 4 3 3 5 WMc. B 2008 www. radbiol. ucla. edu