R for psychologists Professor Thom Baguley Psychology Nottingham

R for psychologists Professor Thom Baguley, Psychology, Nottingham Trent University Thomas. Baguley@ntu. ac. uk

0. Overview 1. What is R? 2. Why use R? 3. R basics 4. R graphics 5. Linear models in R 6. ANOVA and ANCOVA 6. Writing your own R functions 8. Simulation and bootstrapping 2

1. What is R? R is a software environment for statistical analysis “the lingua franca of computational statistics” De Leeuw & Mair (2007) 3

2. Why use R? It is: - free - open source - cross-platform (Mac, Linux, PC) It has: - excellent basic statistics functions - powerful and versatile graphics - hundreds of user-contributed packages - a large community of users 4

3. R basics Some common R objects: - characters (text strings e. g. , 'a' or ’ 1’) - numbers (numbers like 2 or 1 e+3) - vectors (1 D set of numbers or other elements) - data frames (like vectors organized in columns) - matrices (r by c array of numbers) - lists (objects that contain other objects) 5
![Assignment Input: > vector 1 <- 6 > vector 1 Output: [1] 6 6 Assignment Input: > vector 1 <- 6 > vector 1 Output: [1] 6 6](http://slidetodoc.com/presentation_image/19771057d9bfebdb0dc44b36364f9ffb/image-6.jpg)
Assignment Input: > vector 1 <- 6 > vector 1 Output: [1] 6 6
![Arithmetic > 6 + 6 * 2 [1] 18 > vector 1^2 [1] 36 Arithmetic > 6 + 6 * 2 [1] 18 > vector 1^2 [1] 36](http://slidetodoc.com/presentation_image/19771057d9bfebdb0dc44b36364f9ffb/image-7.jpg)
Arithmetic > 6 + 6 * 2 [1] 18 > vector 1^2 [1] 36 7
![Calling functions > c(1, 9, 25) > log(vector 1) [1] 1 9 25 [1] Calling functions > c(1, 9, 25) > log(vector 1) [1] 1 9 25 [1]](http://slidetodoc.com/presentation_image/19771057d9bfebdb0dc44b36364f9ffb/image-8.jpg)
Calling functions > c(1, 9, 25) > log(vector 1) [1] 1 9 25 [1] 1. 791759 > rnorm(1, 100, 15) ? rnorm [1] 123. 5336 help(rnorm) 8

Loading data > dat 1 <- read. csv('pride. csv') > library(foreign) > h 05 <- read. spss('Hayden_2005. sav', to. data. frame = TRUE) 9
![Addressing the contents of an object > vector 1[1] 6 > days <- h Addressing the contents of an object > vector 1[1] 6 > days <- h](http://slidetodoc.com/presentation_image/19771057d9bfebdb0dc44b36364f9ffb/image-10.jpg)
Addressing the contents of an object > vector 1[1] 6 > days <- h 05$days > days[1: 9] [1] 2864 1460 2921 1460 31 10
![Addressing the contents of an object > vector 1[1] 6 > days <- h Addressing the contents of an object > vector 1[1] 6 > days <- h](http://slidetodoc.com/presentation_image/19771057d9bfebdb0dc44b36364f9ffb/image-11.jpg)
Addressing the contents of an object > vector 1[1] 6 > days <- h 05$days > days[1: 9] [1] 2864 1460 2921 1460 31 Or combine the steps with h 05$days[1: 9] 11
![Data frames > is. data. frame(h 05) [1] TRUE > dim(h 05) > names(h Data frames > is. data. frame(h 05) [1] TRUE > dim(h 05) > names(h](http://slidetodoc.com/presentation_image/19771057d9bfebdb0dc44b36364f9ffb/image-12.jpg)
Data frames > is. data. frame(h 05) [1] TRUE > dim(h 05) > names(h 05) [1] 43 [1] "names" "days” 2 - this data frame has 43 rows and 2 named columns - it can be helpful to think of a data frame as a set of named variables (vectors) bound into columns 12

Matrix objects (matrices) > cells <- c(3677, 56, 3924, 31) > cat. names <- list(c("Before", "After"), c("Alive", "Dead")) > checklist <- matrix(cells, 2, 2, byrow=TRUE, dimnames= cat. names) > checklist Alive Dead Before 3677 56 After 3924 31 13

The power of objects > chisq. test(checklist) Pearson's Chi-squared test with Yates' continuity correction data: checklist X-squared = 8. 1786, df = 1, p-value = 0. 004239 > chisq. test(c(2, 18)) Chi-squared test for given probabilities data: c(2, 18) X-squared = 12. 8, df = 1, p-value = 0. 0003466 14

Calling functions: defaults and named arguments > chisq. test(checklist, correct=FALSE) Pearson's Chi-squared test data: checklist X-squared = 8. 8072, df = 1, p-value = 0. 003000 - the default argument (for matrix input is) correct=TRUE - setting correct=FALSE over-rides the default - because unnamed arguments to functions are matched in order, this argument must be named (otherwise naming the arguments is optional) 15

Exercise 1 (R Basics) - entering data - working with objects - simple statistical functions 16
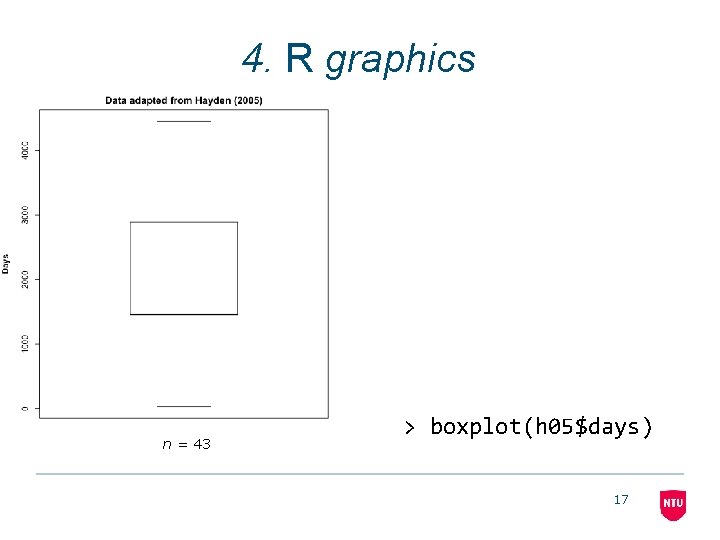
4. R graphics n = 43 > boxplot(h 05$days) 17
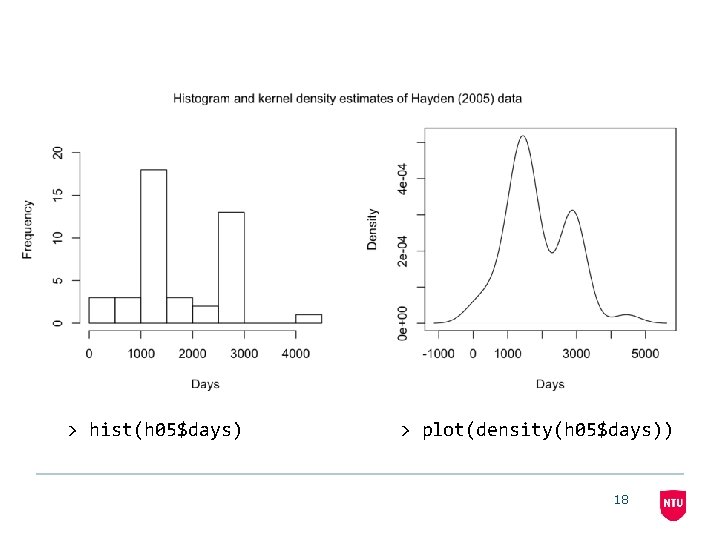
> hist(h 05$days) > plot(density(h 05$days)) 18

Distribution functions e. g. , Normal distribution dnorm(x, pnorm(q, qnorm(p, rnorm(n, mean = = 0, 0, sd sd = = 1) 1, lower. tail = TRUE) 1) > rdat <- rnorm(30, 100, 15) 19
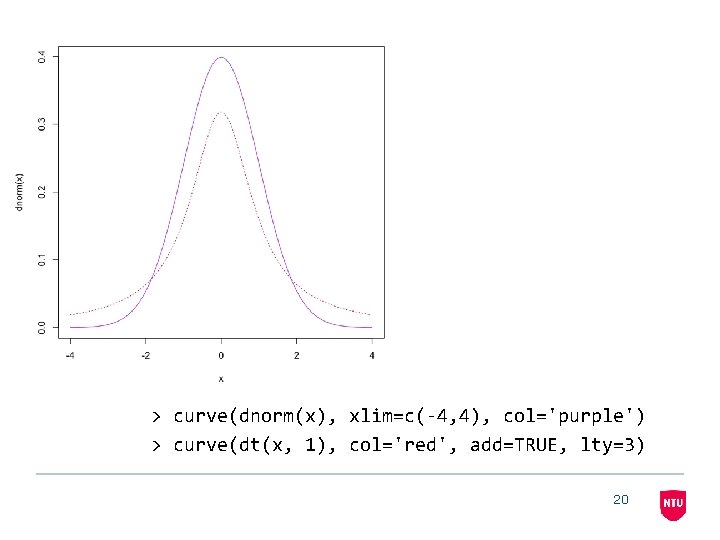
> curve(dnorm(x), xlim=c(-4, 4), col='purple') > curve(dt(x, 1), col='red', add=TRUE, lty=3) 20
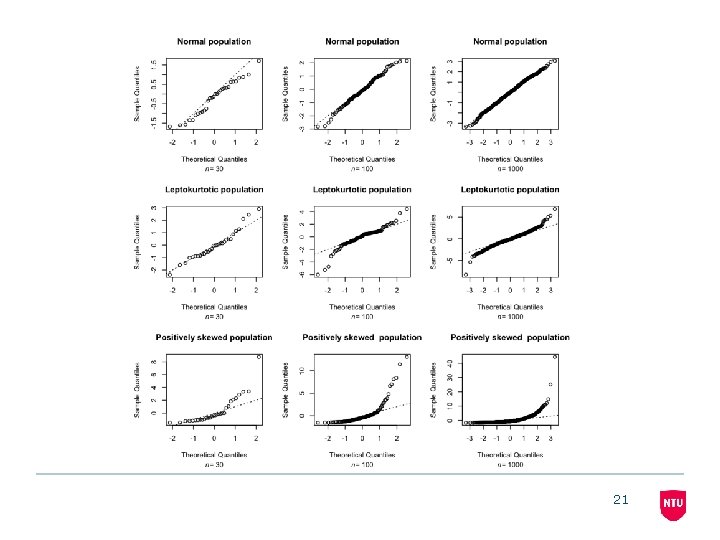
21

Exercise 2 (R Graphics) - exploratory plots - plot parameters - plotting distribution functions - plotting a serial position curve (optional) 22

5. Linear models in R > expenses <- read. csv('expenses. csv') > t. test(majority ~ problem, data = expenses) Welch Two Sample t-test data: majority by problem t = -3. 7477, df = 505. 923, p-value = 0. 000199 alternative hypothesis: true difference in means is not equal to 0 95 percent confidence interval: -2044. 457 -638. 160 sample estimates: mean in group 0 mean in group 1 7092. 712 8434. 021 Data courtesy of Mark Thompson http: //markreckons. blogspot. com/ 23

> expenses <- read. csv('expenses. csv') > lm(majority ~ problem, data = expenses) Call: lm(formula = majority ~ problem, data = expenses) Coefficients: (Intercept) 7093 problem 1341 24
![> lm. out <- lm(majority ~ problem, data = expenses) > max(rstandard(lm. out)) [1] > lm. out <- lm(majority ~ problem, data = expenses) > max(rstandard(lm. out)) [1]](http://slidetodoc.com/presentation_image/19771057d9bfebdb0dc44b36364f9ffb/image-25.jpg)
> lm. out <- lm(majority ~ problem, data = expenses) > max(rstandard(lm. out)) [1] 2. 629733 > AIC(lm. out) [1] 12674. 85 > lm. out$coefficients (Intercept) 7092. 712 problem 1341. 308 > ? lm 25

> summary(lm. out) Call: lm(formula = majority ~ problem, data = expenses) Residuals: Min 1 Q -8397. 0 -3403. 5 Median -260. 9 3 Q Max 3118. 8 11543. 3 Coefficients: Estimate Std. Error t value Pr(>|t|) (Intercept) 7092. 7 218. 9 32. 397 < 2 e-16 *** problem 1341. 3 357. 0 3. 758 0. 000187 *** --Signif. codes: 0 ‘***’ 0. 001 ‘**’ 0. 01 ‘*’ 0. 05 ‘. ’ 0. 1 ‘ ’ 1 Residual standard error: 4395 on 644 degrees of freedom Multiple R-squared: 0. 02145, Adjusted R-squared: 0. 01993 F-statistic: 14. 12 on 1 and 644 DF, p-value: 0. 0001871 26

> glm(problem~I(majority/10000), family='binomial', data = expenses) Call: glm(formula = problem ~ I(majority/10000), family = "binomial", data = expenses) Coefficients: (Intercept) I(majority/10000) -1. 0394 0. 6878 Degrees of Freedom: 645 Total (i. e. Null); Null Deviance: 644 Residual 855. 5 Residual Deviance: 841. 6 AIC: 845. 6 27
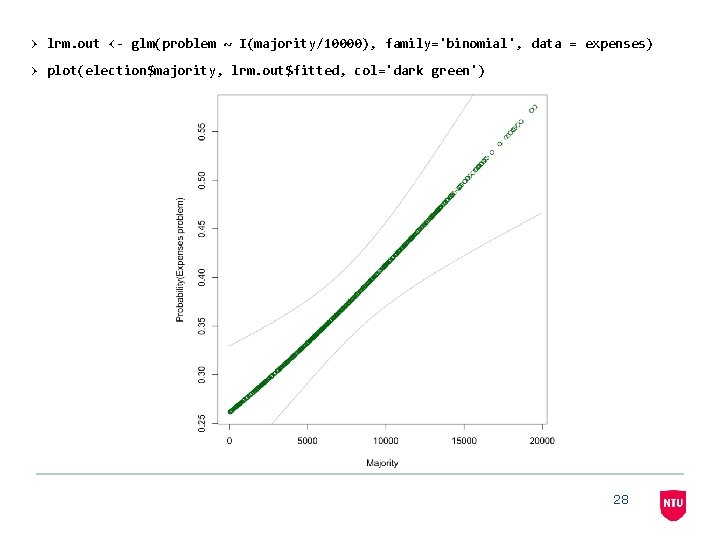
> lrm. out <- glm(problem ~ I(majority/10000), family='binomial', data = expenses) > plot(election$majority, lrm. out$fitted, col='dark green') 28

Exercise 3 (Linear models) - using a formula in a call to a model function - linear models - plotting a regression line - generalized linear models (optional) - plotting confidence bands (optional) 29
![6. ANOVA > factor 1 <- gl(10, 4) > factor 1 [1] 1 1 6. ANOVA > factor 1 <- gl(10, 4) > factor 1 [1] 1 1](http://slidetodoc.com/presentation_image/19771057d9bfebdb0dc44b36364f9ffb/image-30.jpg)
6. ANOVA > factor 1 <- gl(10, 4) > factor 1 [1] 1 1 1 1 1 2 2 2 2 2 3 3 3 3 3 4 4 4 4 4 Levels: 1 2 3 4 > library(foreign) > diag. data <- read. spss("diagram. sav", to. data. frame = T) > diag. fit <- lm(descript ~ group, data=diag. data) 30

Regression model (dummy coding) > diag. fit Call: lm(formula = descript ~ group, data = diag. data) Coefficients: (Intercept) group. Picture 17. 8 0. 5 group. Full Diagram 4. 6 group. Segmented 9. 5 31

aov() > aov(diag. fit) Call: aov(formula = diag. fit) Terms: group Residuals Sum of Squares Deg. of Freedom 583. 7 2440. 2 3 36 32

anova() > summary(aov(diag. fit)) > anova(diag. fit) Analysis of Variance Table Response: descript Df Sum Sq Mean Sq F value group 3 583. 7 194. 567 Residuals 36 2440. 2 Pr(>F) 2. 8704 0. 04977 * 67. 783 --Signif. codes: 0 ‘***’ 0. 001 ‘**’ 0. 01 ‘*’ 0. 05 ‘. ’ 0. 1 ‘ ’ 1 33

Factorial ANOVA and ANCOVA > lm(DV ~ factor 1 + factor 2 + factor 1: factor 2, data=data 2) > lm(DV ~ factor 1 * factor 2, data=data 2) > lm(descript ~ group + time, diag. data) > lm(descript ~ 0 + group + scale(time, scale=F), diag. data) > diag. ancov <- lm(descript ~ group + scale(time, scale=F), diag. data) 34

Type I and Type II Sums of squares > drop 1(diag. ancov, test = 'F') Single term deletions Model: descript ~ group + scale(time, scale = F) Df Sum of Sq <none> RSS AIC F value Pr(F) 2154. 4 169. 46 group 3 821. 45 2975. 9 176. 38 4. 4484 0. 009478 ** scale(time, scale = F) 1 285. 78 2440. 2 172. 44 4. 6427 0. 038149 * - sequential sums of squares (Type I) tests are given by default - hierarchical sums of squares (Type II) tests are given by the drop 1() function [Software such as SAS and SPSS defaults to unique (Type III) SS] 35

Repeated measures (within-subjects) factors - a weakness in the base R installation, but easily done using packages such as ez, nlme or lme 4 (the latter two able to fit a wider range of repeated measures models) > pride. long <- read. csv("pride_long. csv") > pride. mixed <- aov(accuracy ~ emotion*condition + Error(factor(participant)/emotion), pride. long) > summary(pride. mixed) [Software such as SAS and SPSS defaults to unique (Type III) SS] 36

Multiple comparisons and contrasts - you can run contrasts by changing default contrasts in R - Fisher LSD, Tukey’s HSD and p adjustment (e. g. , Hochberg, Holm, Bonferroni, FDR) in R base installation - powerful modified Bonferroni (e. g. , Shaffer, Westfall) and general linear contrasts available in multcomp package - flexible contrast and estimable functions in gmodels package 37

Exercise 4 (ANOVA) - factors - aov() and anova() - ANCOVA - repeated measures (within-subjects) - contrasts and multiple comparisons (optional) 38

7. Writing your own functions sd. pool <- function(sd. 1, n. 1, sd. 2, n. 2 = n. 1){ num <- (n. 1 -1)*sd. 1^2+(n. 2 -1)*sd. 2^2 denom <- n. 1+n. 2 -2 output <- sqrt(num/denom) output } > sd. pool(6. 1, 20, 5. 3, 25) [1] 5. 66743 39

Exercise 5 (Write a function) - pick a simple statistical procedure - write a function to automate it e. g. , pooled standard deviation 40

8. Simulation and bootstrapping - distribution functions > rnorm(100, 1) - R boot package - sample() and replicate() 41
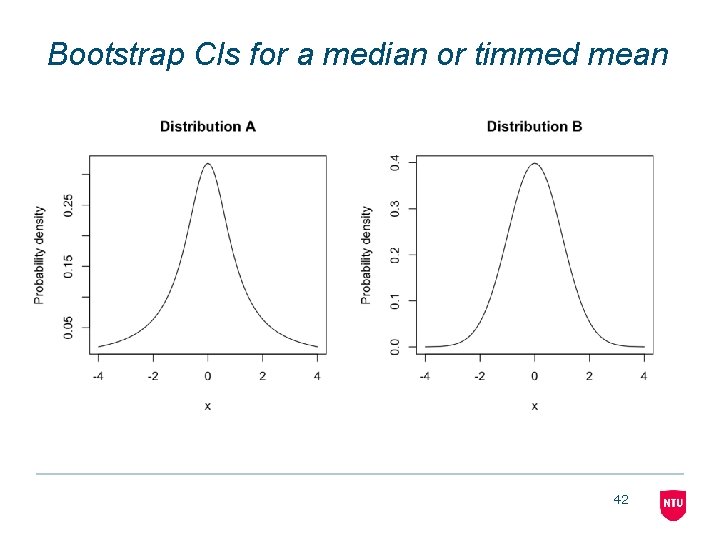
Bootstrap CIs for a median or timmed mean 42

The bootstrap (and other Monte Carlo methods) Bootstrapping involves repeatedly resampling with replacement from a data set e. g. , Data set = {0, 1} 7 simulated samples: > replicate(7, sample(x, 2, replace=TRUE)) {1, 0}, {0, 1}, {0, 0}, {1, 1} Bootstrapping effectively treats the sample distribution as the population distribution 43
![Bootstrapping using R > library(boot) > x 1. boot <- boot(samp, function(x, i) median(x[i]), Bootstrapping using R > library(boot) > x 1. boot <- boot(samp, function(x, i) median(x[i]),](http://slidetodoc.com/presentation_image/19771057d9bfebdb0dc44b36364f9ffb/image-44.jpg)
Bootstrapping using R > library(boot) > x 1. boot <- boot(samp, function(x, i) median(x[i]), R=10000) > boot. ci(x 1. boot) BOOTSTRAP CONFIDENCE INTERVAL CALCULATIONS Based on 10000 bootstrap replicates CALL : boot. ci(boot. out = x 1. boot) Intervals : Level Normal 95% (-0. 1747, 0. 6948 ) Basic (-0. 1648, 0. 6614 ) Level Percentile BCa 95% (-0. 1538, 0. 6724 ) (-0. 2464, 0. 6620 ) Calculations and Intervals on Original Scale Normal 95% CI for mean (using t) [-2. 00, 1. 33] 44

Exercise 6 (Bootstrapping) - resample data - construct a simple percentile bootstrap - run a BCa bootstrap using the boot package (optional) 45
46

9. Advanced statistical modelling in R Multilevel modeling nlme package lme 4 package MCMCglmm package Meta-analysis meta package metafor package … and many, many more other packages 47

metafor package > m. e <- c(10. 7, 16. 24, 10. 03, 3. 65, 5. 73) > n. e <- c(31, 57, 9, 17, 10) > m. c <- c(2. 87, 6. 83, 7. 18, 4. 65, 7. 47) > n. c <- c(14, 52, 9, 18, 10) > sd. pooled <- c(7. 88, 9. 84, 8. 67, 6. 34, 5. 74) > diff <- m. c - m. e > se. diffs <- sd. pooled * sqrt(1/n. e + 1/n. c) 48

> > install. packages('metafor') library(metafor) meta. out <- rma(yi=diff, sei=se. diffs, method = 'FE') meta. out Fixed-Effects Model (k = 5) Test for Heterogeneity: Q(df = 4) = 21. 0062, p-val = 0. 0003 Model Results: estimate -4. 1003 se 1. 0750 --Signif. codes: zval -3. 8141 pval 0. 0001 ci. lb -6. 2073 ci. ub -1. 9933 *** 0 ‘***’ 0. 001 ‘**’ 0. 01 ‘*’ 0. 05 ‘. ’ 0. 1 ‘ ’ 1 49
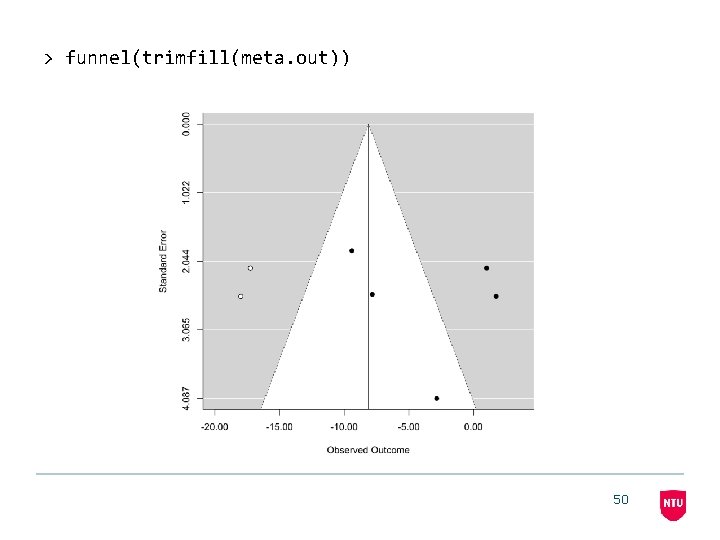
> funnel(trimfill(meta. out)) 50
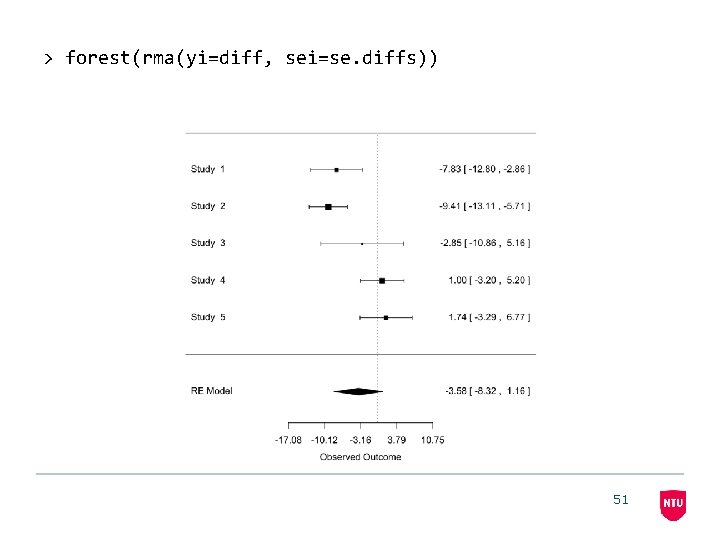
> forest(rma(yi=diff, sei=se. diffs)) 51
52
53
54
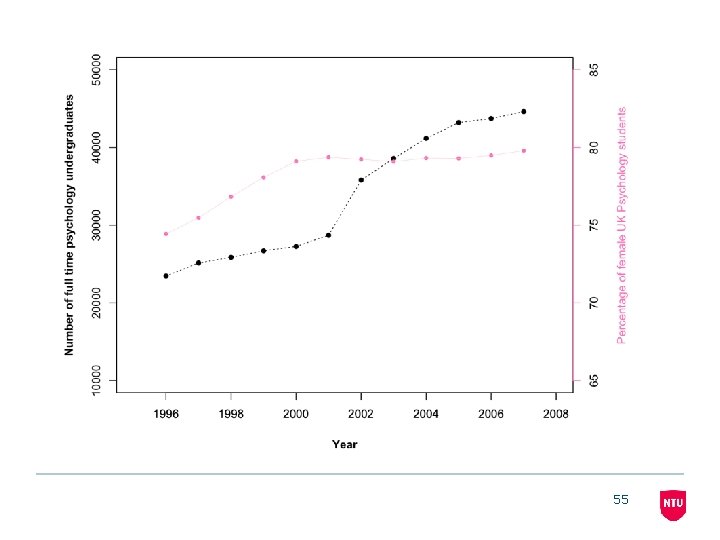
55
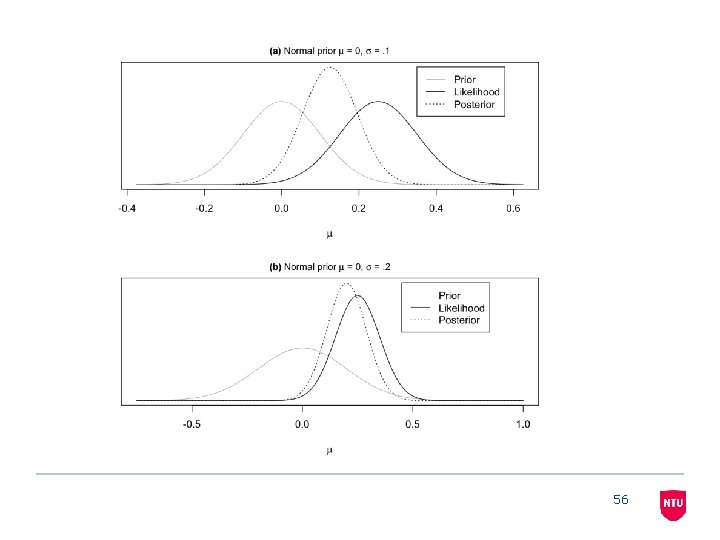
56
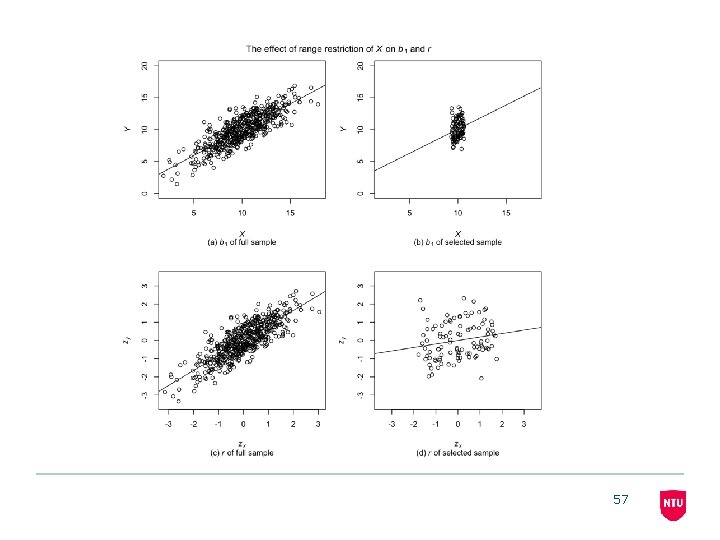
57
- Slides: 57