Quiz 22008 Chpt 3 ECB 1 A chemical

Quiz (2/20/08, Chpt 3, ECB) 1. A chemical process where there is a net gain of electrons is reduction called ________. A chemical process where there is a oxidation net loss of electrons is called _________. lower 2. Enzymes are catalysts, they often _______ the free energy of a reaction by favoring a transition state. 3. This nucleotide cofactor prominently featured as an electron NAD+ NADP+ carriers are _______ and _______. 4. This is the constant at which an enzyme is operating at half of Michalis-Menten constant its maximum speed. Km, ___________ 5. The maximum number of catalytic cycles an enzyme can perform per Turnover rate. unit time is called the _______

© Copyright 2008 Walt Disney Company Fluorescence Imaging with One Nanometer Accuracy (1. 5 nm, 1 -500 msec)
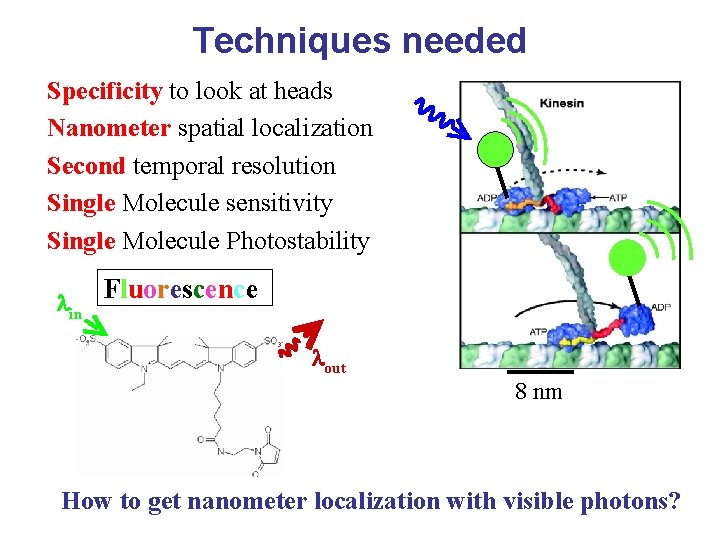
Techniques needed Specificity to look at heads Nanometer spatial localization Second temporal resolution Single Molecule sensitivity Single Molecule Photostability lin Fluorescence lout 8 nm How to get nanometer localization with visible photons?
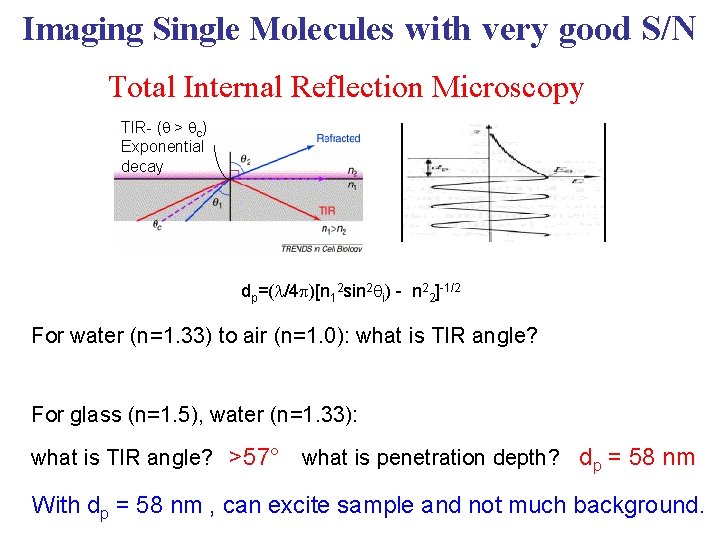
Imaging Single Molecules with very good S/N Total Internal Reflection Microscopy TIR- ( > c) Exponential decay dp=(l/4 p)[n 12 sin 2 i) - n 22]-1/2 For water (n=1. 33) to air (n=1. 0): what is TIR angle? For glass (n=1. 5), water (n=1. 33): what is TIR angle? >57° what is penetration depth? dp = 58 nm With dp = 58 nm , can excite sample and not much background.
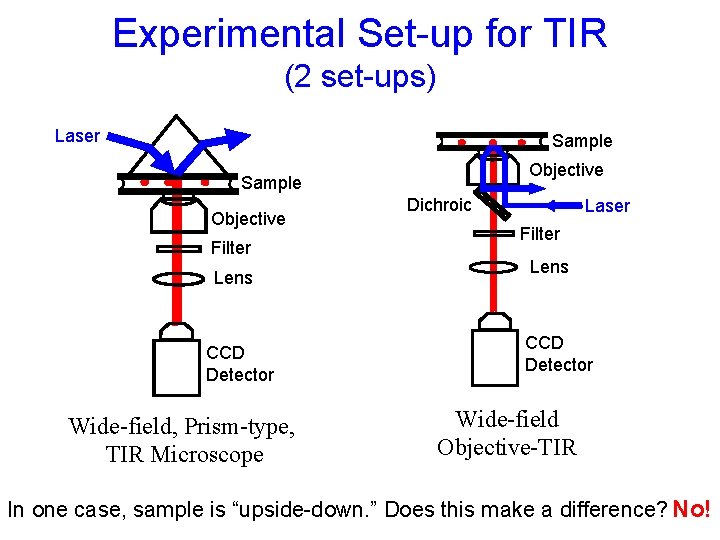
Experimental Set-up for TIR (2 set-ups) Laser Sample Objective Filter Lens CCD Detector Wide-field, Prism-type, TIR Microscope Dichroic Laser Filter Lens CCD Detector Wide-field Objective-TIR In one case, sample is “upside-down. ” Does this make a difference? No!
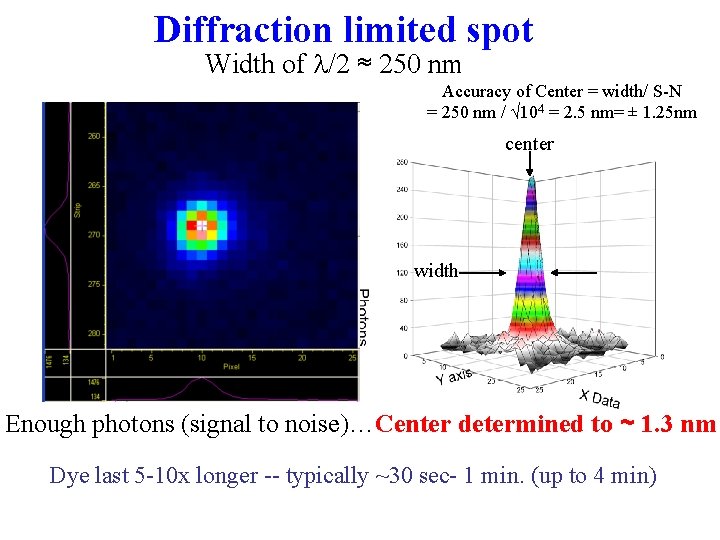
Diffraction limited spot Width of l/2 ≈ 250 nm Accuracy of Center = width/ S-N = 250 nm / √ 104 = 2. 5 nm= ± 1. 25 nm center width Enough photons (signal to noise)…Center determined to ~ 1. 3 nm Dye last 5 -10 x longer -- typically ~30 sec- 1 min. (up to 4 min)

How well can you localize? What does it depend on? (3 things) center 1. # of Photons Detected (N) 2. Pixel size of Detector (a) width 3. Noise (Background) of Detector (b) (includes background fluorescence and detector noise) = derived by Thompson et al. (Biophys. J. ).
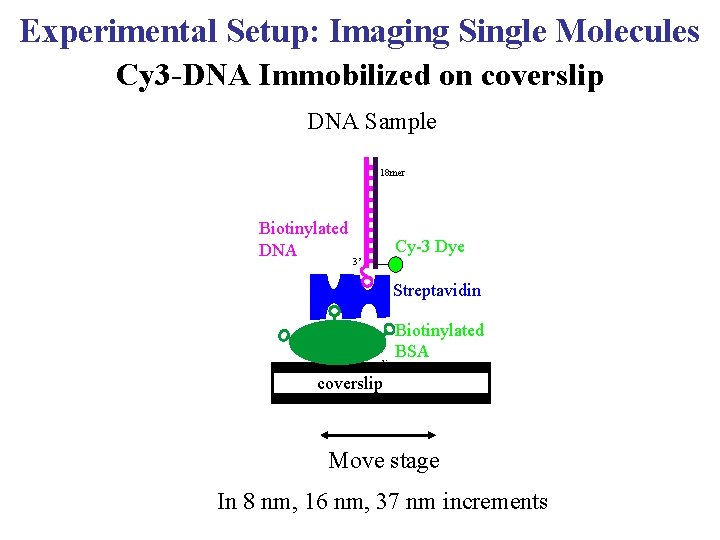
Experimental Setup: Imaging Single Molecules Cy 3 -DNA Immobilized on coverslip DNA Sample 18 mer Biotinylated DNA Cy-3 Dye 3’ Streptavidin coverslip Biotinylated BSA coverslip Move stage In 8 nm, 16 nm, 37 nm increments
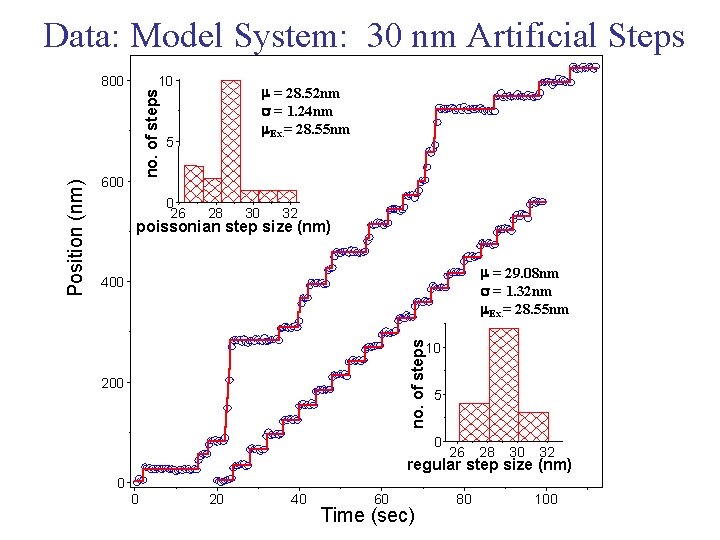
Data: Model System: 30 nm Artificial Steps 10 no. of steps 600 m = 28. 52 nm s = 1. 24 nm m. Ex. = 28. 55 nm 5 0 26 28 30 32 poissonian step size (nm) m = 29. 08 nm s = 1. 32 nm m. Ex. = 28. 55 nm 400 no. of steps Position (nm) 800 10 200 5 0 26 28 30 32 regular step size (nm) 0 0 20 40 60 Time (sec) 80 100
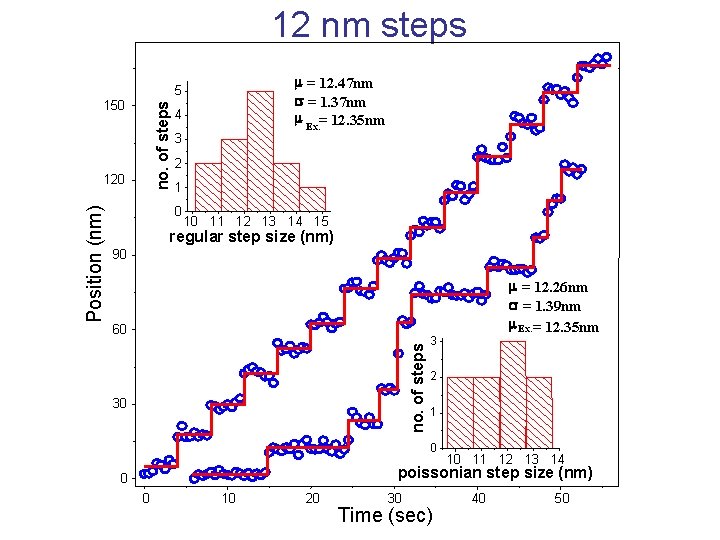
12 nm steps no. of steps 150 120 4 3 2 1 0 10 11 12 13 14 15 regular step size (nm) 90 60 no. of steps Position (nm) m = 12. 47 nm s = 1. 37 nm m Ex. = 12. 35 nm 5 30 m = 12. 26 nm s = 1. 39 nm m Ex. = 12. 35 nm 3 2 1 0 10 11 12 13 14 poissonian step size (nm) 0 0 10 20 30 Time (sec) 40 50
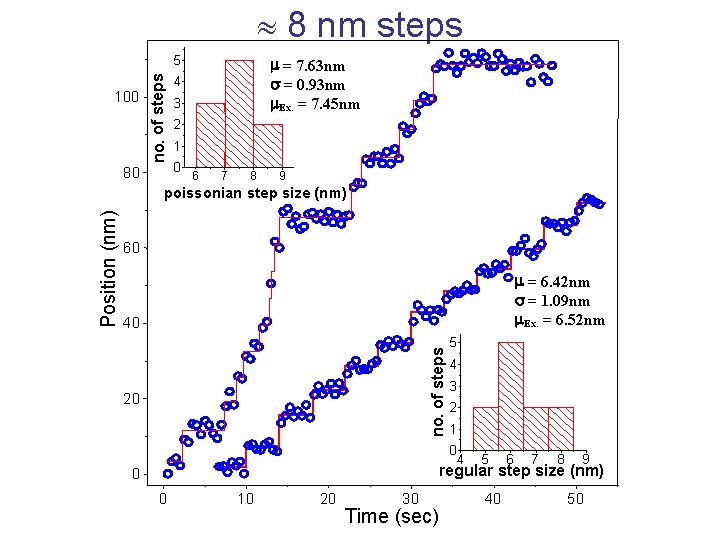
8 nm steps 100 no. of steps 5 80 m = 7. 63 nm s = 0. 93 nm m. Ex. = 7. 45 nm 4 3 2 1 0 6 7 8 9 60 m = 6. 42 nm s = 1. 09 nm m. Ex. = 6. 52 nm 40 no. of steps Position (nm) poissonian step size (nm) 20 5 4 3 2 1 0 4 5 6 7 8 9 regular step size (nm) 0 0 10 20 30 Time (sec) 40 50
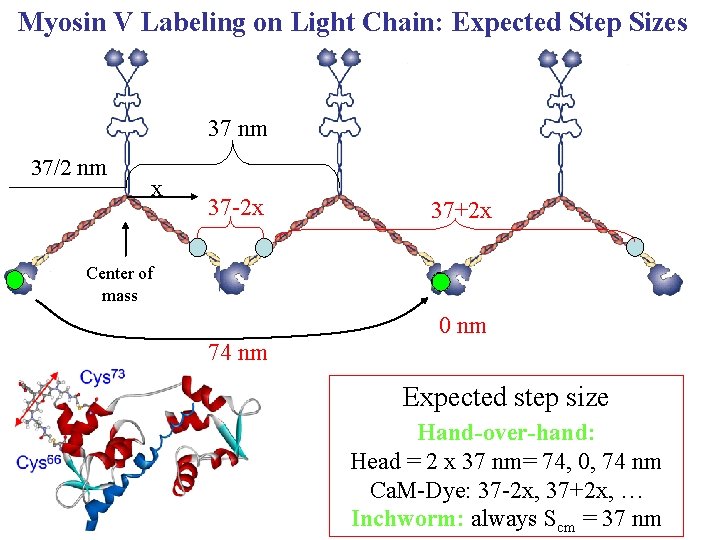
Myosin V Labeling on Light Chain: Expected Step Sizes 37 nm 37/2 nm x 37 -2 x 37+2 x Center of mass 0 nm 74 nm Expected step size Hand-over-hand: Head = 2 x 37 nm= 74, 0, 74 nm Ca. M-Dye: 37 -2 x, 37+2 x, … Inchworm: always Scm = 37 nm
![A Single Myosin V moving [ATP] = 300 n. M (Low) 86 nm pixel A Single Myosin V moving [ATP] = 300 n. M (Low) 86 nm pixel](http://slidetodoc.com/presentation_image_h/b1cffc51ebe5d48204c0b9a1cfabc195/image-13.jpg)
A Single Myosin V moving [ATP] = 300 n. M (Low) 86 nm pixel 37 nm or 74 nm?
![Myosin V steps: 74 nm +/- 5 nm [ATP] = 300 n. M higher Myosin V steps: 74 nm +/- 5 nm [ATP] = 300 n. M higher](http://slidetodoc.com/presentation_image_h/b1cffc51ebe5d48204c0b9a1cfabc195/image-14.jpg)
Myosin V steps: 74 nm +/- 5 nm [ATP] = 300 n. M higher [ATP] (>400 n. M) +/- 1. 3 nm +/- 1. 5 nm +/- 1. 3 nm
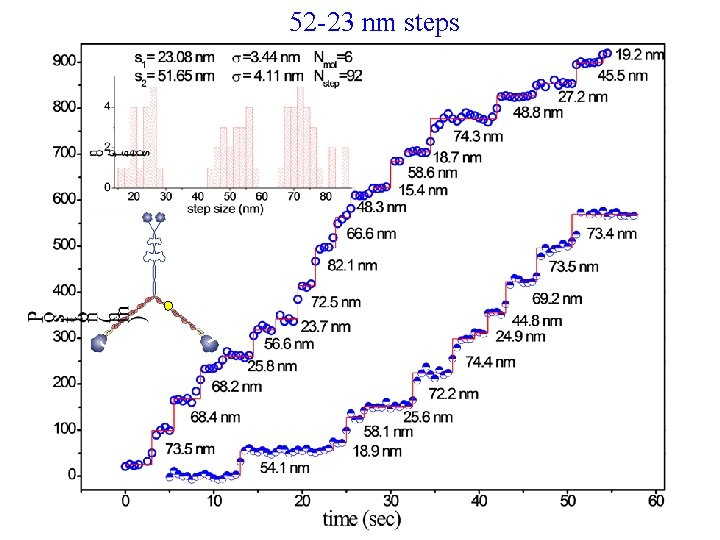
52 -23 nm steps
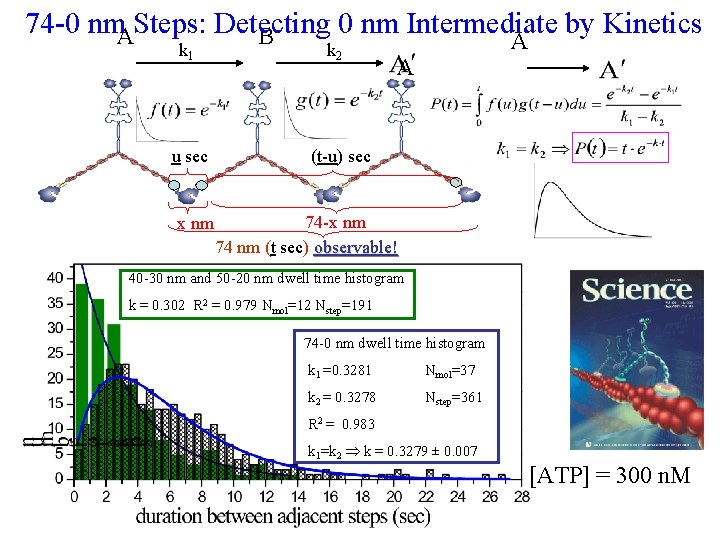
74 -0 nm. ASteps: Detecting 0 nm Intermediate by Kinetics B k 1 u sec x nm k 2 A A (t-u) sec 74 -x nm 74 nm (t sec) observable! 40 -30 nm and 50 -20 nm dwell time histogram k = 0. 302 R 2 = 0. 979 Nmol=12 Nstep=191 74 -0 nm dwell time histogram k 1 =0. 3281 Nmol=37 k 2 = 0. 3278 Nstep=361 R 2 = 0. 983 k 1=k 2 k = 0. 3279 ± 0. 007 [ATP] = 300 n. M

Class evaluation 1. What was the most interesting thing you learned in class today? 2. What are you confused about? 3. Related to today’s subject, what would you like to know more about? 4. Any helpful comments. Answer, and turn in at the end of class.
- Slides: 17