Quantifying the distribution of variation Variation within individuals

Quantifying the distribution of variation Variation within individuals within subpopulations among subpopulations (in total population)

Quantifying the distribution of variation Variation within individuals within subpopulations among subpopulations (in total population)

Quantifying the distribution of variation I = individuals S = subpopulations T = total population H is observed heterozygosity (# heterozygotes/N) in a population HI is observed heterozygosity (# heterozygotes/N) averaged over individuals in all subpopulations HS is expected heterozygosity in each subpopulation if it was in H-W equilibrium, averaged across subpopulations HT is expected heterozygosity if subpopulations are combined as one population

Quantifying the distribution of variation I = individuals S = subpopulations T = total population H is observed heterozygosity (# heterozygotes/N) in a population HI is observed heterozygosity (# heterozygotes/N) averaged over individuals in all subpopulations HS is expected heterozygosity in each subpopulation if it was in H-W equilibrium, averaged across subpopulations HT is expected heterozygosity if subpopulations are combined as one population HT HS HS HS

Quantifying the distribution of variation HI observed heterozygosity averaged over individuals in all subpopulations HS expected heterozygosity in each subpopulation (if it was in H-W equilib. ) averaged across subpopulations HT expected heterozygosity if subpopulations are combined as one population p AA AA Aa aa aa 0. 5 H 1/5 = 0. 2 Aa Aa AA 0. 6 4/5 = 0. 8

Quantifying the distribution of variation HI observed heterozygosity averaged over individuals in all subpopulations HS expected heterozygosity in each subpopulation (if it was in H-W equilib. ) averaged across subpopulations HT expected heterozygosity if subpopulations are combined as one population p AA AA Aa aa aa 0. 5 H 1/5 = 0. 2 HI Aa Aa AA 0. 6 4/5 = 0. 8 (0. 2 + 0. 8)/2 = 0. 5
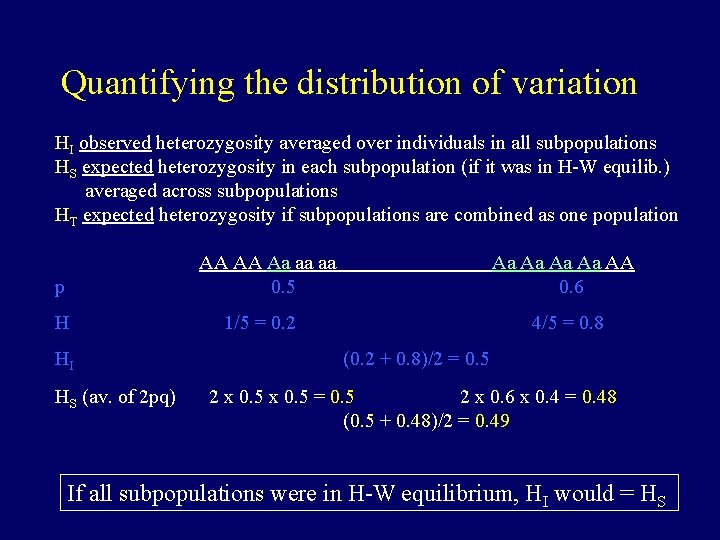
Quantifying the distribution of variation HI observed heterozygosity averaged over individuals in all subpopulations HS expected heterozygosity in each subpopulation (if it was in H-W equilib. ) averaged across subpopulations HT expected heterozygosity if subpopulations are combined as one population p AA AA Aa aa aa 0. 5 H 1/5 = 0. 2 HI HS (av. of 2 pq) Aa Aa AA 0. 6 4/5 = 0. 8 (0. 2 + 0. 8)/2 = 0. 5 2 x 0. 5 = 0. 5 2 x 0. 6 x 0. 4 = 0. 48 (0. 5 + 0. 48)/2 = 0. 49 If all subpopulations were in H-W equilibrium, HI would = HS

Quantifying the distribution of variation HI observed heterozygosity averaged over individuals in all subpopulations HS expected heterozygosity in each subpopulation (if it was in H-W equilib. ) averaged across subpopulations HT expected heterozygosity if subpopulations are combined as one population p AA AA Aa aa aa 0. 5 H 1/5 = 0. 2 HI HS (av. of 2 pq) HT (2 x pav x qav) Aa Aa AA 0. 6 4/5 = 0. 8 (0. 2 + 0. 8)/2 = 0. 5 2 x 0. 5 = 0. 5 2 x 0. 6 x 0. 4 = 0. 48 (0. 5 + 0. 48)/2 = 0. 49 2 x 0. 55 x 0. 45 = 0. 495

Quantifying the distribution of variation HI observed heterozygosity averaged over individuals in all subpopulations HS expected heterozygosity in each subpopulation (if it was in H-W equilib. ) averaged across subpopulations HT expected heterozygosity if subpopulations are combined as one population p AA AA Aa aa aa 0. 5 H 1/5 = 0. 2 HI Aa Aa AA 0. 6 4/5 = 0. 8 (0. 2 + 0. 8)/2 = 0. 5 HS (av. of 2 pq) HT (2 x pav x qav) 2 x 0. 5 = 0. 5 2 x 0. 6 x 0. 4 = 0. 48 (0. 5 + 0. 48)/2 = 0. 49 2 x 0. 55 x 0. 45 = 0. 495 If all subpopulations were the same, HS would = HT

HI observed heterozygosity averaged over individuals in all subpopulations HS expected heterozygosity in each subpopulation averaged across subpopulations HT expected heterozygosity if subpopulations are combined as one population FIT – reduction in heterozygosity of individuals relative to whole population FIT = HT – HI HT

HI observed heterozygosity averaged over individuals in all subpopulations HS expected heterozygosity in each subpopulation averaged across subpopulations HT expected heterozygosity if subpopulations are combined as one population FIT – reduction in heterozygosity of individuals relative to whole population FIT = HT – HI HT - any departure from single panmictic population will lead to significant value - used to detect departures from Hardy-Weinberg equilibrium in total population

HI observed heterozygosity averaged over individuals in all subpopulations HS expected heterozygosity in each subpopulation averaged across subpopulations HT expected heterozygosity if subpopulations are combined as one population FIS - inbreeding coefficient of individual relative to its subpopulation (change in H due to non-random mating) FIS = HS – HI HS

HI observed heterozygosity averaged over individuals in all subpopulations HS expected heterozygosity in each subpopulation averaged across subpopulations HT expected heterozygosity if subpopulations are combined as one population FIS - inbreeding coefficient of individual relative to its subpopulation FIS = HS – HI HS used to detect departures from Hardy-Weinberg equilibrium in "good" populations positive value = heterozygote deficiency (Wahlund effect) zero value = all sub-populations in Hardy-Weinberg equilibrium (random mating within subpopulations) negative value = heterozygote excess

HI observed heterozygosity averaged over individuals in all subpopulations HS expected heterozygosity in each subpopulation averaged across subpopulations HT expected heterozygosity if subpopulations are combined as one population FST - inbreeding coefficient of sub-population relative to the whole population = fixation index FST = HT – HS HT

HI observed heterozygosity averaged over individuals in all subpopulations HS expected heterozygosity in each subpopulation averaged across subpopulations HT expected heterozygosity if subpopulations are combined as one population FST - inbreeding coefficient of sub-population relative to the whole population = fixation index FST = HT – HS HT measures degree of population differentiation within species (always positive) 0. 00 = sub-popns have same allele frequencies 0. 05 -0. 15 = moderate differentiation 0. 15 -0. 25 = great differentiation >0. 25 = extremely different 1. 0 = popns fixed for different alleles
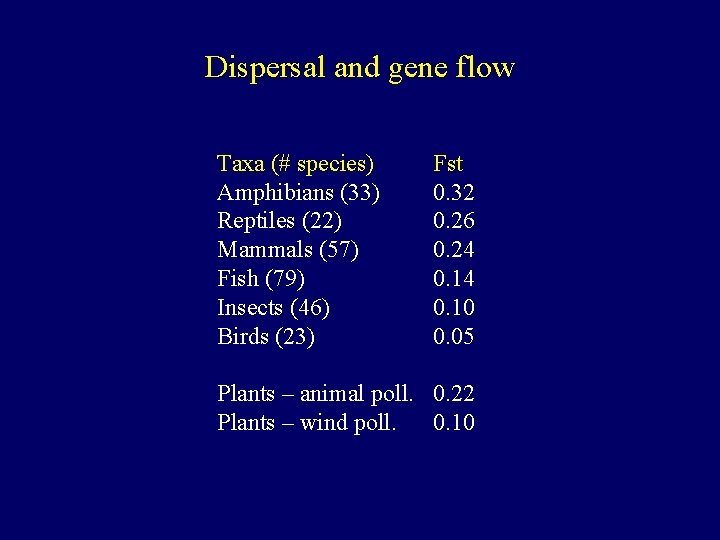
Dispersal and gene flow Taxa (# species) Amphibians (33) Reptiles (22) Mammals (57) Fish (79) Insects (46) Birds (23) Fst 0. 32 0. 26 0. 24 0. 10 0. 05 Plants – animal poll. 0. 22 Plants – wind poll. 0. 10

Quantifying the distribution of variation HI observed heterozygosity averaged over individuals in all subpopulations (# heterozygotes/ total N) HS expected heterozygosity in each subpopulation (if it was in H-W equilib. ) averaged across subpopulations (# heterozygotes/total N) HT expected heterozygosity if subpopulations are combined as one population AA, AA BB, BB, BB HI HS HT 0 0 0. 5 AA, AB, AB, BB 0. 5 AB, AB, AB, AB 1 0. 5 AA, BB, BB AB, AA, BB 0. 25 0. 44 0. 47

Quantifying the distribution of variation FIS - inbreeding coefficient of individual relative to its subpopulation FIS = HS – HI HS positive value = heterozygote deficiency zero value = all sub-populations in Hardy-Weinberg equilibrium negative value = heterozygote excess AA, AA BB, BB, BB HI HS HT 0 0 0. 5 FIS 0 AA, AB, AB, BB 0. 5 0 AB, AB, AB, AB 1 0. 5 -1 AA, BB, BB AB, AA, BB 0. 47 0. 25 0. 44 0. 4

9 populations examined at 21 loci FIS = 0. 49 high level of inbreeding? dioecious – so likely due to clustering of relatives Pacific yew (Taxus brevifolia)

Quantifying the distribution of variation FIT - reduction in heterozygosity of individuals relative to whole population FIT = HT – HI HT used to detect departures from Hardy-Weinberg equilibrium in total population AA, AA BB, BB, BB HI HS HT 0 0 0. 5 FIT 1 AA, AB, AB, BB 0. 5 0 AB, AB, AB, AB 1 0. 5 -1 AA, BB, BB AB, AA, BB 0. 47 0. 25 0. 44 0. 4

Quantifying the distribution of variation FST - inbreeding coefficient of sub-population relative to the whole population FST = HT – HS HT measures degree of population differentiation within species (always positive) AA, AA BB, BB, BB HI HS HT 0 0 0. 5 FST 1 AA, AB, AB, BB 0. 5 0 AB, AB, AB, AB 1 0. 5 0 AA, BB, BB AB, AA, BB 0. 47 0. 25 0. 44 0. 0

9 populations examined at 21 loci FST = 0. 078 Low to moderate level of population differentiation Pacific yew (Taxus brevifolia)
- Slides: 22