Quality and mappability of K 562GM stranded pairedend

Quality and mappability of K 562/GM stranded paired-end libraries Gingeras lab 1
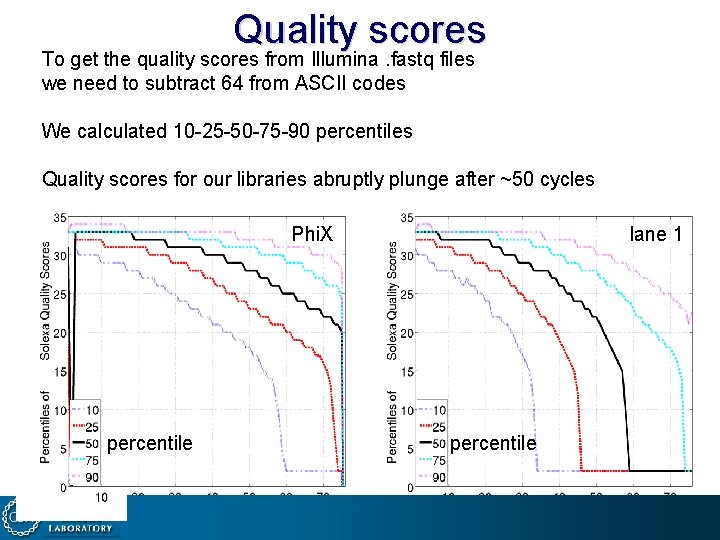
Quality scores To get the quality scores from Illumina. fastq files we need to subtract 64 from ASCII codes We calculated 10 -25 -50 -75 -90 percentiles Quality scores for our libraries abruptly plunge after ~50 cycles Phi. X percentile lane 1 percentile 2
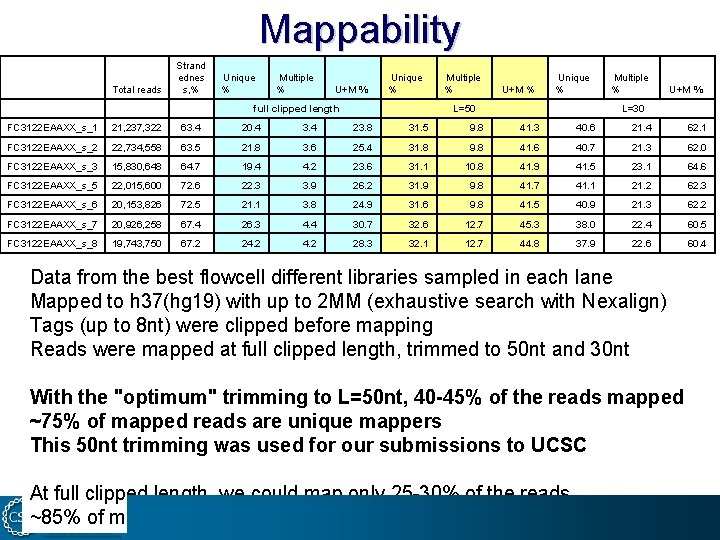
Mappability Strand ednes s, % Unique % Multiple % FC 3122 EAAXX_s_1 21, 237, 322 63. 4 20. 4 3. 4 23. 8 31. 5 9. 8 41. 3 40. 6 21. 4 62. 1 FC 3122 EAAXX_s_2 22, 734, 558 63. 5 21. 8 3. 6 25. 4 31. 8 9. 8 41. 6 40. 7 21. 3 62. 0 FC 3122 EAAXX_s_3 15, 830, 648 64. 7 19. 4 4. 2 23. 6 31. 1 10. 8 41. 9 41. 5 23. 1 64. 6 FC 3122 EAAXX_s_5 22, 015, 600 72. 6 22. 3 3. 9 26. 2 31. 9 9. 8 41. 7 41. 1 21. 2 62. 3 FC 3122 EAAXX_s_6 20, 153, 826 72. 5 21. 1 3. 8 24. 9 31. 6 9. 8 41. 5 40. 9 21. 3 62. 2 FC 3122 EAAXX_s_7 20, 926, 258 67. 4 26. 3 4. 4 30. 7 32. 6 12. 7 45. 3 38. 0 22. 4 60. 5 FC 3122 EAAXX_s_8 19, 743, 750 67. 2 24. 2 28. 3 32. 1 12. 7 44. 8 37. 9 22. 6 60. 4 full clipped length U+M % Unique % Total reads U+M % Unique % L=50 U+M % L=30 Data from the best flowcell different libraries sampled in each lane Mapped to h 37(hg 19) with up to 2 MM (exhaustive search with Nexalign) Tags (up to 8 nt) were clipped before mapping Reads were mapped at full clipped length, trimmed to 50 nt and 30 nt With the "optimum" trimming to L=50 nt, 40 -45% of the reads mapped ~75% of mapped reads are unique mappers This 50 nt trimming was used for our submissions to UCSC At full clipped length, we could map only 25 -30% of the reads ~85% of mapped reads are unique mappers 3
- Slides: 3