QTL Mapping in Humans Lon Cardon Stacey Cherny





- Slides: 8

QTL Mapping in Humans Lon Cardon, Stacey Cherny University of Oxford Goncalo Abecasis University of Michigan Pak Sham Institute of Psychiatry, King’s College, London

3 Stages of Genetic Mapping • Are there genes influencing this trait? – Epidemiological studies • Where are those genes? – Linkage analysis • What are those genes? – Association analysis

3 Stages of Genetic Mapping • Are there genes influencing this trait? – Epidemiological studies • Where are those genes? – Linkage analysis • What are those genes? – Association analysis

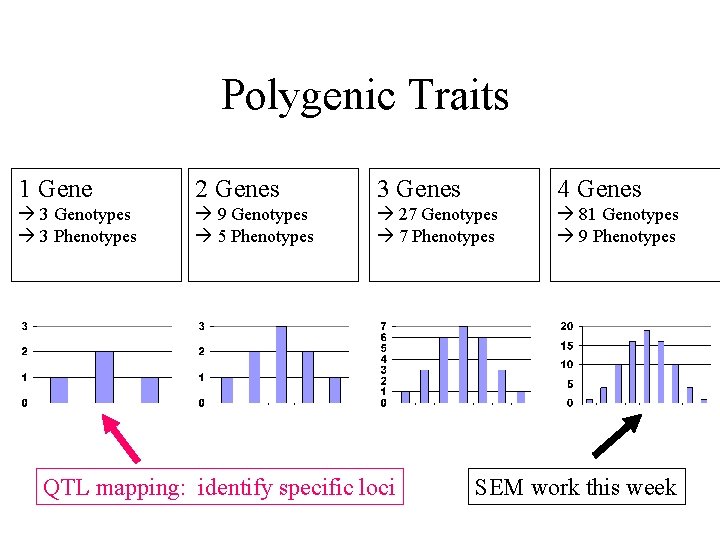
Polygenic Traits 1 Gene 2 Genes 3 Genes 4 Genes 3 Genotypes 3 Phenotypes 9 Genotypes 5 Phenotypes 27 Genotypes 7 Phenotypes 81 Genotypes 9 Phenotypes QTL mapping: identify specific loci SEM work this week

QTL Linkage Analysis • Aim: Identify regions of genome that contain (‘are linked with’) gene(s) influencing trait • Follows directly from biometrical model – Builds on all concepts developed this week – Twins informative, as are extended sibships larger kindreds • Only one new data type added to design: genotypes – ‘identity by descent’ derived from genotypes

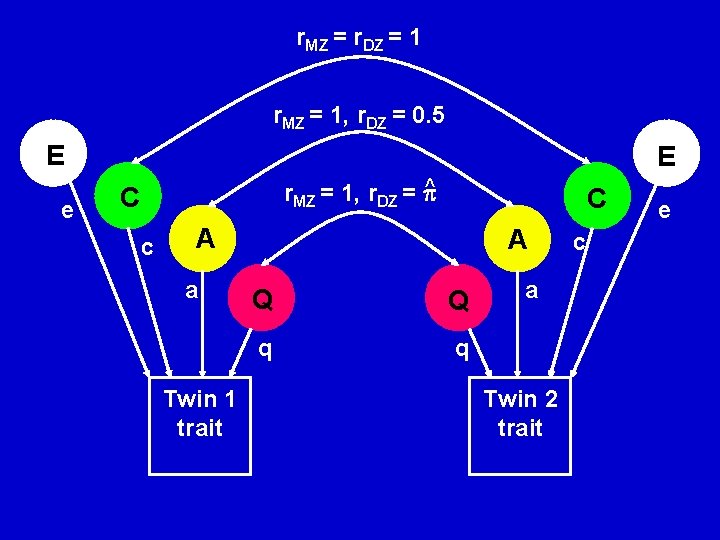
r. MZ = r. DZ = 1 r. MZ = 1, r. DZ = 0. 5 E e E ^ r. MZ = 1, r. DZ = C c C A a Twin 1 trait A Q Q q q a Twin 2 trait c e

Some QTL linkage analysis software programs • ACT: Chris Amos’ variance components suite • Genehunter: Leonid Kruglyak’s discrete-trait package, extended to simple VC • SOLAR: John Blangero’s variance components package • QTDT/Merlin: Goncalo Abecasis’ association/linkage VC program and likelihood engine • Many others, large and small • Mx: Mike Neale’s variance components program

Friday afternoon schedule • 1: 30 -2: 30: Linkage analysis theory: Abecasis/Sham • 2: 30 -3: 00: QTL linkage analysis in Mx: Cardon/Cherny • 3: 00 -3: 30: Tea • 3: 30 -4: 00: Multivariate QTL linkage analysis: Cardon/Cherny • 4: 00 -4: 45: Beyond linkage: some current issues in QTL identification (aka: gratuitous promotion of advanced workshop): Cardon