QSite Finder an energybased method for the prediction

Q-Site. Finder: an energy-based method for the prediction of protein-ligand binding sites Bioinformatics Vol. 21 no. 9 2005 (Pages 1908 -1916) Reporter: Yu Lun Kuo (D 95922037) E-mail: sscc 6991@gmail. com Date: June 5, 2008

Motivation • 3 D structure available for protein whose interaction with small molecules (ligands) are not known – Describe a new method of ligand binding site prediction called Q-Site. Finder • Use the interaction energy 2

Introduction • Goal – Given a protein structure, predicts its ligand bindings • Flexible ligand docking • Lead optimisation • Applicat ion – Function prediction – Drug discovery – etc. 3
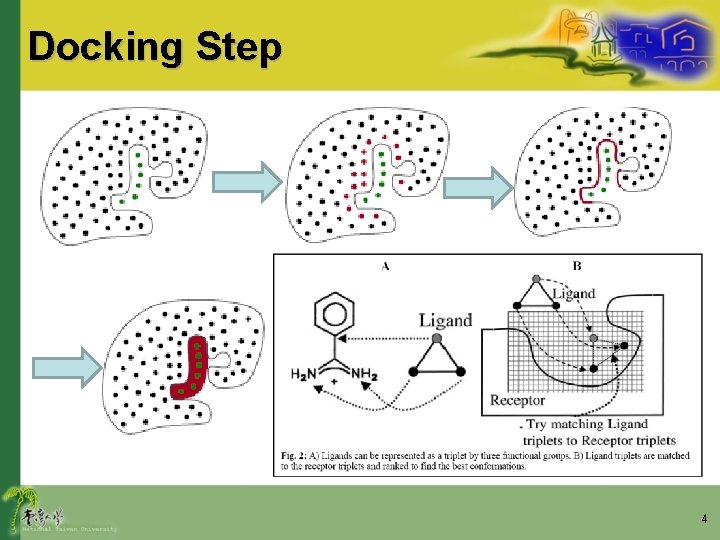
Docking Step 4
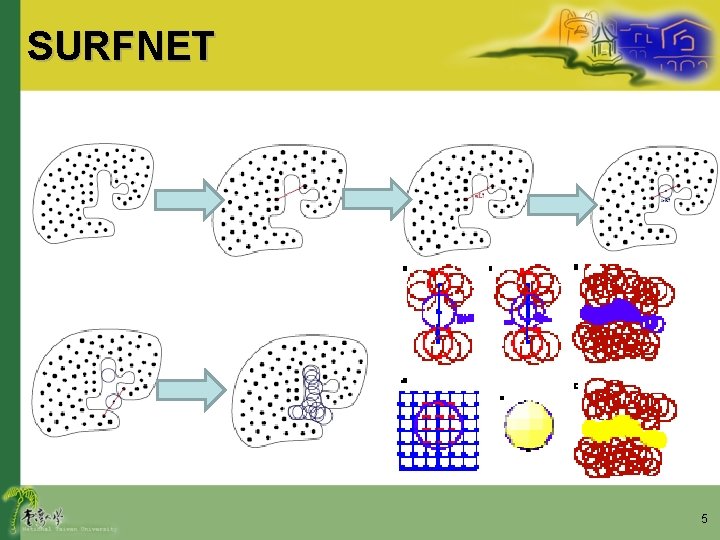
SURFNET 5

SURFNET 6

Introduction • Detection and characterization of functional sites on protein – Identify functional sites • In addition de novo drug design – Lead to the creation of novel ligands not found in molecular databases 7

Introduction • The ligand binding site is usually in the largest pocket – SURFNET (Laskowski et al. , 1996) • The ligand binding site was found to be in the largest pocket in 83% of cases – LIGSITE (Hendlich et al. , 1997) • The ligand binding site was found in the largest pocket in all 10 proteins tested – etc. 8

Introduction • Q-Site. Finder – Defined only by energetic criteria • Calculates the van der Waals interaction energies of a methyl probe with the protein • Probes are ranked according to their total interaction energies 9

Introduction • Several techniques have been developed for estimating the interaction energy – GRID (Wade and Goodford) • Identify the hydrogen bonding potential of drug-like molecules • The interaction energies – Using a conventional molecular mechanics function • Van der Waals, electrostatic, and solvation terms 10

Introduction • Q-Site. Finder – Keep the predicted ligand binding site as small as possible without compromising accuracy – Provide a threshold for success 11

Methods • Datasets – Consisted of 134 records obtained from the PDB • Correspond to the GOLD protein-ligand docking dataset (305 proteins) • Remove those with high levels of structural similarity – Which could bias the results – Solvent molecules were discarded • Phosphate, sulphate and metal ions • Q-Site. Finder is not designed to detect the binding site of small solvent molecules 12

Q-Site. Finder • Simply uses the van der Waals interaction (of a methyl probe) and an interaction energy threshold to determine favourable binding clefts 13

Results (Q-Site. Finder) • Define a successful prediction using a precision threshold – A threshold of 25% precision was used to define success in al the result here • A precision of 26% is considered a success • 17% is not 14
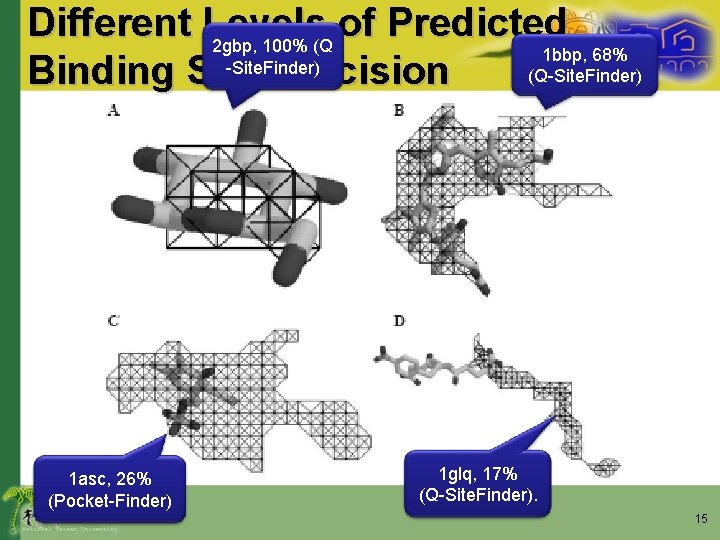
Different Levels of Predicted 2 gbp, 100% (Q 1 bbp, 68% -Site. Finder) (Q-Site. Finder) Binding Site Precision 1 asc, 26% (Pocket-Finder) 1 glq, 17% (Q-Site. Finder). 15

Results (Q-Site. Finder) • If a ligand is successfully predicted in more than one site on a protein – It is counted as a success only in the higher ranking site – If more than one ligand is found in the same site • Only the success with the highest precision is counted for this site 16
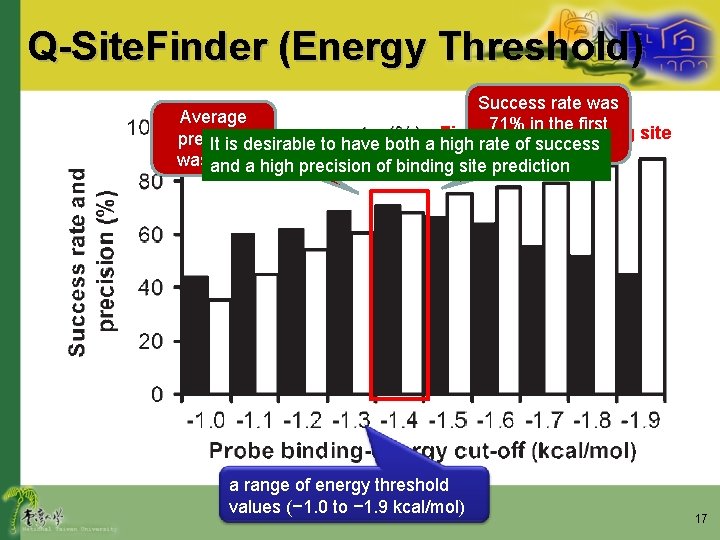
Q-Site. Finder (Energy Threshold) Success rate was Average Precision of first 0% 71% in the First predicted binding site precision excluded It is desirable to have both a highwere rate predicted of success wasand 68%a high precision of binding site prediction a range of energy threshold values (− 1. 0 to − 1. 9 kcal/mol) 17

Results (Pocket-Finder) • Use a variable, MINPSP – PSP (protein-site-protein) • A pocket is identified if an interaction occurs followed by a period of no interaction, followed by another interaction • Measure the extent to which each grid point is buried in the protein • Each grid point has seven scanning lines passing through it – x, y and z direction and the four cubic diagonals 18

Results (Picket-Finder) • MINPSP (minimum number of PSP) – Thought of a burial threshold – PSP values for each grid point vary from 0 to 7 • 0: not a pocket • 7: deeply buried 19
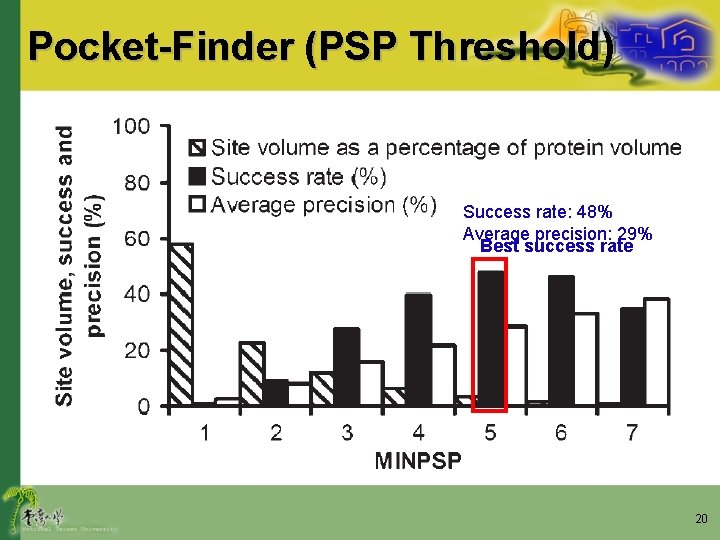
Pocket-Finder (PSP Threshold) Success rate: 48% Average precision: 29% Best success rate 20

Results • Hendlich et al. (1997) – Recommend a MINPSP of 2 • Our implementation of Pocket-Finder – Low average precision: 8% – Large site volume: 8700 A 3 (23% of the average protein volume) • No significant benefit in the success rate was observed on using a MINPSP of 2 rather than 5 21

Results • Smaller sites have a higher average precision – Sites with high volume will usually incorporate locations on the protein surface • That are not part of binding site 22

Comparison • Q-Site. Finder – Energy threshold value: -1. 4 kcal/mol • Success rate: 71% average precision: 68% – At least one successful prediction in the • Top three predicted sites for 90% of the proteins • Top ten predicted sites for 96% of the proteins • Pocket-Finder – MINPSP threshold of 5 • Success rate: 48% average precision: 29% – At least one successful prediction in the • Top three predicted sites for 65% of the proteins • Top ten predicted sites for 74% of the proteins 23

Comparison of the success rates • Q-Site. Finder has a higher success rate in each of the top three predicted binding sites 24
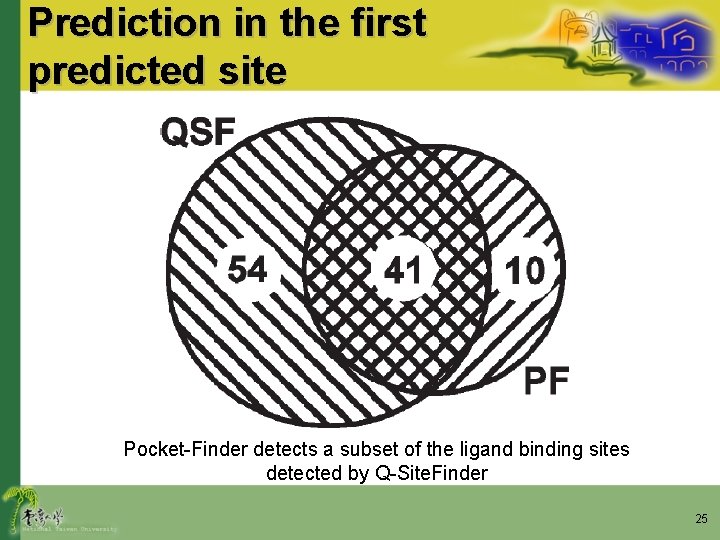
Prediction in the first predicted site Pocket-Finder detects a subset of the ligand binding sites detected by Q-Site. Finder 25

Application of Q-Site. Finder rate in thebinding first predictedsites on • Q-Site. Finder for Success detecting Unbound state: 51% state: 80% unbound protein Ligand-bound The average precision of the first predicted binding site 71% for the unbound state At least one success 74% for the ligand-bound state. in the top 3 Unbound state: 86% Ligand-bound state: 97% 26

Average Volume of Successfully Predicted Sites • Relax our threshold to allow any non-zero value (success requires a precision > 0%) Average precision of Pocket-Finder is 29% Q-Site. Finder is 68% Q-Site. Finder would appear to be more robust than Pocket-Finder, and better able to pinpoint the location of the ligand binding site 27

Conclusion • Q-Site. Finder is better able to pinpoint the location of the ligand binding site than Pocket-Finder – High precision – Closely as possible to the actual binding site – Keep the predicted ligand binding site as small as possible without compromising accuracy • Given the high level of success in unbound protein sites – Do not have a ligand already bound 28
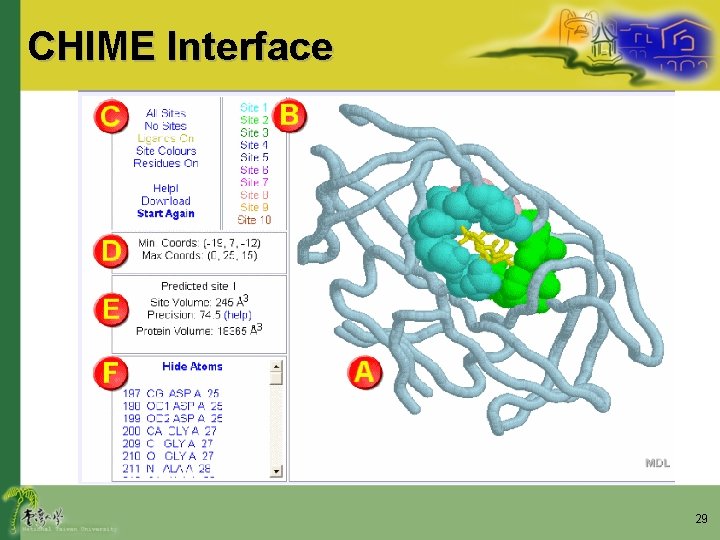
CHIME Interface 29
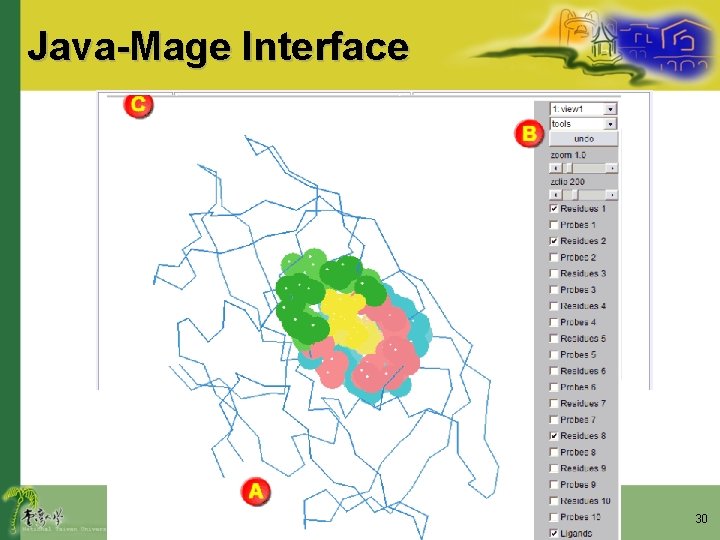
Java-Mage Interface 30

Reference • Q-Site. Finder: Ligand Binding Site Prediction – http: //bmbpcu 36. leeds. ac. uk/qsitefinder/ • Pocket-Finder: Pocket Detection – http: //bmbpcu 36. leeds. ac. uk/pocketfinder/ 31

Thanks for your attention 2020/10/7 32
- Slides: 32