QMMM Calculations and Applications to Biophysics Marcus Elstner

QM/MM Calculations and Applications to Biophysics Marcus Elstner Physical and Theoretical Chemistry, Technical University of Braunschweig
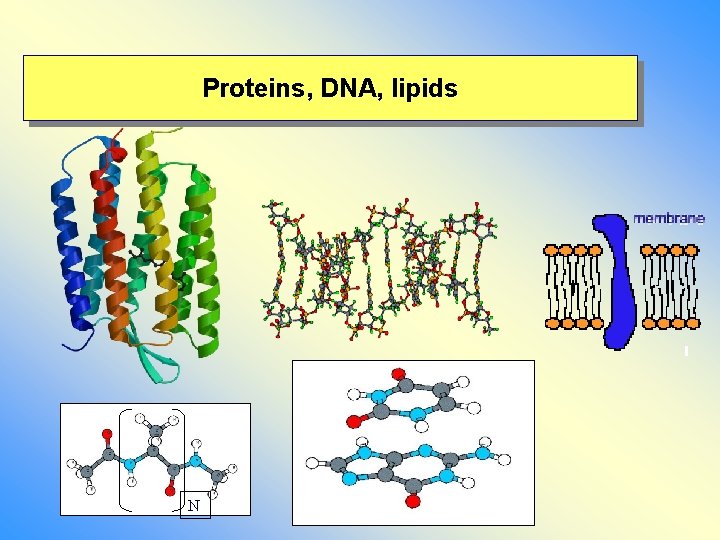
Proteins, DNA, lipids N
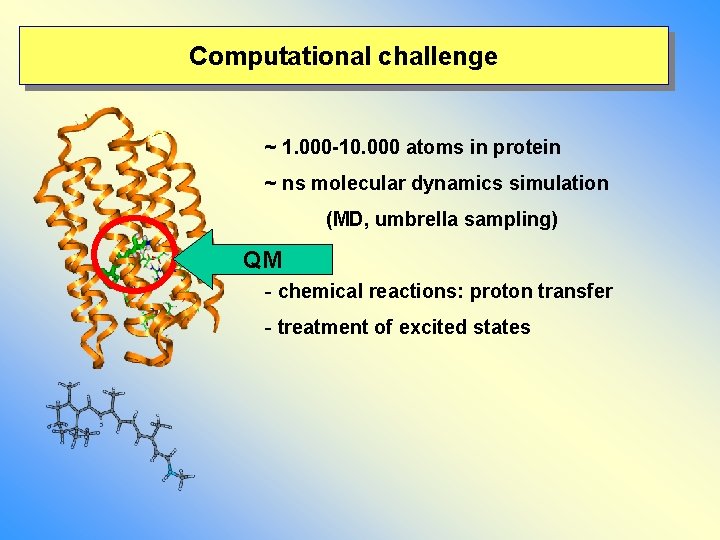
Computational challenge ~ 1. 000 -10. 000 atoms in protein ~ ns molecular dynamics simulation (MD, umbrella sampling) QM - chemical reactions: proton transfer - treatment of excited states

Computational problem I: number of atoms § chemical reaction which needs QM treatment § immediate environment: electrostatic and steric interactions § solution, membrane: polarization and structural effects on protein and reaction! 10. 000. . . - several 100. 000 atoms

Computational problem II: sampling with MD § flexibility: not one global minimum conformational entropy § solvent relaxation ps – ns timescale (timestep ~ 1 fs) (folding anyway out of reach!)
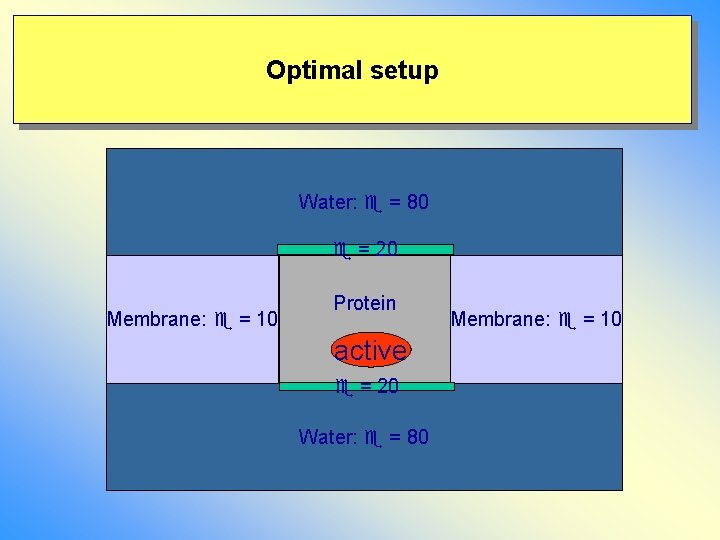
Optimal setup Water: = 80 = 20 Membrane: = 10 Protein active = 20 Water: = 80 Membrane: = 10
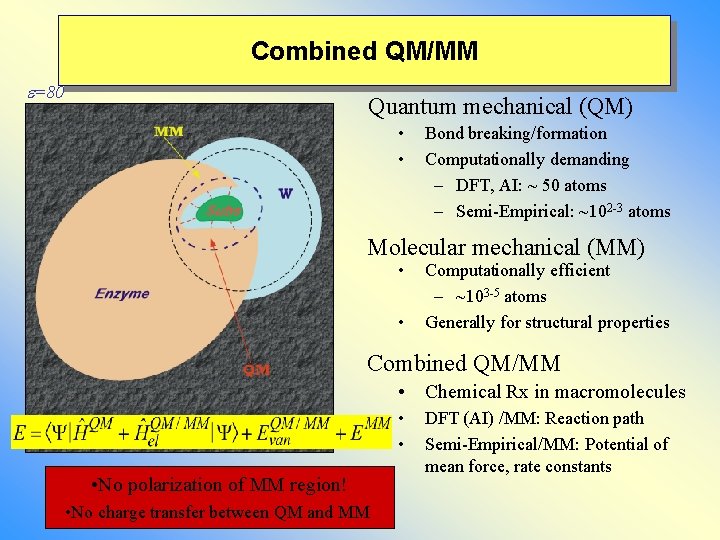
Combined QM/MM e=80 Quantum mechanical (QM) • • Bond breaking/formation Computationally demanding – DFT, AI: ~ 50 atoms – Semi-Empirical: ~102 -3 atoms Molecular mechanical (MM) • • Computationally efficient – ~103 -5 atoms Generally for structural properties Combined QM/MM • Chemical Rx in macromolecules • • • No polarization of MM region! • No charge transfer between QM and MM DFT (AI) /MM: Reaction path Semi-Empirical/MM: Potential of mean force, rate constants

Combined QM/MM 1976 Warshel and Levitt 1986 Singh and Kollman 1990 Field, Bash and Karplus QM • Semi-empirical • quantum chemistry packages: DFT, HF, MP 2, LMP 2 • DFT plane wave codes: CPMD MM • CHARMM, AMBER, GROMOS, SIGMA, TINKER, . . .
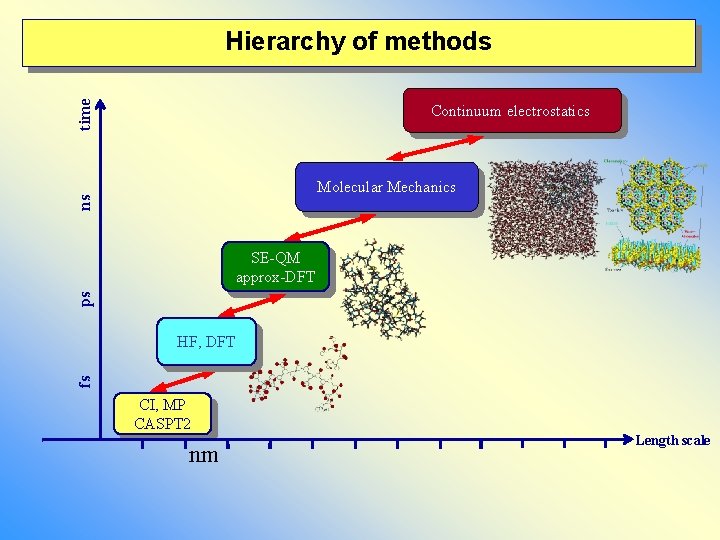
time Hierarchy of methods Continuum electrostatics ns Molecular Mechanics ps SE-QM approx-DFT fs HF, DFT CI, MP CASPT 2 nm Length scale
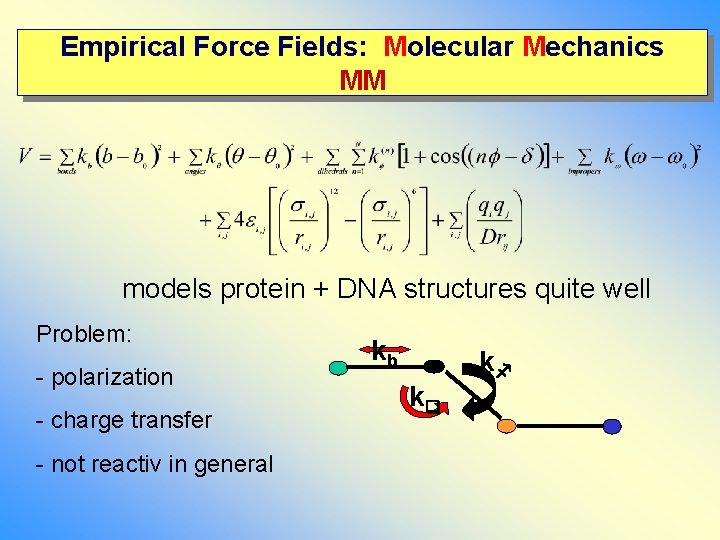
Empirical Force Fields: Molecular Mechanics MM models protein + DNA structures quite well Problem: - polarization - charge transfer - not reactiv in general kb k k

QM/MM Methods § Mechanical embedding: only steric effects § Electrostatic embedding: polarization of QM due to MM § Electrostatic embedding + polarizable MM § Larger environment: - box + Ewald summ. - continuum electrostatics - coarse graining ? ? QM MM

Ho to study reactions and (rare) dynamical events § direct MD § accelerated MD - hyperdynamics (Voter) - chemical flooding (Grubmüller) - metadynamics (Parinello) § reaction path methods - NEB (nudged elastic band, Jonsson) - CPR (conjugate peak refinement, Fischer, Karplus) - dimer method (Jonsson) § free energy sampling techniques - umbrella sampling - free energy perturbation - transition path sampling
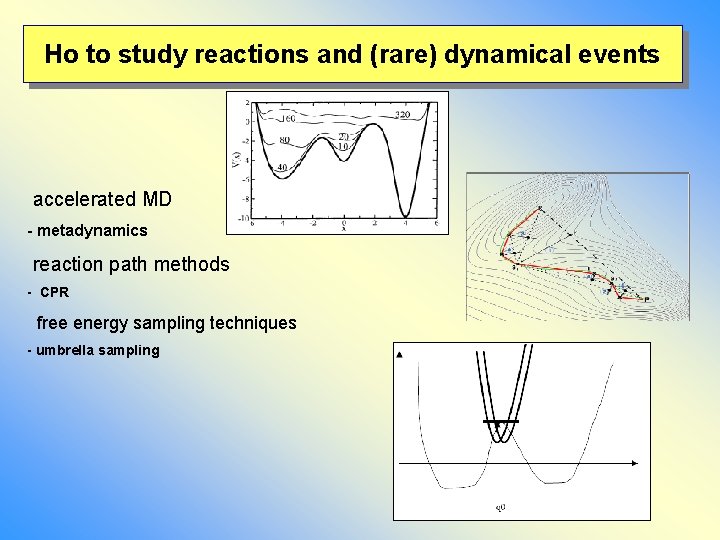
Ho to study reactions and (rare) dynamical events accelerated MD - metadynamics reaction path methods - CPR free energy sampling techniques - umbrella sampling

QM/MM Methods

Subtractive vs. additive models - subtractive: several layers: QM-MM doublecounting on the regions is subtracted - additive: different methods in different regions + interaction between the regions QM MM
Additive QM/MM total energy = + QM MM + interaction QM MM
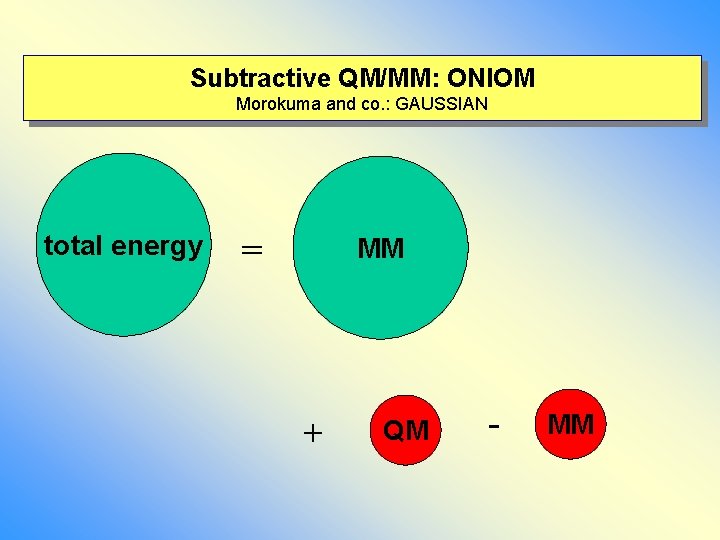
Subtractive QM/MM: ONIOM Morokuma and co. : GAUSSIAN total energy = MM + QM - MM

The ONIOM Method (an ONION-like method) Example: The binding energy of f 3 C-C f 3 (HPE) from S. Irle
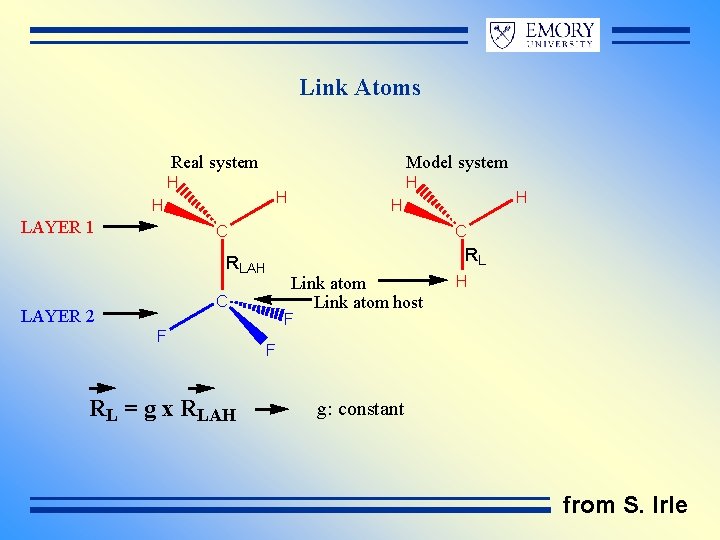
Link Atoms Real system Model system H H LAYER 1 C C RLAH C LAYER 2 F RL = g x RLAH H H RL Link atom host H F F g: constant from S. Irle
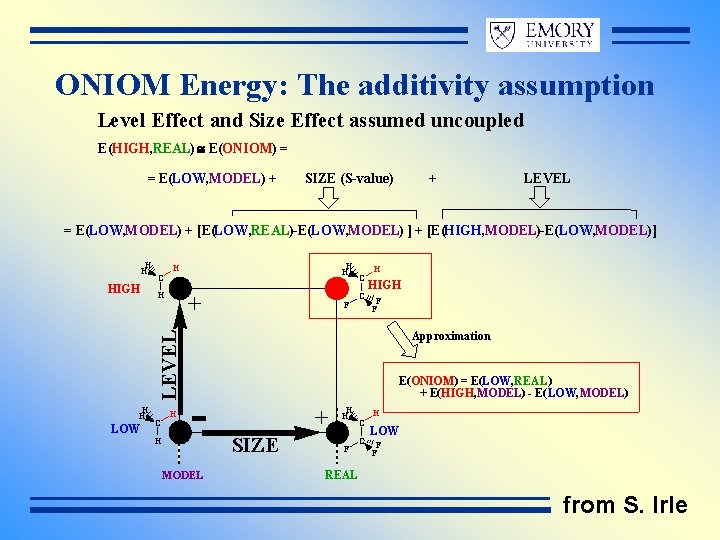
ONIOM Energy: The additivity assumption Level Effect and Size Effect assumed uncoupled E(HIGH, REAL) @ E(ONIOM) = = E(LOW, MODEL) + SIZE (S-value) + LEVEL = E(LOW, MODEL) + [E(LOW, REAL)-E(LOW, MODEL) ] + [E(HIGH, MODEL)-E(LOW, MODEL)] H H + H LOW C H H LEVEL HIGH C H F C C H HIGH F F Approximation - MODEL E(ONIOM) = E(LOW, REAL) + E(HIGH, MODEL) - E(LOW, MODEL) + SIZE H H F C C H LOW F F REAL from S. Irle
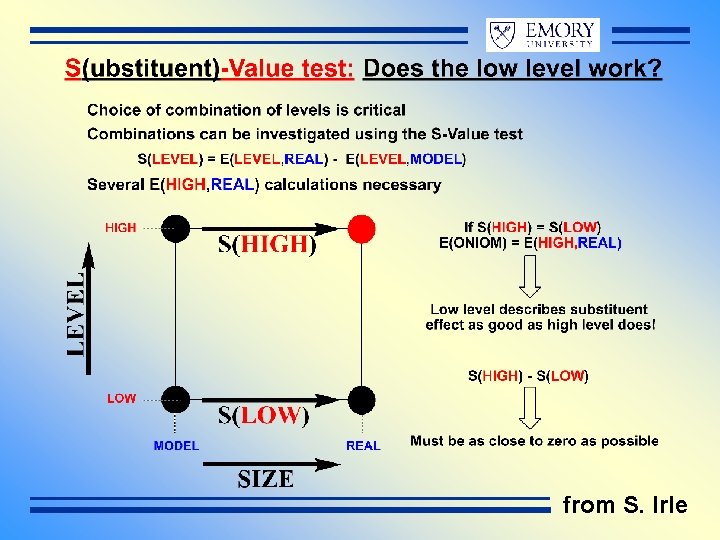
from S. Irle

ONIOM Potential Energy Surface and Properties ONIOM energy E(ONIOM, Real) = E(Low, Real) + E(High, Model) - E(Low, Model) Potential energy surface well defined, and also derivatives are available. ONIOM gradient G(ONIOM, Real) = G(Low, Real) + G(High, Model) x J - E(Low, Model) x J J = (Real coord. )/ (Model coord. ) is the Jacobian that converts the model system coordinate to the real system coordinate ONIOM Hessian H(ONIOM, Real) = H(Low, Real) + JT x H(High, Model) x J - JT x H(Low, Model) x J Scale each Hessian by s(Low)�**2 or s(High)**2 to get scaled H(ONIOM density r(ONIOM, Real) = r(Low, Real) + r(High, Model) - r(Low, Model) ONIOM properties < o (ONIOM, Real)> = < o (Low, Real) > + < o (High, Model) > - < o (Low, Model) > from S. Irle
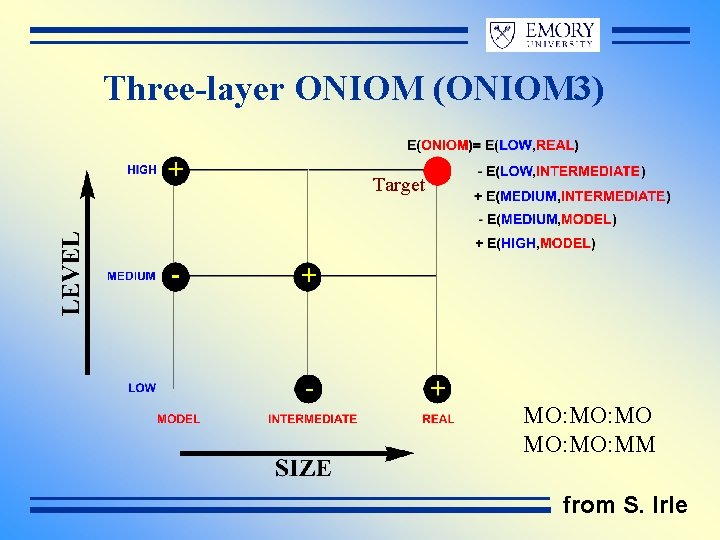
Three-layer ONIOM (ONIOM 3) Target MO: MO MO: MM from S. Irle

Additive QM/MM: linking
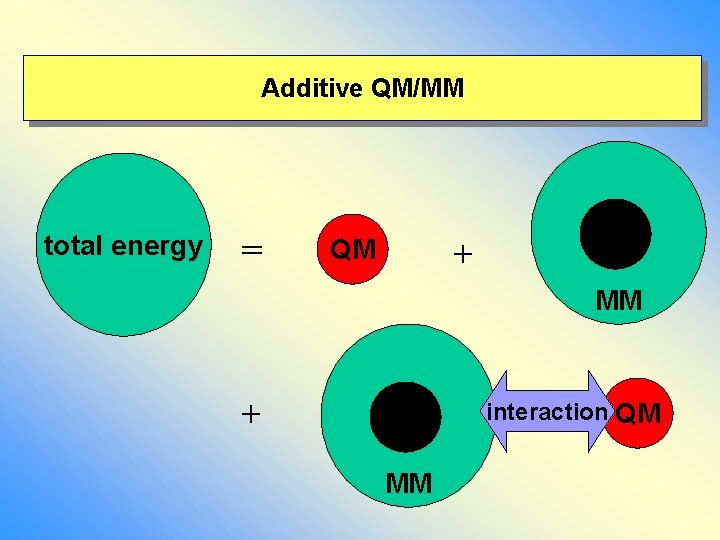
Additive QM/MM total energy = + QM MM + interaction QM MM
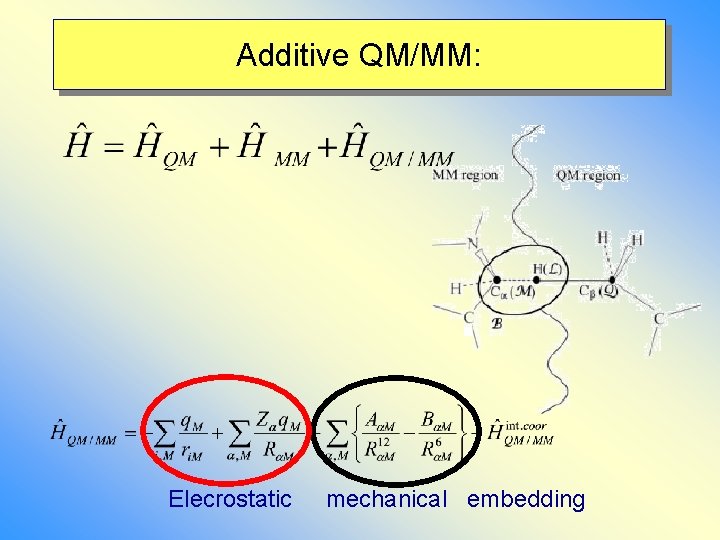
Additive QM/MM: Elecrostatic mechanical embedding
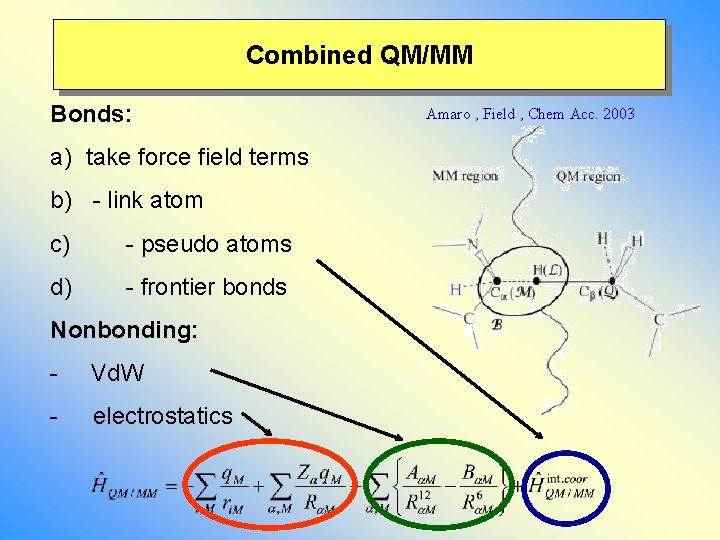
Combined QM/MM Bonds: a) take force field terms b) - link atom c) - pseudo atoms d) - frontier bonds Nonbonding: - Vd. W - electrostatics Amaro , Field , Chem Acc. 2003
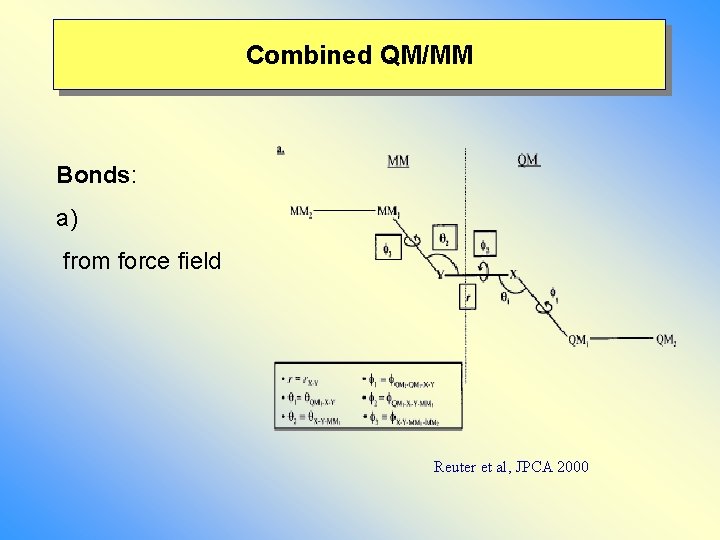
Combined QM/MM Bonds: a) from force field Reuter et al, JPCA 2000

Combined QM/MM: link atom a) constrain or not? b) (artificial forces) c) relevant for MD d) b) Electrostatics - LA included – excluded (include!) - QM-MM: exclude MM-hostgroup - Amaro & Field , T. Chem Acc. 2003 DFT, HF: gaussian broadening of MM point charges, pseudopotentails (e spill out)
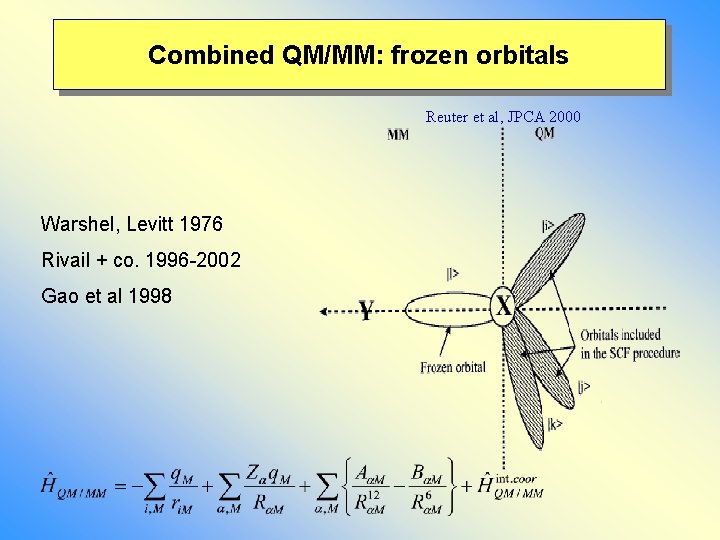
Combined QM/MM: frozen orbitals Reuter et al, JPCA 2000 Warshel, Levitt 1976 Rivail + co. 1996 -2002 Gao et al 1998
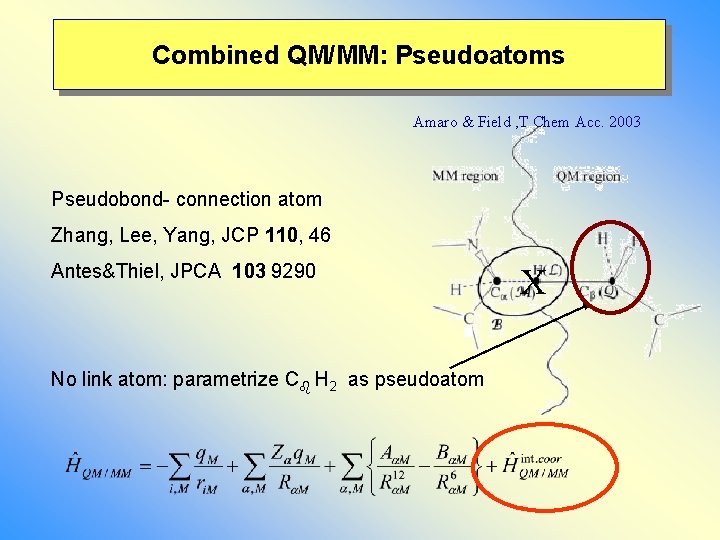
Combined QM/MM: Pseudoatoms Amaro & Field , T Chem Acc. 2003 Pseudobond- connection atom Zhang, Lee, Yang, JCP 110, 46 Antes&Thiel, JPCA 103 9290 No link atom: parametrize C H 2 as pseudoatom X
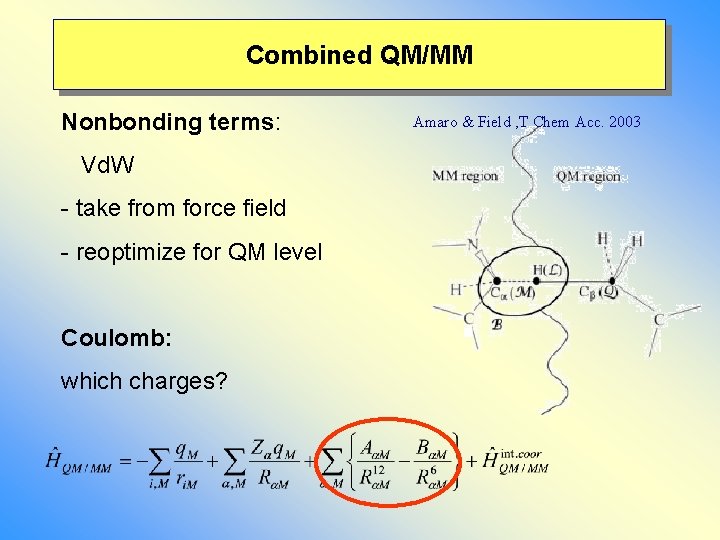
Combined QM/MM Nonbonding terms: Vd. W - take from force field - reoptimize for QM level Coulomb: which charges? Amaro & Field , T Chem Acc. 2003

Combined QM/MM Tests: - C-C bond lengths, vib. frequencies - C-C torsional barrier - H-bonding complexes - proton affinities, deprotonation energies

Subtractive vs. additive QM/MM - parametrization of methods for all regions required e. g. MM for Ligands SE for metals + QM/QM/MM conceptionally simple and applicable

Local Orbital vs. plane wave approaches: PW implementations (most implementations in LCAO) - periodic boundary conditions and large box! lots of empty space in unit cell - hybride functionals have better accuracy: B 3 LYP, PBE 0 etc. + no BSSE + parallelization (e. g. DNA with ~1000 Atoms)

Problems • QM and MM accuracy • QM/MM coupling • model setup: solvent, restraints • PES vs. FES: importance of sampling All these factors CAN introduce errors in similar magnitude

Modelling Stratgies
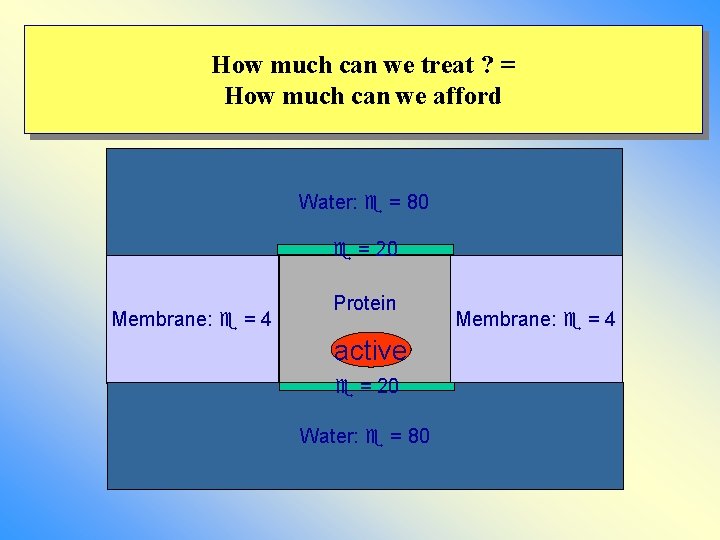
How much can we treat ? = How much can we afford Water: = 80 = 20 Membrane: = 4 Protein active = 20 Water: = 80 Membrane: = 4

How to model the environment 1) Only QM (implicit solvent) 2) QM/MM w/wo MM polarization 3) Truncated systems and charge scaling 4) System in water with periodic boundary conditions: pbc and Ewald summation 5) Truncated system and implicit solvent models

How much can we treat ? = How much can we afford Don‘t have or don‘t trust QM/MM or too complicated Only active site models active = ? ?

How much can we treat ? = How much can we afford Small protein Simple QM/MM: - fix most of the protein - neglect polarization of environment Protein active
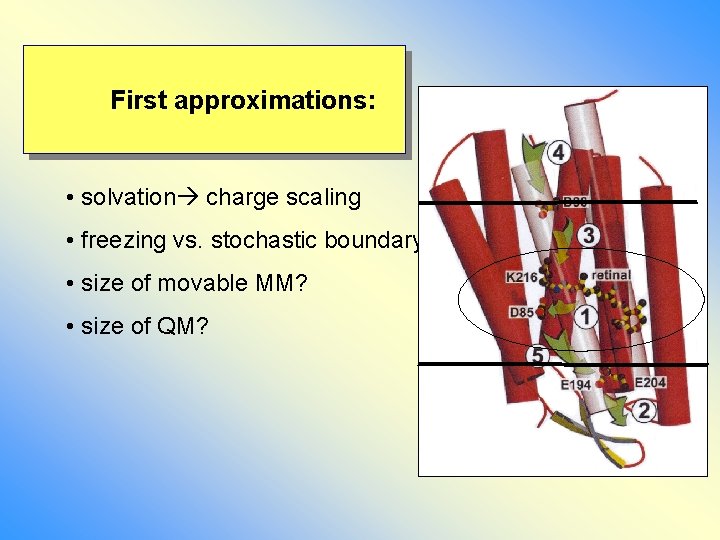
First approximations: • solvation charge scaling • freezing vs. stochastic boundary • size of movable MM? • size of QM?

How much can we treat ? = How much can we afford Small protein Simple QM/MM: - fix most of the protein - include polarization from environment Protein: polarizable active

Absolute excitation energies S 1 excitation energy (e. V) exp vacuum b. R (QM: RET) 2. 18 1 Vreven[2003] 2 TDB 3 LYP 1 TDDFTB OM 2/ CIS CASSCF 2 OM 2/ MRCI SORCI 2. 42 2. 14 2. 34 2. 86 2. 13 1. 86 2. 53 2. 21 2. 54 3. 94 2. 53 2. 34 0. 2 1. 0 Hayashi[2000] 0. 1 • TDDFT nearly zero • CIS shifts still too small ~50% • SORCI, CASPT 2 • OM 2/MRCI compares very well 0. 5

Polarizable force field for environment • MM charges • MM polarization RESP charges for residues in gas phase atomic polarizabilities: = E Polarization red shift of about 0. 1 e. V:
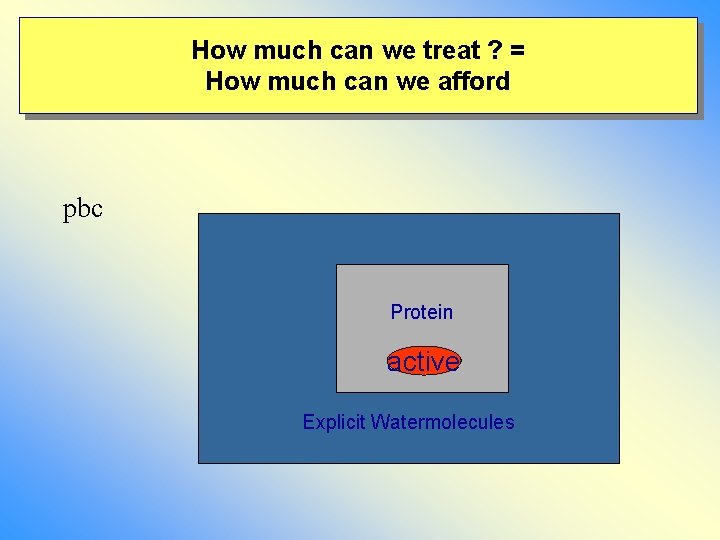
How much can we treat ? = How much can we afford pbc Protein active Explicit Watermolecules
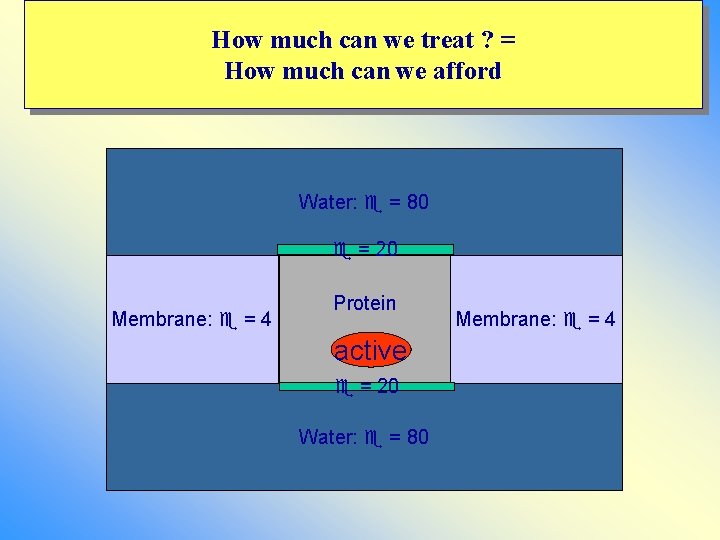
How much can we treat ? = How much can we afford Water: = 80 = 20 Membrane: = 4 Protein active = 20 Water: = 80 Membrane: = 4

Ion channels Water: = 80 Explicit water Membrane: = 4 Explicit water Water: = 80
Implicit solvent: Generalized Solvent Boundary Potential (GSBP, B. Roux) • Drawback of conventional implicit solvation: e. g. specific water molecules important • Compromise: 2 layers, one explicit solvent layer before implicit solvation model. • inner region: MD, geomopt • outer region: fixed QM/MM explicit MM implicit

GSBP Solvation free energy of point charges
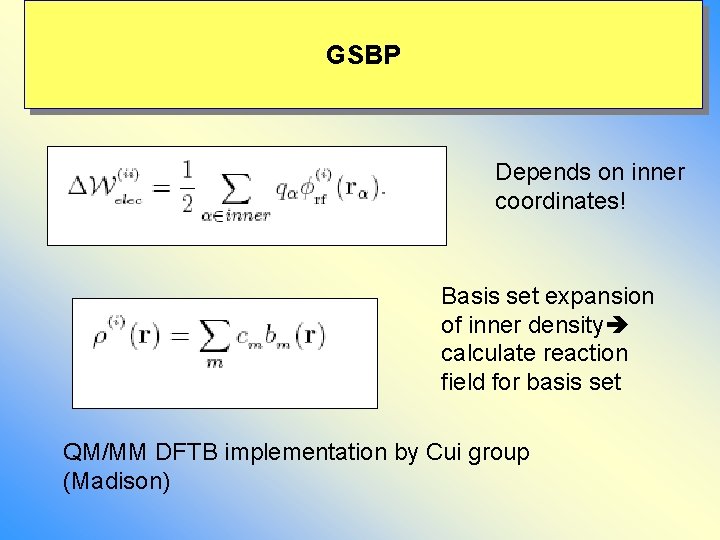
GSBP Depends on inner coordinates! Basis set expansion of inner density calculate reaction field for basis set QM/MM DFTB implementation by Cui group (Madison)
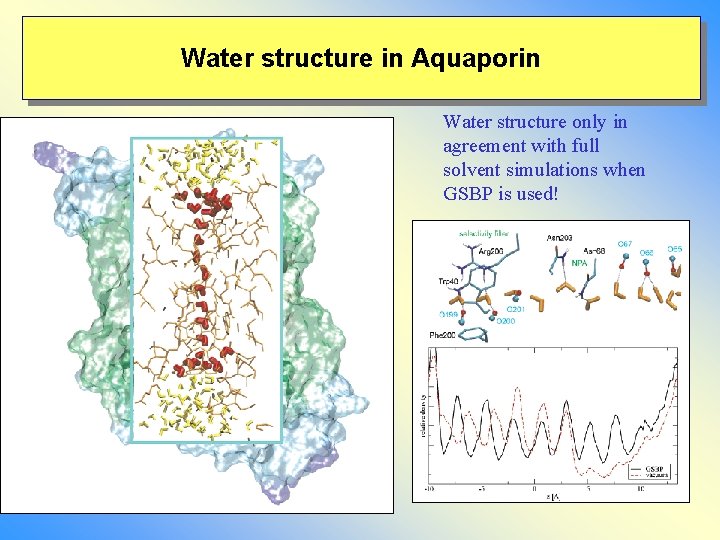
Water structure in Aquaporin Water structure only in agreement with full solvent simulations when GSBP is used!
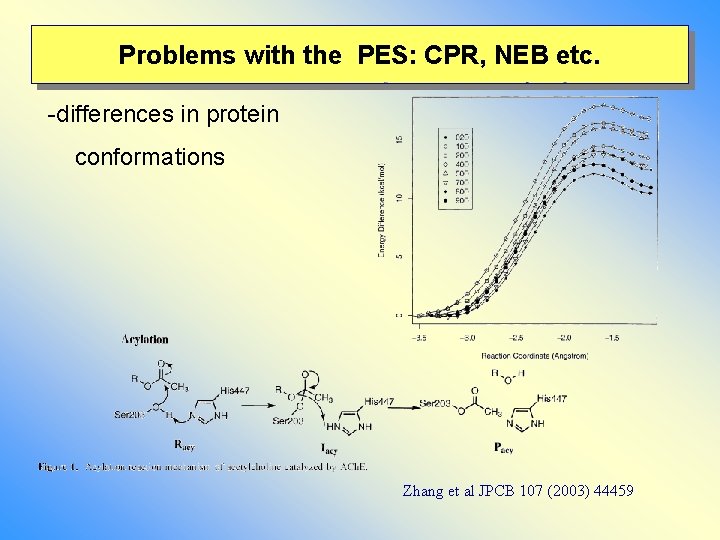
Problems with the PES: CPR, NEB etc. -differences in protein conformations Zhang et al JPCB 107 (2003) 44459

Problems with the PES: complex energy landscape - differences in protein conformations (starting the reaction path calculation) - problems along the reaction pathway * flipping of water molecules * size of movable MM region different H-bonding pattern average over these effects: potential of mean force/free energy
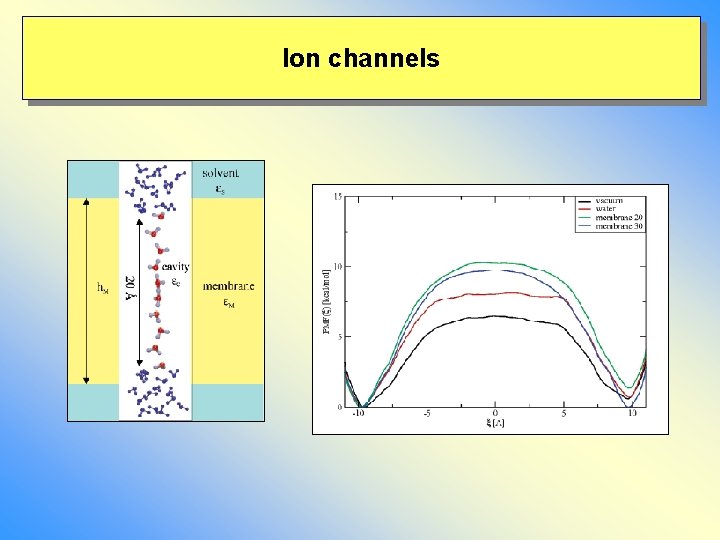
Ion channels
- Slides: 55