QIBA CT Volumetrics CrossPlatform Study Group 1 C

QIBA CT Volumetrics Cross-Platform Study (Group 1 C) March 29, 2009 Interclinic Comparison of CT Volumetry Quantitative Imaging Biomarker Alliance

Charge 1. 2. 3. 4. 5. To agree on the scanner settings and other protocol elements under which imagery is to be collected. To agree on requirements of phantoms to be imaged/measured. To agree on the platforms and centers to be selected for imagery collected. To identify the measurements and the algorithms for use in image processing. To specify the analysis of the measurements.

Goals 1. Measure the volume of nodules on CT imagery collected from several CT scanners and sites (may include multiple settings on single scanners). 1. Measure image noise and other image quality factors and determine their impact on the measurement of volume. 2. Compare the accuracy and precision of volume measurements for these phantom datasets. 3. Determine the minimum detectable level of change that can be achieved when measuring nodules in phantom datasets.

Goal 1 1. Measure nodule volume on CT imagery collected from several CT scanners/sites (including single scanners with varying settings). Determine the systems to be used and the system settings to be varied. (a) (b) (c) (d) (e) k. Vp may be specified. m. As may be specified. collimation fixed (+) field of view (skin-to-skin = closest possible view)reconstruction filters – follow-up Wendy & radiologists - Find “equivalent” filters. Site selection – poll the team for potential image collection sites.

Goal 2 2. Measure “image noise” and determine its impact on the measurement of volume. Facilitates inter-comparison of scanner results. (a) Characterizing / specifying image quality (a) Expert visual assessment of image quality. (Deni)

QIBA CT 1 -C Protocol - first branch Using protocol for NLST (ACRIN 6678? ) specify k. Vp, slice thickness, m. As, rotation time, pitch, reconstruction kernel (affects MTF). Use (water and ACR) phantoms to characterize the resolution and noise levels under this branch of the protocol.
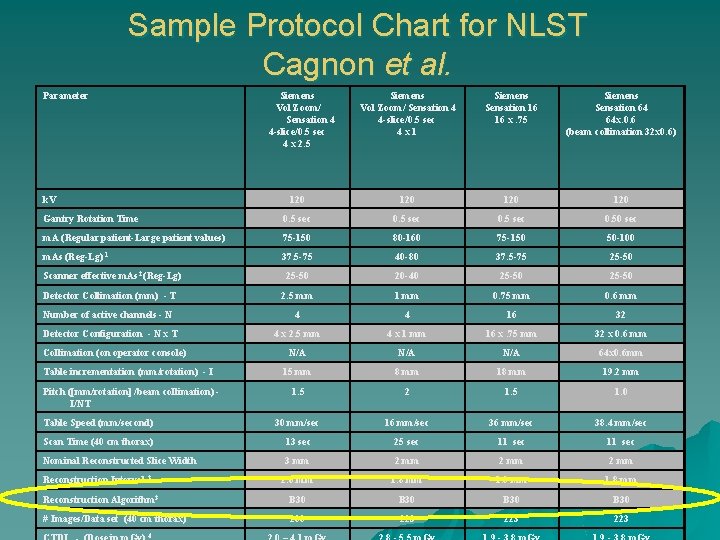
Sample Protocol Chart for NLST Cagnon et al. Parameter Siemens Vol Zoom/ Sensation 4 4 -slice/0. 5 sec 4 x 2. 5 Siemens Vol Zoom/ Sensation 4 4 -slice/0. 5 sec 4 x 1 Siemens Sensation 16 16 x. 75 Siemens Sensation 64 64 x. 0. 6 (beam collimation 32 x 0. 6) 120 120 Gantry Rotation Time 0. 5 sec 0. 50 sec m. A (Regular patient-Large patient values) 75 -150 80 -160 75 -150 50 -100 m. As (Reg-Lg) 1 37. 5 -75 40 -80 37. 5 -75 25 -50 Scanner effective m. As 2 (Reg-Lg) 25 -50 20 -40 25 -50 Detector Collimation (mm) - T 2. 5 mm 1 mm 0. 75 mm 0. 6 mm Number of active channels - N 4 4 16 32 Detector Configuration - N x T 4 x 2. 5 mm 4 x 1 mm 16 x. 75 mm 32 x 0. 6 mm N/A N/A 64 x 0. 6 mm 15 mm 8 mm 19. 2 mm 1. 5 2 1. 5 1. 0 Table Speed (mm/second) 30 mm/sec 16 mm/sec 38. 4 mm/sec Scan Time (40 cm thorax) 13 sec 25 sec 11 sec Nominal Reconstructed Slice Width 3 mm 2. 0 mm 1. 8 mm Reconstruction Algorithm 3 B 30 # Images/Data set (40 cm thorax) 200 223 223 k. V Collimation (on operator console) Table incrementation (mm/rotation) - I Pitch ([mm/rotation] /beam collimation) I/NT Reconstruction Interval 3 4

QIBA CT 1 -C Profiles - second branch Specify PERFORMANCE metrics such as simple spatial resolution and noise metrics. u k. Vp (affects contrast difference between materials) u Slice thickness, recon interval (affects z-axis resolution & noise) u Rotation time and pitch (coverage, breath hold, etc. ) u Recon kernel OR recon kernel performance - EXAMPLE: – Choose kernel such that you can see 6 or 7 (but no more than 7) lp/cm on ACR phantom…. or – 10% MTF should be between 6 and 7 lp/cm u m. A level performance – Choose effective m. As level so that std dev is between 20 and 30 HU in a 20 cm water phantom

Goal 3 Compare the accuracy and precision of radiologists’ measurements of RECIST and Volume for these phantom datasets. (image mask? ) a) b) c) d) RECIST vs. volume. Investigate variance & bias. Inter-system variation. Intra-system variation.

Goal 4 4. DEFER: Determine the minimum detectable level of change that can be achieved when measuring nodules in phantom datasets.

Required resources u u u Use of FDA phantom, water and ACR phantoms. Recruit clinical image collection sites through QIBA-CT group. Use QIBA CT 1 -A mark-up at Rad. Pharm, generating u u RECIST Segmented volume.

Future steps Refine Questions and Experimental Design. u Recruit participating clinics. Share and discuss plans with associated medical physicists. u Confirm availability and discuss reading with Rad-Pharm. u Confirm availability and schedule FDA phantom. u
- Slides: 12