Psalm 102 25 25 In the beginning you

Psalm 102: 25 25 In the beginning you laid the foundations of the earth, and the heavens are the work of your hands. © 2001 Timothy G. Standish

Initiation of Transcription: In Eukaryotes Timothy G. Standish, Ph. D. © 2001 Timothy G. Standish

Expression Control In Eukaryotes �Some of the general methods used to control expression in prokaryotes are used in eukaryotes, but nothing resembling operons is known �Eukaryotic genes are controlled individually and each gene has specific control sequences preceding the transcription start site �In addition to controlling transcription, there additional ways in which expression can be controlled in eukaryotes © 2001 Timothy G. Standish

Eukaryotes Have Large Complex Geneomes �The human genome is about 3 x 109 base pairs or ≈ 1 m of DNA �Because humans are diploid, each nucleus contains 6 x 109 base pairs or ≈ 2 m of DNA �Some gene families are located close to one another on the same chromosome �Genes with related functions appear to be distributed almost at random throughout the genome © 2001 Timothy G. Standish

Highly Packaged DNA Cannot be Expressed �Because of its size, eukaryotic DNA must be packaged �Heterochromatin, the most highly packaged form of DNA, cannot be transcribed, therefore expression of genes is prevented �Chromosome puffs on some insect chomosomes illustrate areas of active gene expression © 2001 Timothy G. Standish

�It Only a Subset of Genes is Expressed at any Given Time takes lots of energy to express genes �Thus it would be wasteful to express all genes all the time �By differential expression of genes, cells can respond to changes in the environment �Differential expression allows cells to specialize in multicelled organisms. �Differential expression also allows organisms to develop over time. © 2001 Timothy G. Standish
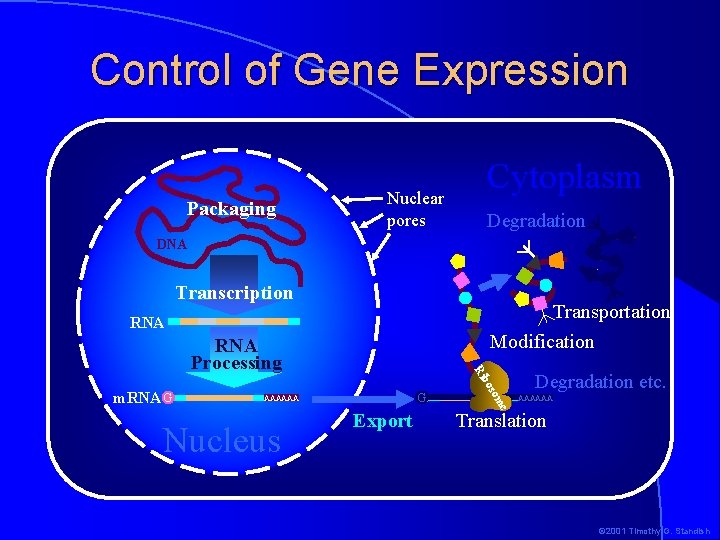
Control of Gene Expression Packaging Nuclear pores Cytoplasm Degradation DNA Transcription Transportation Modification RNA G AAAAAA Degradation etc. AAAAAA e Nucleus Export som bo m. RNA G Ri RNA Processing Translation © 2001 Timothy G. Standish

Logical Expression Control Points packaging � Transcription � RNA processing � m. RNA export � m. RNA masking/unmasking and/or modification � m. RNA degradation � Translation � Protein modification � Protein transport � Protein degradation Increasing cost � DNA The logical place to control expression is before the gene is transcribed © 2001 Timothy G. Standish

Three Eukaryotic RNA Polymerases 1 RNA Polymerase I - Produces r. RNA in the nucleolus, accounts for 50 - 70 % of transcription 2 RNA Polymerase II - Produces m. RNA in the nucleoplasm - 20 - 40 % of transcription 3 RNA Polymerase III - Produces t. RNA in the nucleoplasm - 10 % of transcription © 2001 Timothy G. Standish
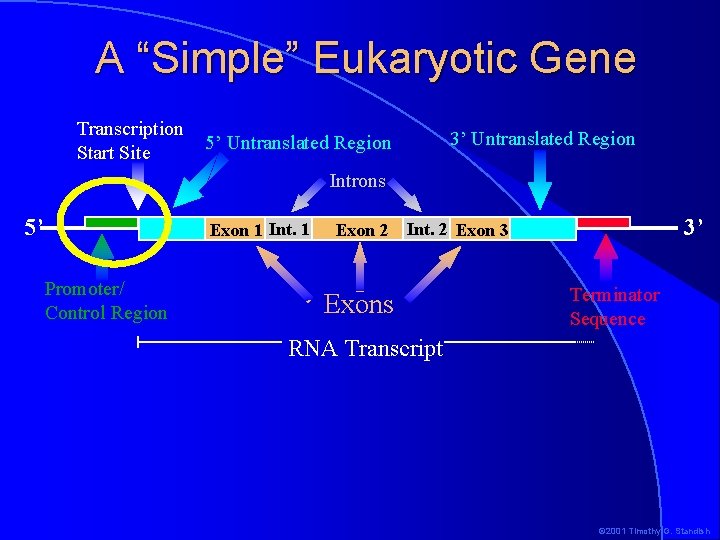
A “Simple” Eukaryotic Gene Transcription Start Site 3’ Untranslated Region 5’ Untranslated Region Introns 5’ Exon 1 Int. 1 Promoter/ Control Region Exon 2 3’ Int. 2 Exon 3 Exons Terminator Sequence RNA Transcript © 2001 Timothy G. Standish
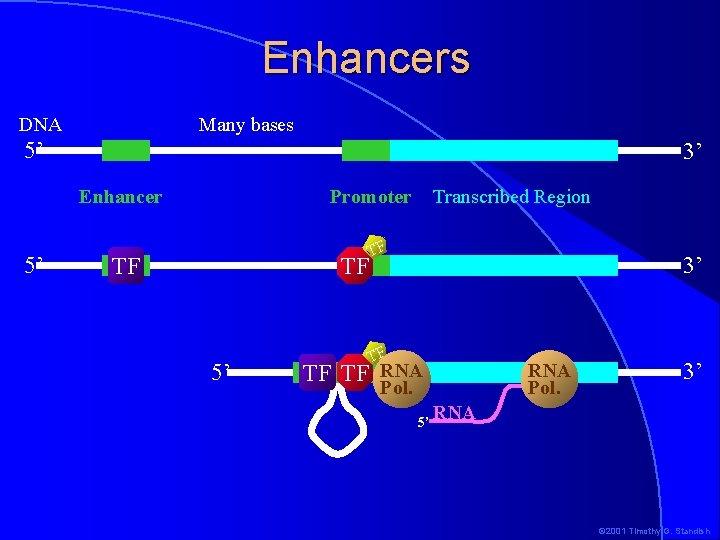
Enhancers Many bases DNA 5’ 3’ Enhancer 5’ Promoter TF TF 5’ Transcribed Region TF 3’ TF TF TF RNA Pol. 5’ 3’ RNA © 2001 Timothy G. Standish

Eukaryotic RNA Polymerase II �RNA polymerase is a very fancy enzyme that does many tasks in conjunction with other proteins �RNA polymerase II is a protein complex of over 500 k. D with more than 10 subunits: © 2001 Timothy G. Standish

Eukaryotic RNA Polymerase II Promoters �Several sequence elements spread over about 200 bp upstream from the transcription start site make up RNA Pol II promoters �Enhancers, in addition to promoters, influence the expression of genes �Eukaryotic expression control involves many more factors than control in prokaryotes �This allows much finer control of gene expression © 2001 Timothy G. Standish
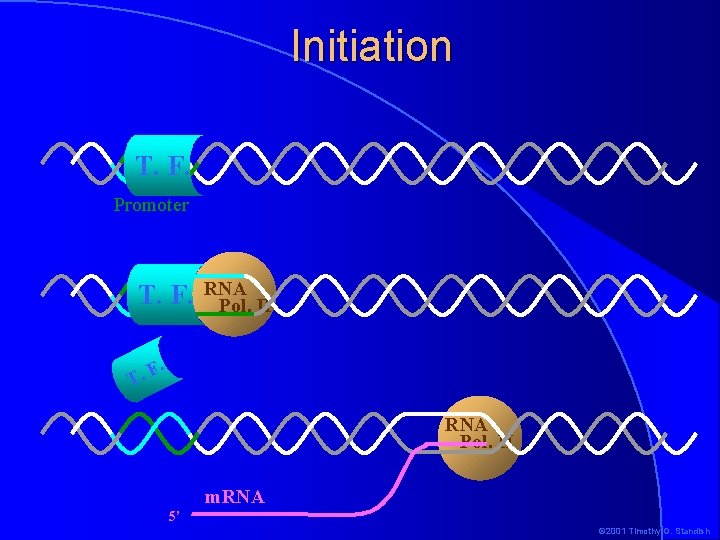
Initiation T. F. Promoter T. F. RNA Pol. II . F T. RNA Pol. II m. RNA 5’ © 2001 Timothy G. Standish

Eukaryotic Promoters Promoter 5’ Exon 1 Sequence elements TATA ~200 bp “TATA Box” Initiator Transcription start site SSTATAAAASSSSSNNNNNNNNNYYCAYY (Template strand) -1+1 ~-25 S = C or G Y = C or T N = A, T, G or C © 2001 Timothy G. Standish
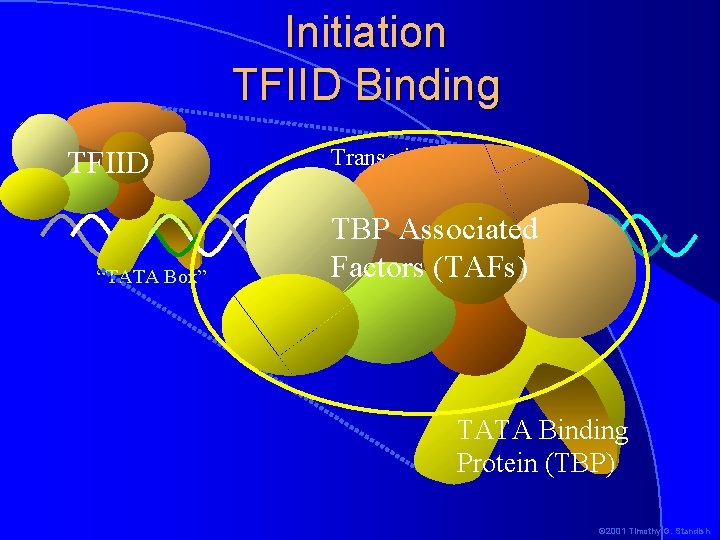
Initiation TFIID Binding TFIID “TATA Box” Transcription start site TBP Associated Factors (TAFs) -1+1 TATA Binding Protein (TBP) © 2001 Timothy G. Standish

Initiation TFIID Binding Transcription start site TFIID -1+1 80 o Bend © 2001 Timothy G. Standish
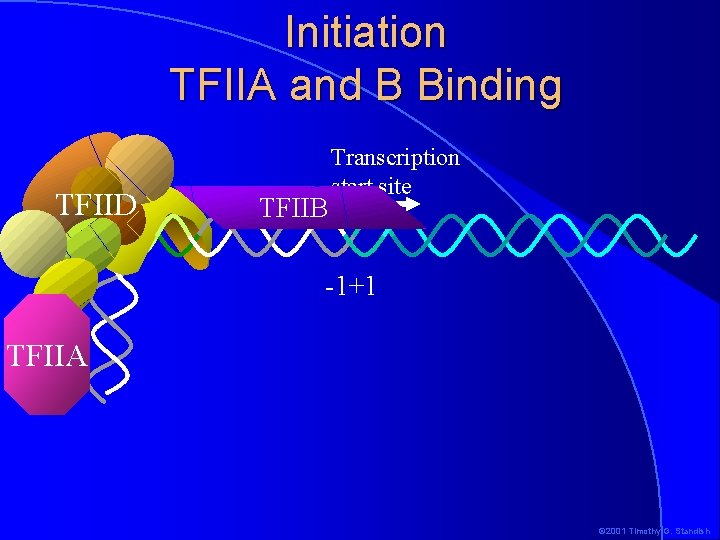
Initiation TFIIA and B Binding TFIID TFIIB Transcription start site -1+1 TFIIA © 2001 Timothy G. Standish

Initiation TFIIF and RNA Polymerase Binding TFIID TFIIB Transcription start site -1+1 TFIIA TFIIF RNA Polymerase © 2001 Timothy G. Standish
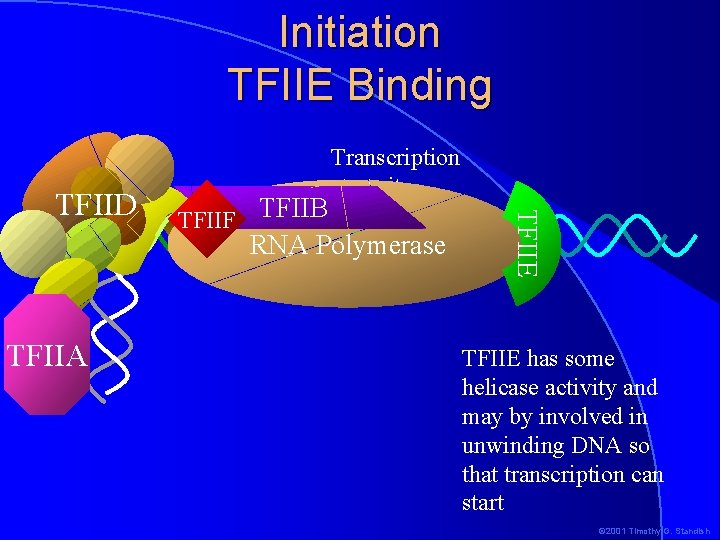
Initiation TFIIE Binding TFIIF TFIIB RNA Polymerase -1+1 TFIIA TFIIE TFIID Transcription start site TFIIE has some helicase activity and may by involved in unwinding DNA so that transcription can start © 2001 Timothy G. Standish
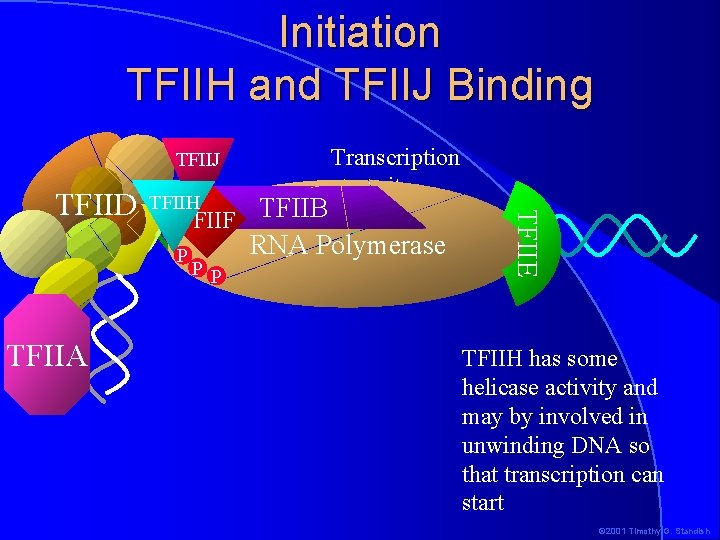
Initiation TFIIH and TFIIJ Binding TFIIJ TFIIH TFIIF TFIIB P TFIIA PP RNA Polymerase -1+1 TFIIE TFIID Transcription start site TFIIH has some helicase activity and may by involved in unwinding DNA so that transcription can start © 2001 Timothy G. Standish
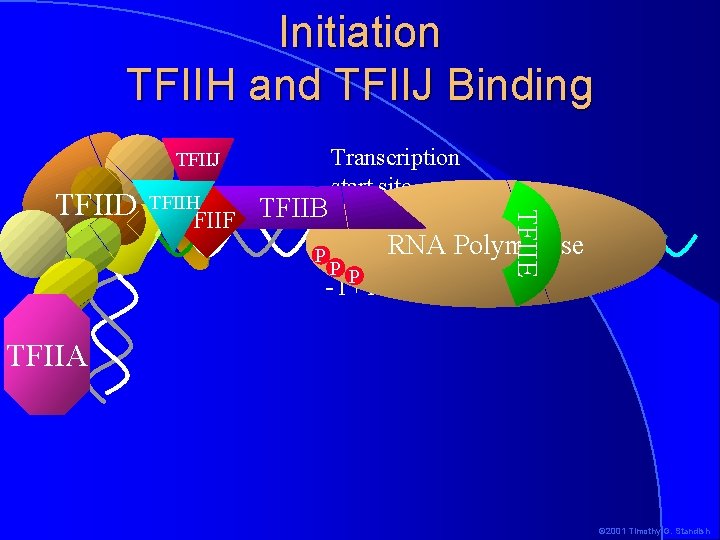
Initiation TFIIH and TFIIJ Binding TFIIJ TFIIH TFIIF TFIIB P PP -1+1 TFIIE TFIID Transcription start site RNA Polymerase TFIIA © 2001 Timothy G. Standish

Initiation TFIIH and TFIIJ Binding Transcription start site P -1+1 PP RNA Polymerase © 2001 Timothy G. Standish

© 2001 Timothy G. Standish
- Slides: 24