Proteus a Grid based Problem Solving Environment PSE

Proteus, a Grid based Problem Solving Environment (PSE) for Bioinformatics: Architecture and Experiments Presenter: Michael Robinson Agnostic: Javier Munoz Advanced Topics in Software Engineering CIS 6612 Florida International University July 31, 2006 Authors: Mario Cannataro 1, Carmela Comito 2, Filippo Lo Schiavo 1, and Pierangelo Veltri 1 (February 2004) 1 University of Magna Graecia of Catanzaro, Italy 2 University of Calabria, Italy

Organization n n n Abstract ~60% is about Bioinformatics Proteus Architecture First Test Implementation Results of First Test Conclusion and Future Work 2

Abstract n Live sciences Bioinformatics Computer Science n Data Files sizes n Computer power 3

The Partners n What is Livesciences n What is Bioinformatics n n Other Sciences used in Bioinformatics What is Computer Science 4

Human Genome n n The sum total of DNA in an organism is its genome. The Human Genome Project (HGP) an international effort, began in October 1990, and was completed in 1999, 2003, 2004. (http: //www. pbs. org/wgbh/nova/genome/program. html) n Project goals were to: n Determine the complete sequence of the 3 billion DNA bases n Identify all human genes n And make them accessible for further biological study 5

Human Genome n The bacterium E. coli and others were used to help develop the technology and interpret human gene function. The Human Genome Project was sponsored by: The U. S. Department of Energy and The U. S. National Institutes of Health n http: //www. preventiongenetics. com/edu/genetics_nutshell. htm 6

DNA (ACGT) n Humans have from 10 to 100 trillion cells n Each Human cell has about 3 billion nucleotides n We have approximately 30, 000 genes n Of the three billion letters of DNA that we have, only 1 to 1. 5 percent of it is gene the rest is STUFF”. n The functions are unknown for over 50% of known genes 7

DNA (ACGT) Human Genome n n n 3, 000, 000 ~ dna bases 30, 000 ~ bases in genes 2, 970, 000 ~ stuff adenine (A) forms a base pair with thymine (T) guanine (G) forms a base pair with cytosine (C) n 8

Similarities to Human DNA Another n human? 99. 9% - All humans have the same genes, but some of these genes contain sequence differences that make each person unique. A chimpanzee? 98. 5% - Chimpanzees are the closest living species to humans. A mouse? 92. 0% - All mammals are quite similar genetically. A fruit fly? 44. 0% - Studies of fruit flies have shown how shared genes govern the growth and structure of both insects and mammals. Yeast? 26. 0% - Yeasts are single-celled organisms, but they have many housekeeping genes that are the same as the genes in humans, such as those that enable energy to be derived from the breakdown of sugars. A weed (thale cress)? 18. 0% - Plants have many metabolic differences from humans. For example, they use sunlight to convert carbon dioxide gas to sugars. But they also have similarities in their housekeeping genes. 9

The gene sizes n n Largest known human gene is dystrophin at 2. 4 million bases. Chromosome 21 is the smallest human chromosome. Three copies of this autosome causes Down syndrome, the most frequent genetic disorder associated with significant mental retardation. Academic groups from Germany and Japan mapped and sequenced it, it has 33, 546, 361 bp of DNA Analysis of the chromosome revealed: n 127 known genes, n 98 predicted genes, n and 59 pseudogenes. n Smallest bacterial genome, Mycoplasma genitalium size of 580 kbp 10
Bioinformatics n DNA RNA PROTEINS MUTATIONS, ILLNESSES MEDICATIONS CLONING 11

DNA (ACGT) n n Pseudomonas Aeruginosas PA 01 6, 264, 403 bases, 5565 genes complement(6264226. . 6264360) 6264181 gcttgtcccg gtcgaagtcc cgactcacca cccgtaccgg ataaatcaga cggtcagacg 6264241 cttacggcct ttggcgcgac gacgcgacag aacctgacgg ccgttcttgg tggccatacg 6264301 ggcgcggaaa ccgtggacgc gagcgcgctt gagggtgctg ggttggaaag tacgtttcat 6264361 gattcggtac ctgggttgac gacttgaggt cgcagtgacc ccg 12

RNA n In RNA, thymine is replaced by uracil (U). DNA 6264181 gcttgtcccg gtcgaagtcc cgactcacca cccgtaccgg ataaatcaga cggtcagacg 6264241 cttacggcct ttggcgcgac gacgcgacag aacctgacgg ccgttcttgg tggccatacg 6264301 ggcgcggaaa ccgtggacgc gagcgcgctt gagggtgctg ggttggaaag tacgtttcat 6264361 gattcggtac ctgggttgac gacttgaggt cgcagtgacc ccg RNA 6264181 gcuugucccg gucgaagucc cgacucacca cccguaccgg auaaaucaga cggucagacg 6264241 cuuacggccu uuggcgcgac gacgcgacag aaccugacgg ccguucuugg uggccauacg 6264301 ggcgcggaaa ccguggacgc gagcgcgcuu gagggugcug gguuggaaag uacguuucau 6264361 gauucgguac cuggguugac gacuugaggu cgcagugacc ccg 13
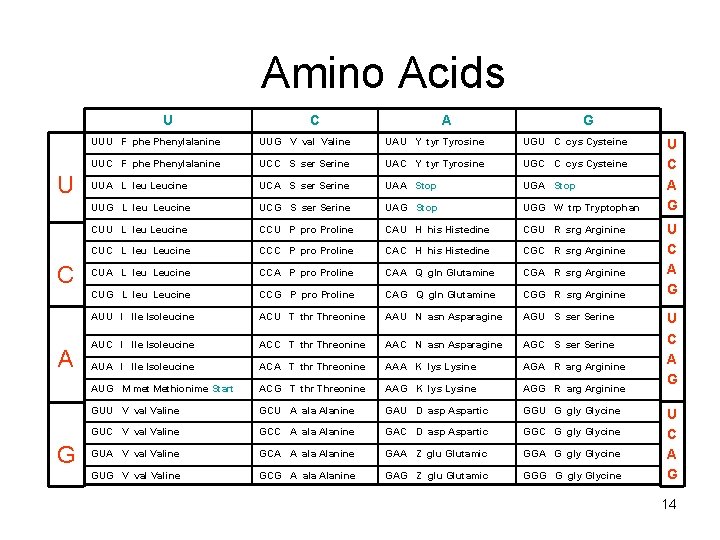
Amino Acids U U C A G UUU F phe Phenylalanine UUG V val Valine UAU Y tyr Tyrosine UGU C cys Cysteine UUC F phe Phenylalanine UCC S ser Serine UAC Y tyr Tyrosine UGC C cys Cysteine UUA L leu Leucine UCA S ser Serine UAA Stop UGA Stop UUG L leu Leucine UCG S ser Serine UAG Stop UGG W trp Tryptophan CUU L leu Leucine CCU P pro Proline CAU H his Histedine CGU R srg Arginine CUC L leu Leucine CCC P pro Proline CAC H his Histedine CGC R srg Arginine CUA L leu Leucine CCA P pro Proline CAA Q gln Glutamine CGA R srg Arginine CUG L leu Leucine CCG P pro Proline CAG Q gln Glutamine CGG R srg Arginine AUU l lle Isoleucine ACU T thr Threonine AAU N asn Asparagine AGU S ser Serine AUC l lle Isoleucine ACC T thr Threonine AAC N asn Asparagine AGC S ser Serine AUA l lle Isoleucine ACA T thr Threonine AAA K lys Lysine AGA R arg Arginine AUG M met Methionime Start ACG T thr Threonine AAG K lys Lysine AGG R arg Arginine GUU V val Valine GCU A ala Alanine GAU D asp Aspartic GGU G gly Glycine GUC V val Valine GCC A ala Alanine GAC D asp Aspartic GGC G gly Glycine GUA V val Valine GCA A ala Alanine GAA Z glu Glutamic GGA G gly Glycine GUG V val Valine GCG A ala Alanine GAG Z glu Glutamic GGG G gly Glycine U C A G 14

Proteins (sequences) DNA 6264181 gcttgtcccg gtcgaagtcc cgactcacca cccgtaccgg ataaatcaga cggtcagacg 6264241 cttacggcct ttggcgcgac gacgcgacag aacctgacgg ccgttcttgg tggccatacg 6264301 ggcgcggaaa ccgtggacgc gagcgcgctt gagggtgctg ggttggaaag tacgtttcat 6264361 gattcggtac ctgggttgac gacttgaggt cgcagtgacc ccg RNA 6264181 gcuugucccg gucgaagucc cgacucacca cccguaccgg auaaaucaga cggucagacg 6264241 cuuacggccu uuggcgcgac gacgcgacag aaccugacgg ccguucuugg uggccauacg 6264301 ggcgcggaaa ccguggacgc gagcgcgcuu gagggugcug gguuggaaag uacguuucau 6264361 gauucgguac cuggguugac gacuugaggu cgcagugacc ccg PROTEIN MKRTFQPSTLKRARVHGFRARMATKNGRQVLSRRRAKGRKRLTV 15
![Proteins: Pattern Matching G-H-E-X(2)-G-X(4, 5)-[GA] 16 Proteins: Pattern Matching G-H-E-X(2)-G-X(4, 5)-[GA] 16](http://slidetodoc.com/presentation_image/02dd13b239271ed6745fe0518f00d6ff/image-16.jpg)
Proteins: Pattern Matching G-H-E-X(2)-G-X(4, 5)-[GA] 16
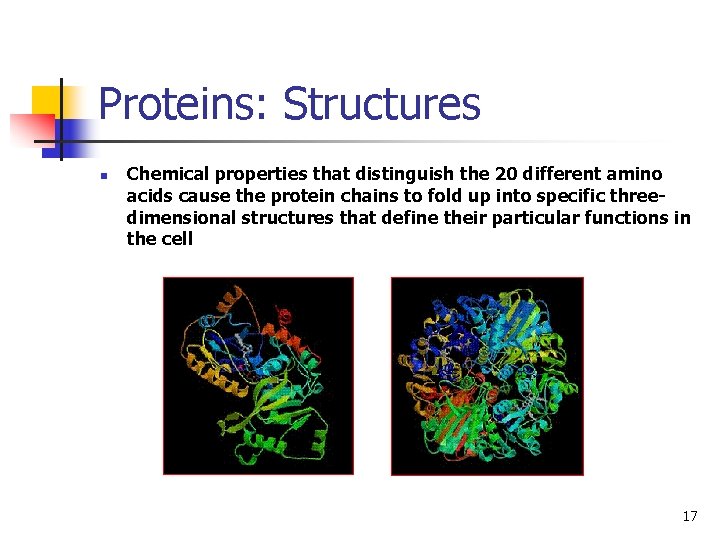
Proteins: Structures n Chemical properties that distinguish the 20 different amino acids cause the protein chains to fold up into specific threedimensional structures that define their particular functions in the cell 17

Reality Somewhere in this dense chemical forest are genes involved in deafness, Alzheimer, cancer, cataracts, etc. But where? This is such a maze scientists need a map. n n Out of three billion base pairs in our DNA, just one single letter can make a difference. 18

Data Locations Gen. Bank in the US, 1974 n 1997 = 1. 26 gigabases http: //www. ncbi. nlm. nih. gov/ 2004 = 39 gigabases 2005 = 100 gigabases EMBL in England, 1980 n http: //www. ebi. ac. uk/embl/ DDBJ in Japan, 1984 n http: //www. ddbj. nig. ac. jp/ 19

Some Databases n The Swiss Institute of Bioinformatics maintains the following databases: Ashbya Genome Database Cancer Immunome Database Eukaryotic Promoter Database (EPD) Germ. Online My. Hits PROSITE Swiss-Prot and Tr. EMBL SWISS-2 DPAGE SWISS-MODEL Repository 20

Specialization Plasmodb http: //www. plasmodb. org/plasmo/home. jsp parasitic eukaryote Plasmodium the causative agent of the disease Malaria. apibugz@delphi. pcbi. upenn. edu n 21
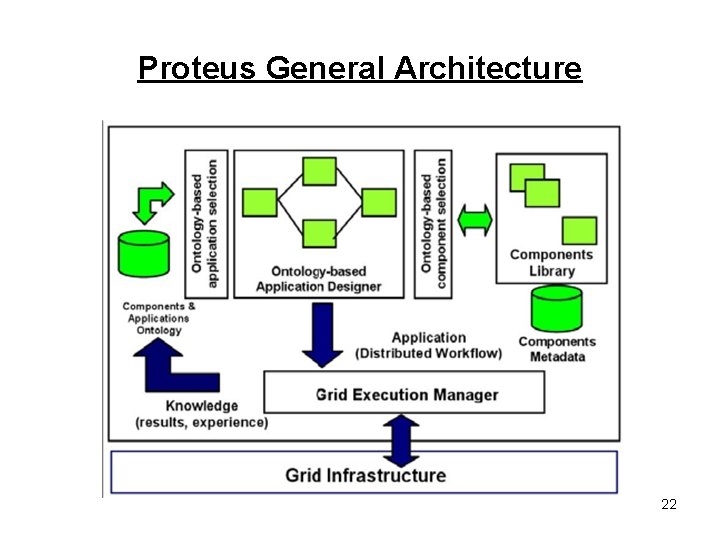
Proteus General Architecture 22
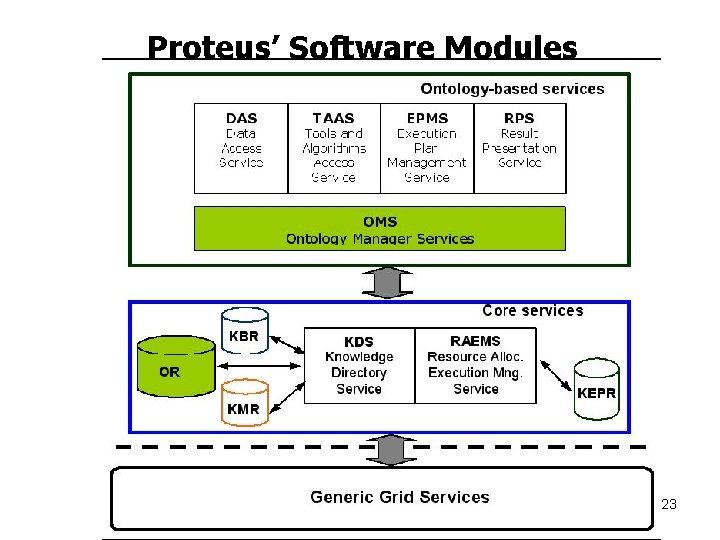
Proteus’ Software Modules 23

Some Taxonomies of the Bioinformatics Ontology 24

Snapshot of the Ontology Browser 25

Human Protein Clustering Workflow 26

Snapshot of VEGA: Workspace 1 of the Data Selection Phase 27

Software Installed in the Example Grid Software Components segret splitfasta blastall cat Tribe-parse Tribe-matrix mcl Tribe-families Grid Nodes Minos k 3 k 4 * * * * 28

Snapshot of the Ontology Browser 29
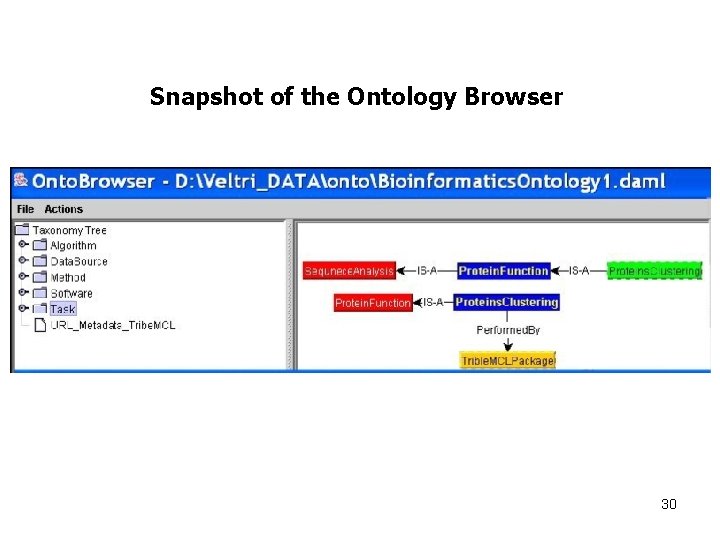
Snapshot of the Ontology Browser 30
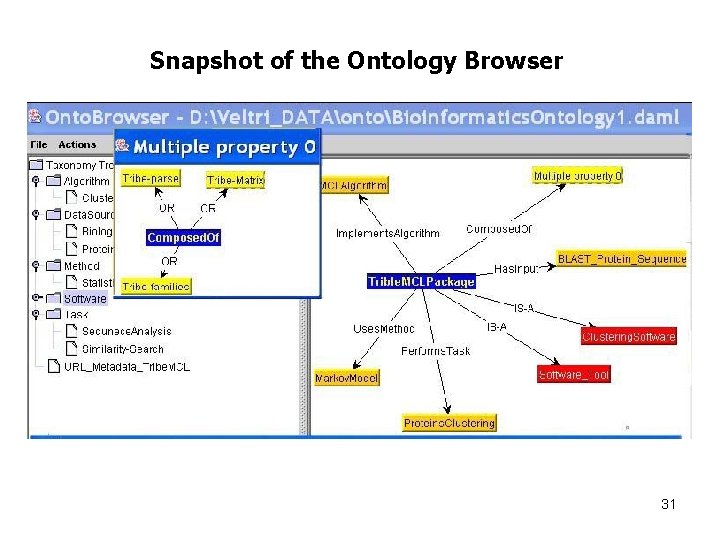
Snapshot of the Ontology Browser 31
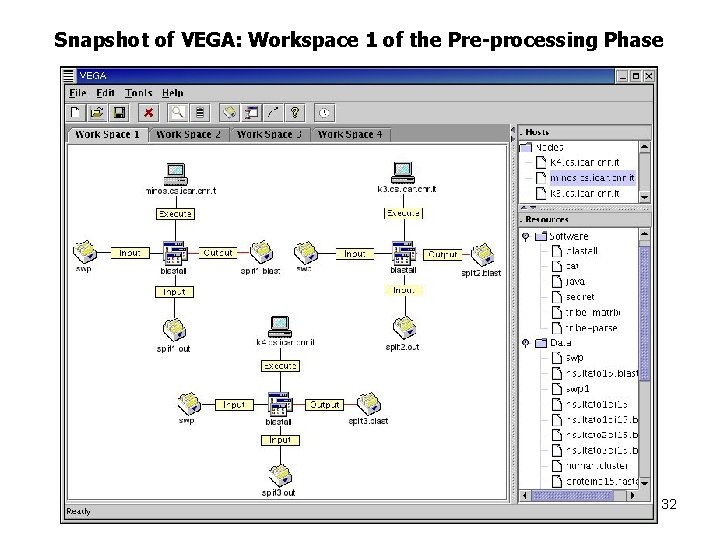
Snapshot of VEGA: Workspace 1 of the Pre-processing Phase 32
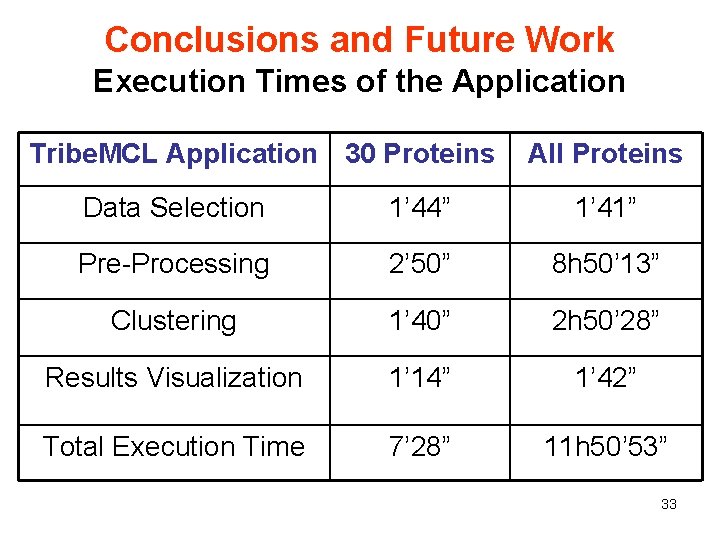
Conclusions and Future Work Execution Times of the Application Tribe. MCL Application 30 Proteins All Proteins Data Selection 1’ 44” 1’ 41” Pre-Processing 2’ 50” 8 h 50’ 13” Clustering 1’ 40” 2 h 50’ 28” Results Visualization 1’ 14” 1’ 42” Total Execution Time 7’ 28” 11 h 50’ 53” 33

References On the paper the authors cited 27 references 34

Questions Thank you 35
- Slides: 35