Proteomics via Mass Spectrometry a bioinformatics perspective Vineet

Proteomics via Mass Spectrometry (a bioinformatics perspective) Vineet Bafna www. cse. ucsd. edu/~vbafna Wi’ 07 Bafna

Nobel Citation 2002 Wi’ 07 Bafna

Nobel Citation, 2002 Wi’ 07 Bafna
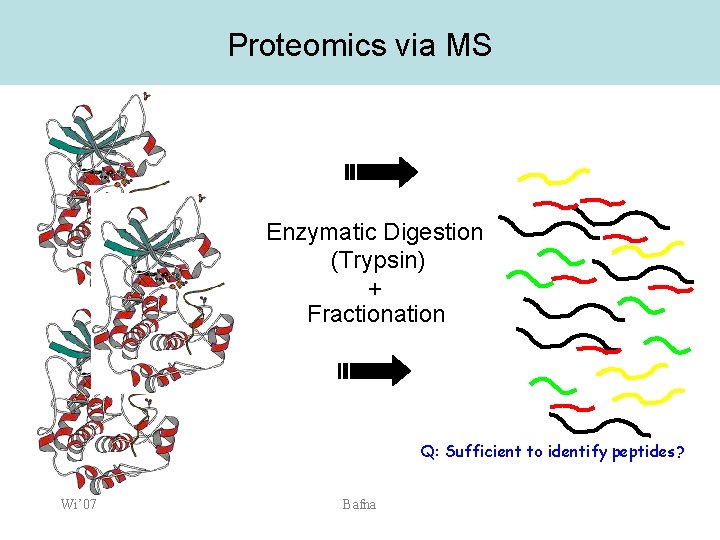
Proteomics via MS Enzymatic Digestion (Trypsin) + Fractionation Q: Sufficient to identify peptides? Wi’ 07 Bafna
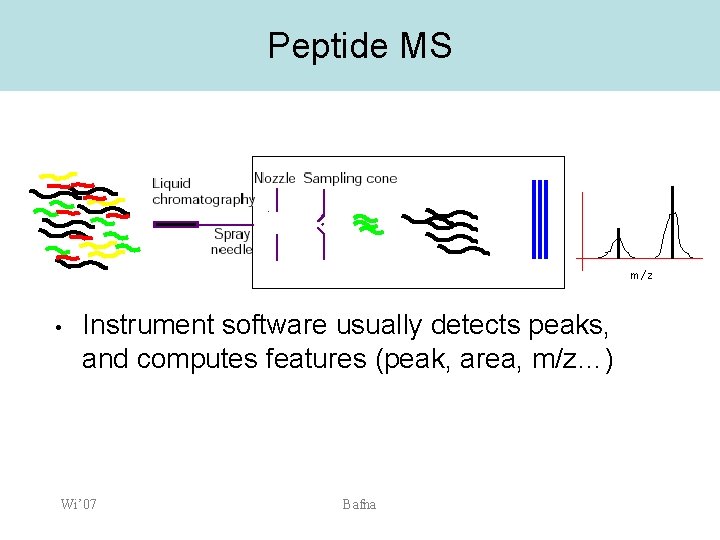
Peptide MS m/z • Instrument software usually detects peaks, and computes features (peak, area, m/z…) Wi’ 07 Bafna
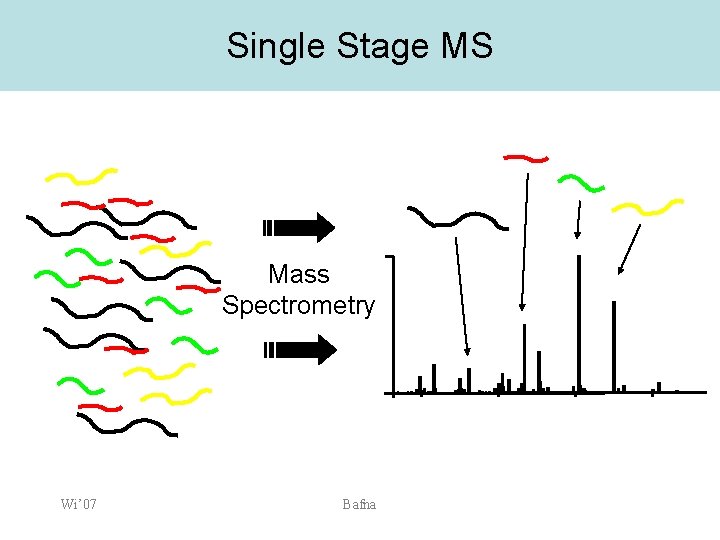
Single Stage MS Mass Spectrometry Wi’ 07 Bafna
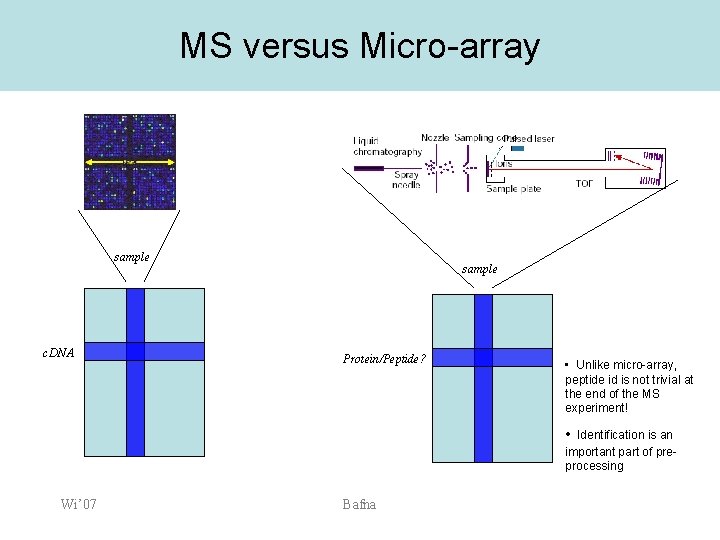
MS versus Micro-array sample c. DNA sample Protein/Peptide? • Unlike micro-array, peptide id is not trivial at the end of the MS experiment! • Identification is an important part of preprocessing Wi’ 07 Bafna

MS based proteomics • Identification – • Quantitation – • Identify all the proteins in the proteome, specific organelles, specific pathways, complexes… Is a protein differentially-expressed in certain conditions? Others – Protein 3 D structure, protein interactions, … We will consider an informatics-centered perspective Wi’ 07 Bafna

Protein Identification • • • The preferred mode is through tandem mass spectrometry of peptides. Is identifying peptides sufficient? Rough probability for co-occurrence of a 15 -aa peptide? With higher accuracy instruments, it may be possible to do intact proteins as well. Wi’ 07 Bafna
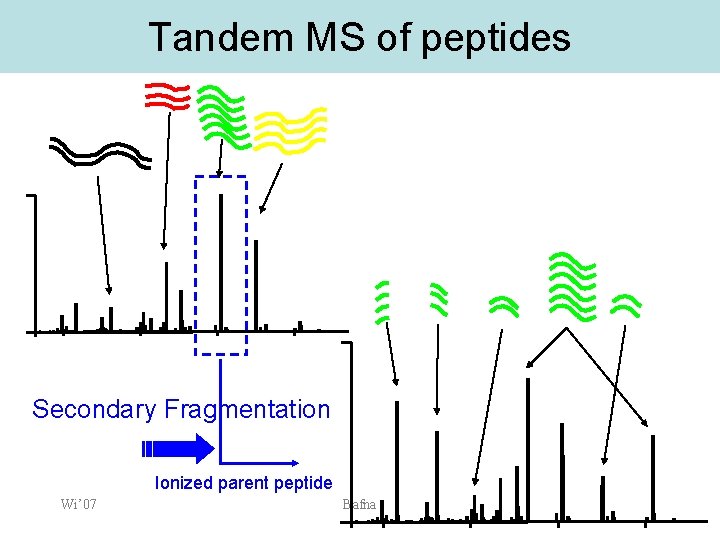
Tandem MS of peptides Secondary Fragmentation Ionized parent peptide Wi’ 07 Bafna
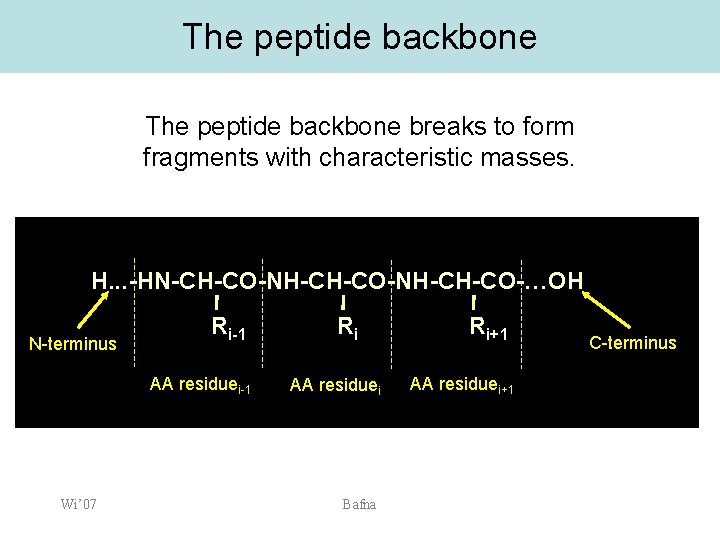
The peptide backbone breaks to form fragments with characteristic masses. H. . . -HN-CH-CO-NH-CH-CO-…OH N-terminus Ri-1 AA residuei-1 Wi’ 07 Ri AA residuei Bafna Ri+1 AA residuei+1 C-terminus
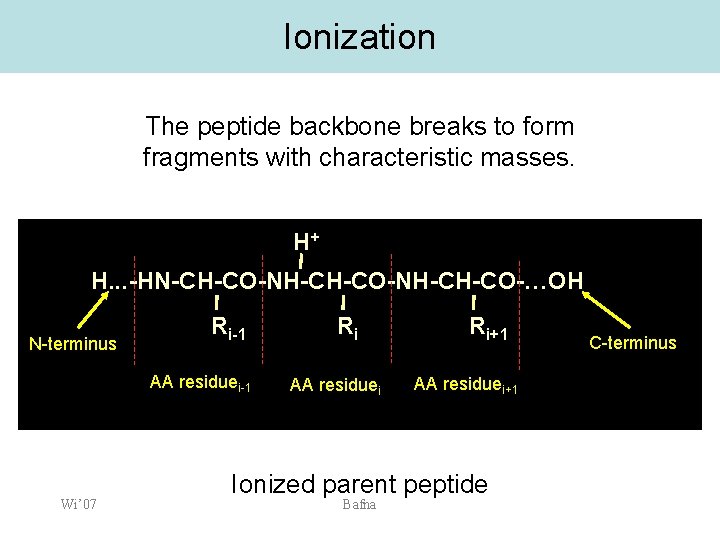
Ionization The peptide backbone breaks to form fragments with characteristic masses. H+ H. . . -HN-CH-CO-NH-CH-CO-…OH N-terminus Ri-1 AA residuei-1 Wi’ 07 Ri AA residuei Ri+1 AA residuei+1 Ionized parent peptide Bafna C-terminus
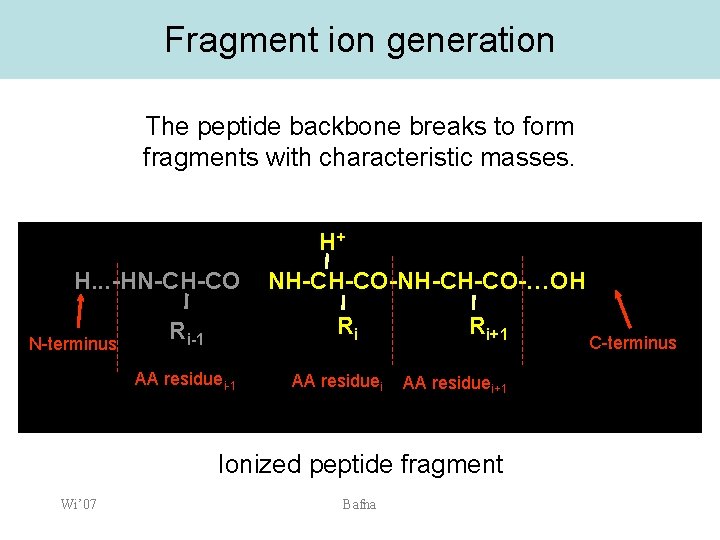
Fragment ion generation The peptide backbone breaks to form fragments with characteristic masses. H+ H. . . -HN-CH-CO N-terminus NH-CH-CO-…OH Ri Ri-1 AA residuei Ri+1 AA residuei+1 Ionized peptide fragment Wi’ 07 Bafna C-terminus
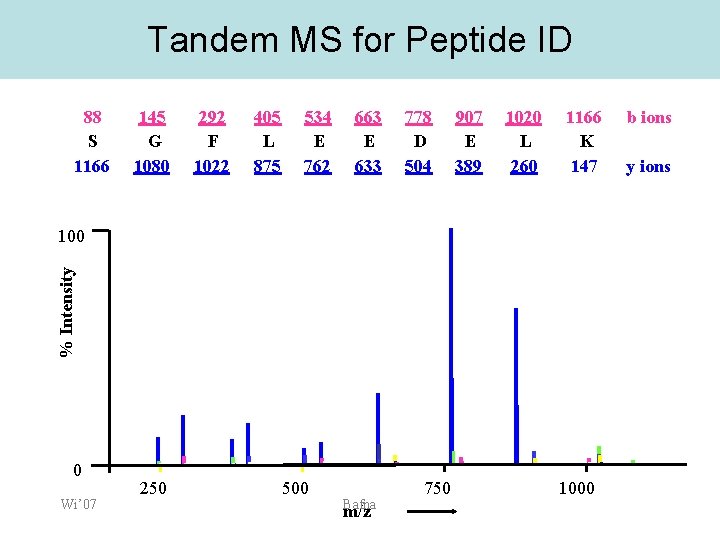
Tandem MS for Peptide ID 88 S 1166 145 G 1080 292 F 1022 405 L 875 534 E 762 663 E 633 778 D 504 907 E 389 1020 L 260 1166 K 147 % Intensity 100 0 Wi’ 07 [M+2 H]2+ 250 500 Bafna m/z 750 1000 b ions y ions
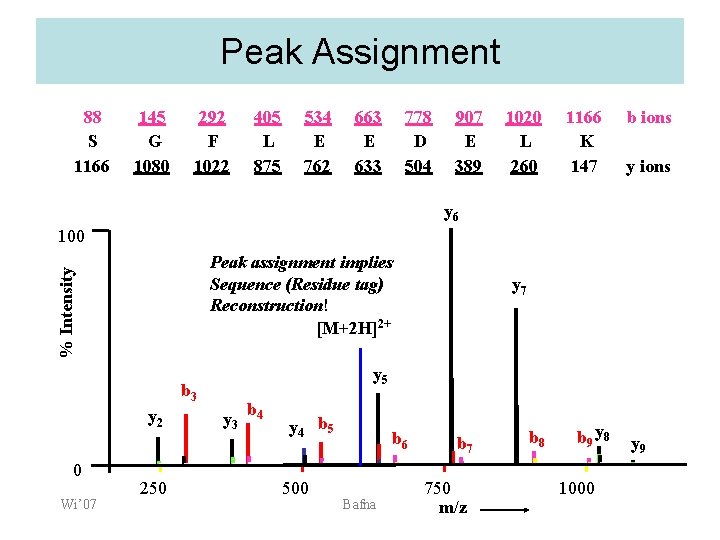
Peak Assignment 88 S 1166 145 G 1080 292 F 1022 405 L 875 534 E 762 663 E 633 778 D 504 907 E 389 1020 L 260 1166 K 147 b ions y 6 100 % Intensity Peak assignment implies Sequence (Residue tag) Reconstruction! [M+2 H]2+ y 5 b 3 y 2 0 Wi’ 07 250 y 7 y 3 b 4 y 4 b 5 500 b 6 Bafna b 7 750 m/z b 8 b 9 y 8 1000 y 9
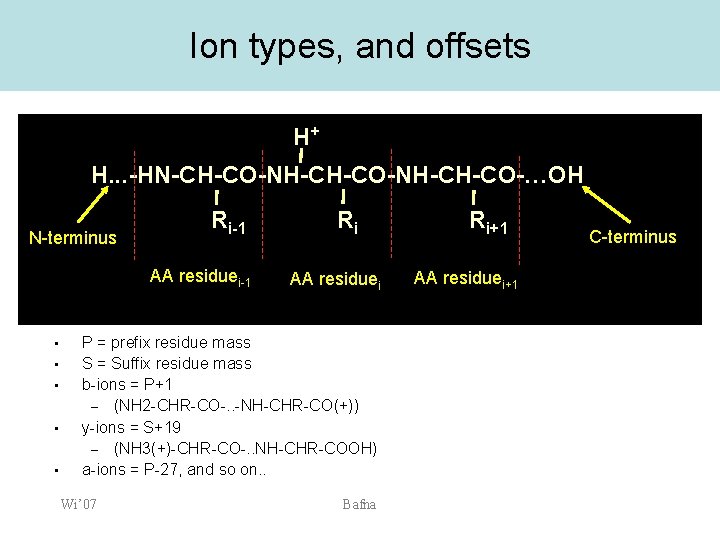
Ion types, and offsets H+ H. . . -HN-CH-CO-NH-CH-CO-…OH N-terminus Ri-1 AA residuei-1 • • • Ri AA residuei P = prefix residue mass S = Suffix residue mass b-ions = P+1 – (NH 2 -CHR-CO-. . -NH-CHR-CO(+)) y-ions = S+19 – (NH 3(+)-CHR-CO-. . NH-CHR-COOH) a-ions = P-27, and so on. . Wi’ 07 Bafna Ri+1 AA residuei+1 C-terminus

MS Quiz: • • Why aren’t all tandem MS peaks of the same intensity? Do the intensities for a peptide vary from spectrum to spectrum? Wi’ 07 Bafna

Database Searching for peptide ID • For every peptide from a database – – • …SARLSQETFSDLWKLLPENNVLSPLP…. Reject if it has the wrong mass, else: Generate a hypothetical spectrum Compute a correlation between observed and experimental spectra Choose the best Database searching is very powerful and is the de facto standard for MS. – Wi’ 07 Sequest, Mascot, Inspect, and many others Bafna

So what’s new? • • The Id picture is very simplistic. Only 20 -30% of spectra are conclusively identified. Many reasons: – – • • Spectra are noisy. Databases are incomplete. Sometimes, we need to do a de novo interpretation Post-translational modifications. Instrument performance is critical. The algorithms for identification must be sensitive to these issues. We present a systematic look at identification software. Wi’ 07 Bafna
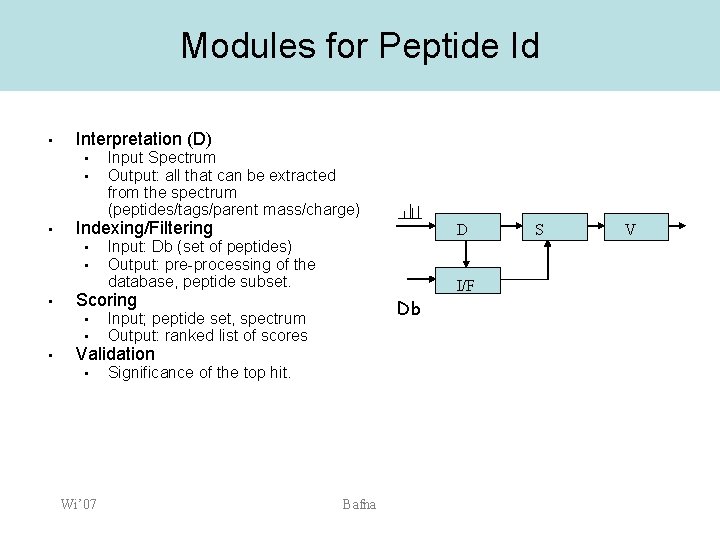
Modules for Peptide Id • Interpretation (D) • • • Indexing/Filtering • • • D Input: Db (set of peptides) Output: pre-processing of the database, peptide subset. I/F Scoring • • • Input Spectrum Output: all that can be extracted from the spectrum (peptides/tags/parent mass/charge) Db Input; peptide set, spectrum Output: ranked list of scores Validation • Wi’ 07 Significance of the top hit. Bafna S V
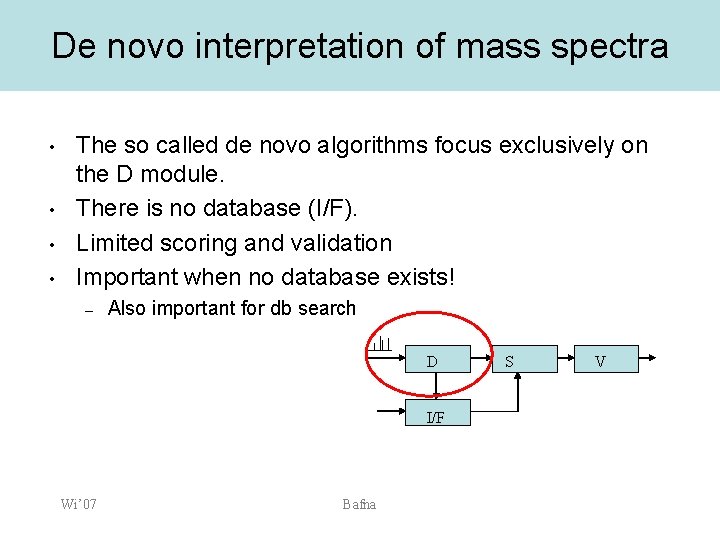
De novo interpretation of mass spectra • • The so called de novo algorithms focus exclusively on the D module. There is no database (I/F). Limited scoring and validation Important when no database exists! – Also important for db search D I/F Wi’ 07 Bafna S V
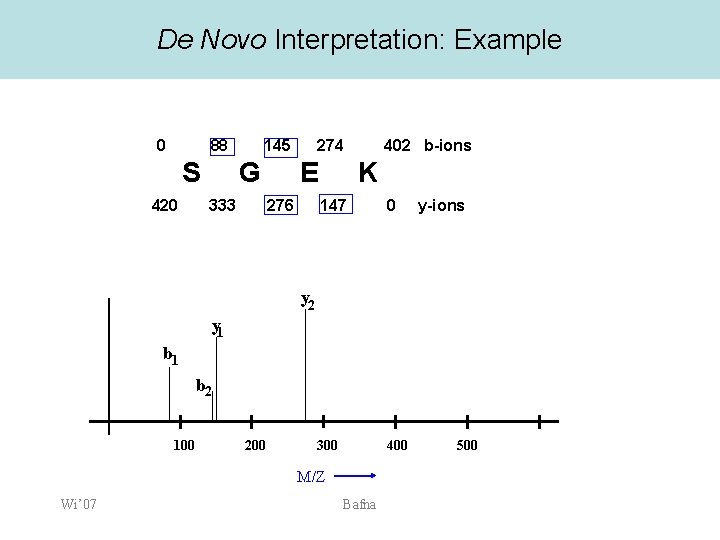
De Novo Interpretation: Example 0 88 S 420 145 G 333 274 E 276 402 b-ions K 147 0 300 400 y-ions y 2 y 1 b 2 100 200 M/Z Wi’ 07 Bafna 500

The simplest case 0 88 S 420 S 145 G 333 88 274 E 276 G K 147 • 0 y-ions 145 E • 402 b-ions 274 K 402 Suppose only (and all) the prefix ions were visible. Would identification be easy? We have two problems: – – Wi’ 07 There is a mix of b and y ions. Separating them is critical! Other ions besides b, y, including neutral losses, noise and so on. We need to account for them. Bafna

• • • Separating b-, and y-ions is solved using a combinatorial formulation (forbidden pairs) Separating b, y from all others is solved using a statistical approach. Together, they form the basis for a de novo sequencer. Wi’ 07 Bafna
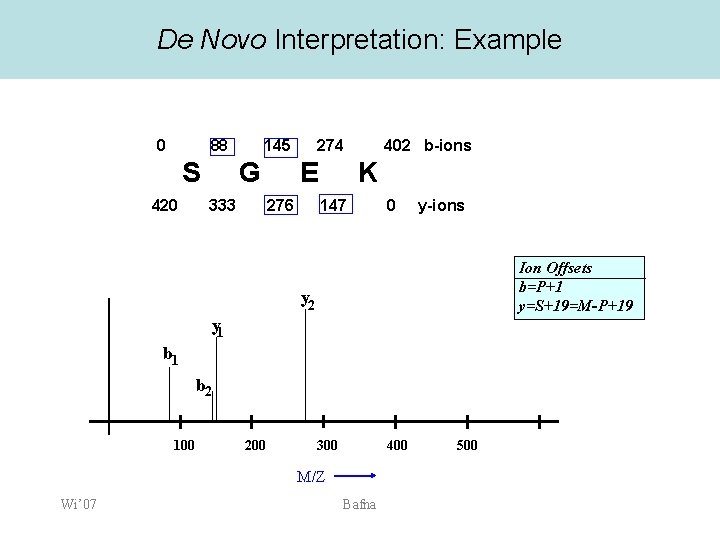
De Novo Interpretation: Example 0 88 S 420 145 G 333 274 E 276 402 b-ions K 147 0 y-ions Ion Offsets b=P+1 y=S+19=M-P+19 y 2 y 1 b 2 100 200 300 400 M/Z Wi’ 07 Bafna 500

Computing possible prefixes • • • We know the parent mass M=401. Consider a mass value 88 Assume that it is a b-ion, or a y-ion If b-ion, it corresponds to a prefix of the peptide with residue mass 88 -1 = 87. If y-ion, y=M-P+19. – Therefore the prefix has mass • P=M-y+19= 401 -88+19=332 Compute all possible Prefix Residue Masses (PRM) for all ions. Wi’ 07 Bafna

Putative Prefix Masses • Prefix Mass M=401 88 145 147 276 S 0 Wi’ 07 b 87 144 146 275 y 332 275 273 144 • G E K 87 144 273 401 Bafna Only a subset of the prefix masses are correct. The correct mass values form a ladder of amino-acid residues
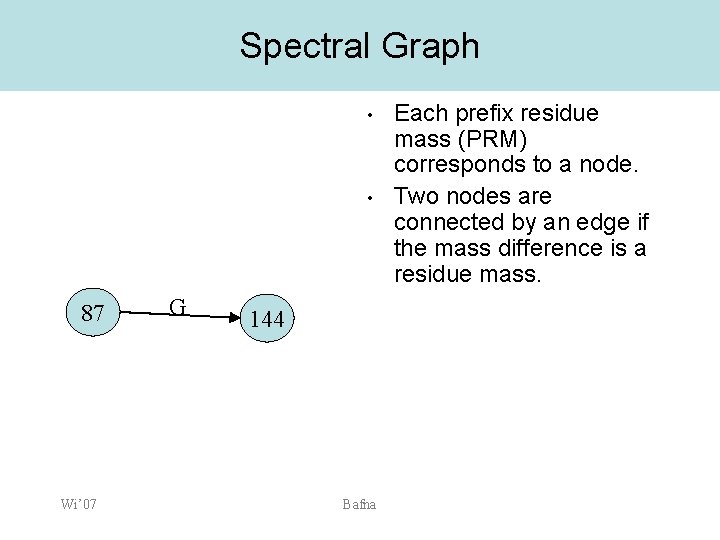
Spectral Graph • • 87 Wi’ 07 G 144 Bafna Each prefix residue mass (PRM) corresponds to a node. Two nodes are connected by an edge if the mass difference is a residue mass.

Spectral Graph • • Each peak, when assigned to a prefix/suffix ion type generates a unique prefix residue mass. Spectral graph: – – – Each node u defines a putative prefix residue M(u). (u, v) in E if M(v)-M(u) is the residue mass of an a. a. (tag) or 0. Paths in the spectral graph correspond to a interpretation 0 87 100 S Wi’ 07 273275 144 146 G 200 332 300 E K Bafna 401
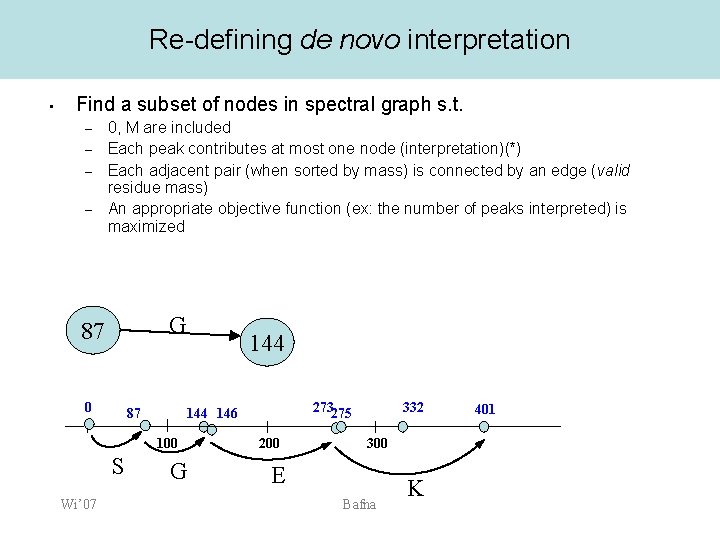
Re-defining de novo interpretation • Find a subset of nodes in spectral graph s. t. – – 0, M are included Each peak contributes at most one node (interpretation)(*) Each adjacent pair (when sorted by mass) is connected by an edge (valid residue mass) An appropriate objective function (ex: the number of peaks interpreted) is maximized G 87 0 87 Wi’ 07 273275 144 146 100 S 144 G 200 332 300 E Bafna K 401

Two problems • Too many nodes. – – – A. Only a small fraction correspond to b/y ions (leading to true PRMs). B. Even if the b/y ions were correctly predicted, each peak generates multiple possibilities, only one of which is correct. We need to find a path that uses each peak only once (algorithmic problem). In general, the forbidden pairs problem is NP-hard 0 87 100 S Wi’ 07 273275 144 146 G 200 332 300 E Bafna K 401

However, . . • • The b, y ions have a special non-interleaving property Consider pairs (b 1, y 1), (b 2, y 2) – – Wi’ 07 Note that b 1+y 1 = b 2+y 2 If (b 1 < b 2), then y 1 > y 2 Bafna
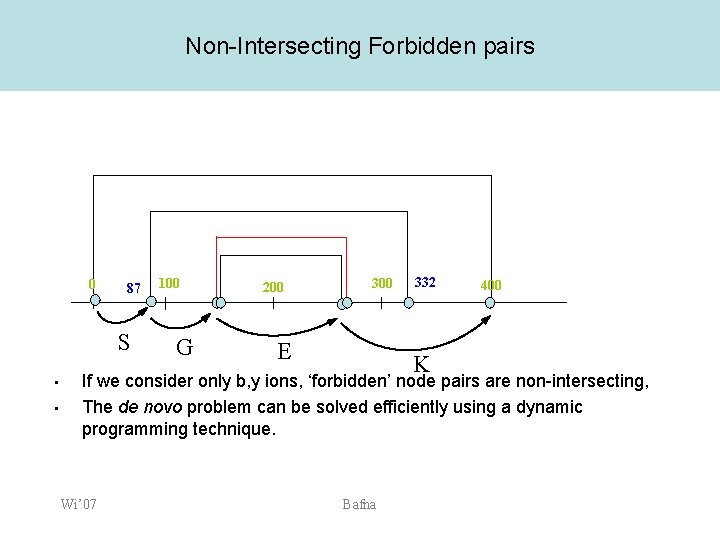
Non-Intersecting Forbidden pairs 0 87 S • • 100 G 200 300 E 332 K 400 If we consider only b, y ions, ‘forbidden’ node pairs are non-intersecting, The de novo problem can be solved efficiently using a dynamic programming technique. Wi’ 07 Bafna

The forbidden pairs method • • There may be many paths that avoid forbidden pairs. We choose a path that maximizes an objective function, – – Wi’ 07 EX: the number of peaks interpreted Here we assume a function , which gives a score to a PRM. The score captures the likelihood that the PRM is correct. Bafna
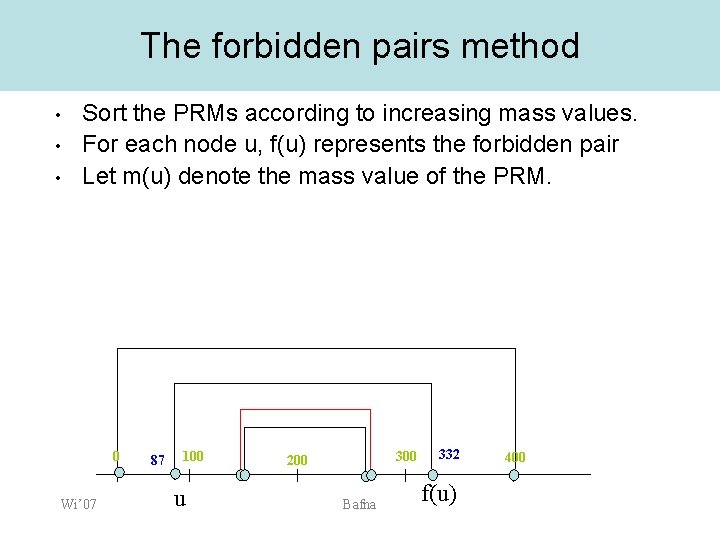
The forbidden pairs method • • • Sort the PRMs according to increasing mass values. For each node u, f(u) represents the forbidden pair Let m(u) denote the mass value of the PRM. 0 Wi’ 07 87 100 u 300 200 Bafna 332 f(u) 400
![D. P. forbidden pairs • Consider all pairs u, v – • • m[u] D. P. forbidden pairs • Consider all pairs u, v – • • m[u]](http://slidetodoc.com/presentation_image/32973d466a93ba3fd383b630575d8e50/image-36.jpg)
D. P. forbidden pairs • Consider all pairs u, v – • • m[u] <= M/2, m[v] >M/2 Define S(u, v) as the best score of a forbidden pair path from 0 ->u, v>M Is it sufficient to compute S(u, v) for all u, v? 0 Wi’ 07 87 100 u 300 200 Bafna 332 400 v
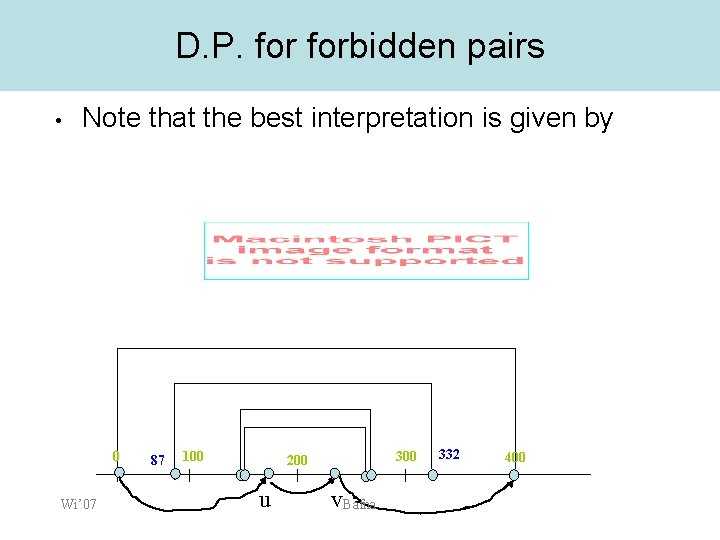
D. P. forbidden pairs • Note that the best interpretation is given by 0 Wi’ 07 87 100 300 200 u v. Bafna 332 400
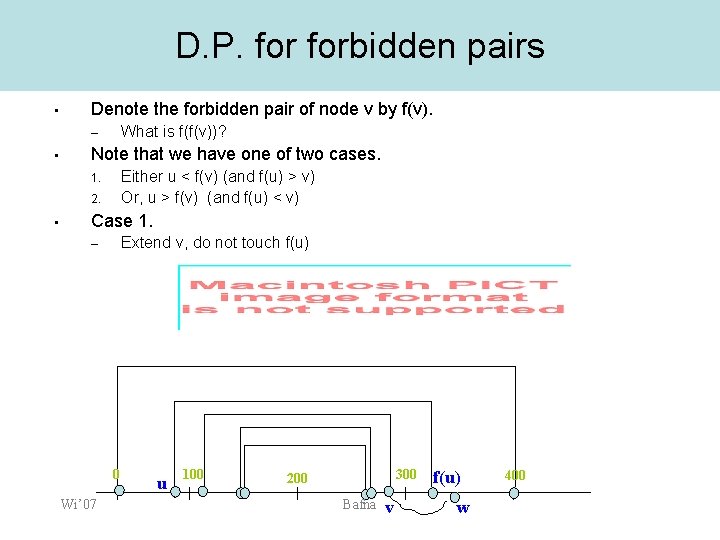
D. P. forbidden pairs • Denote the forbidden pair of node v by f(v). What is f(f(v))? – • Note that we have one of two cases. Either u < f(v) (and f(u) > v) Or, u > f(v) (and f(u) < v) 1. 2. • Case 1. Extend v, do not touch f(u) – 0 Wi’ 07 u 100 300 200 Bafna v f(u) w 400

The complete algorithm for all u /*increasing mass values from 0 to M/2 */ for all v /*decreasing mass values from M to M/2 */ if (u > f[v]) else if (u < f[v]) If (u, v) E /*max. I is the score of the best interpretation*/ max. I = max {max. I, S[u, v]} Wi’ 07 Bafna

De Novo: Second issue • • Given only b, y ions, a forbidden pairs path will solve the problem. However, recall that there are MANY other ion types. – – Wi’ 07 Typical length of peptide: 15 Typical # peaks? 50 -150? #b/y ions? Most ions are “Other” • a ions, neutral losses, isotopic peaks…. Bafna
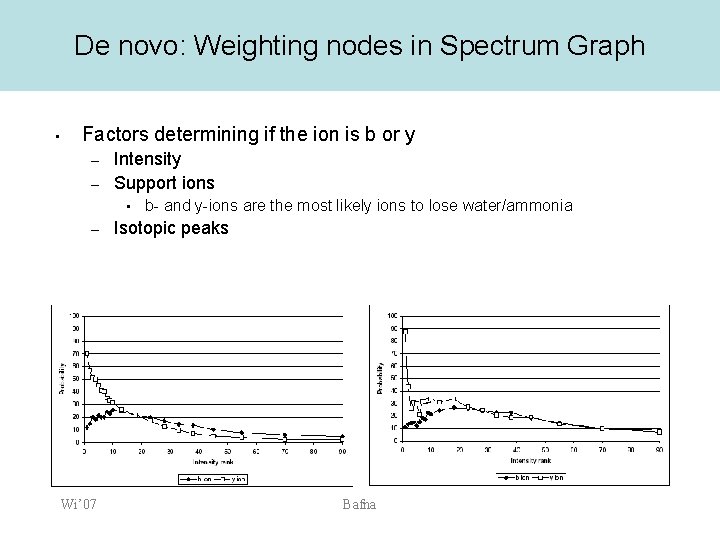
De novo: Weighting nodes in Spectrum Graph • Factors determining if the ion is b or y – – Intensity Support ions • – Wi’ 07 b- and y-ions are the most likely ions to lose water/ammonia Isotopic peaks Bafna
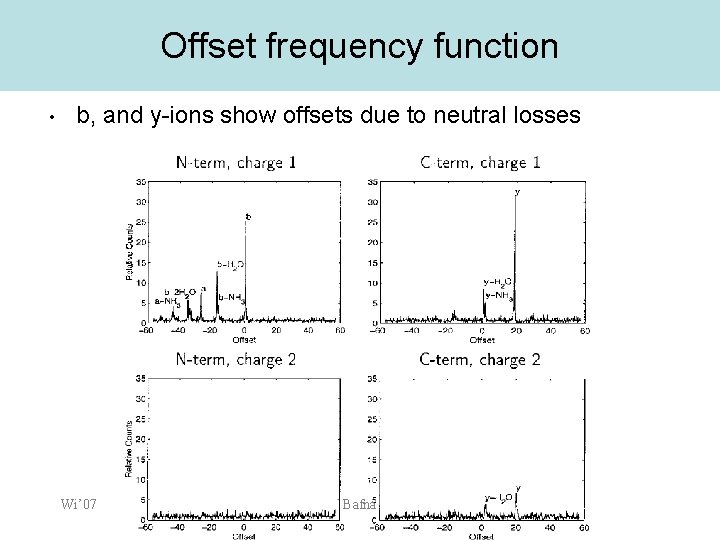
Offset frequency function • b, and y-ions show offsets due to neutral losses Wi’ 07 Bafna
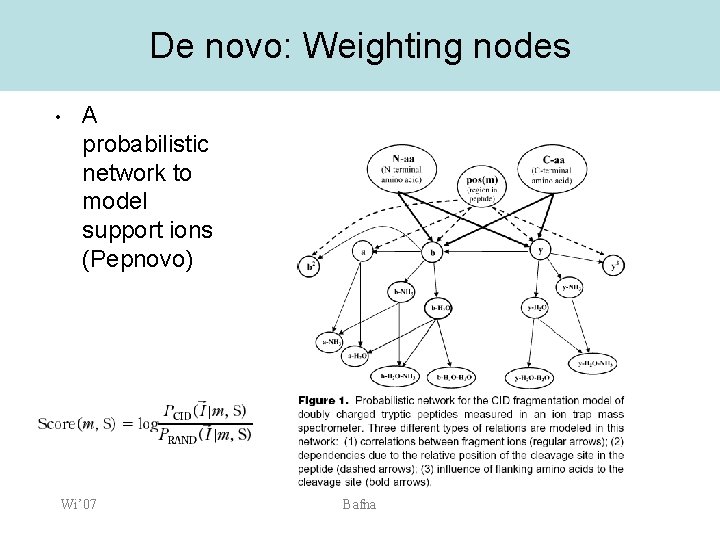
De novo: Weighting nodes • A probabilistic network to model support ions (Pepnovo) Wi’ 07 Bafna

De Novo Interpretation Summary • • The main challenge is to separate b/y ions from everything else (weighting nodes), and separating the prefix ions from the suffix ions (Forbidden Pairs). As always, the abstract idea must be supplemented with many details. – – Wi’ 07 Noise peaks, incomplete fragmentation A PRM is first scored on its likelihood of being correct, and the forbidden pair method is applied subsequently. Bafna
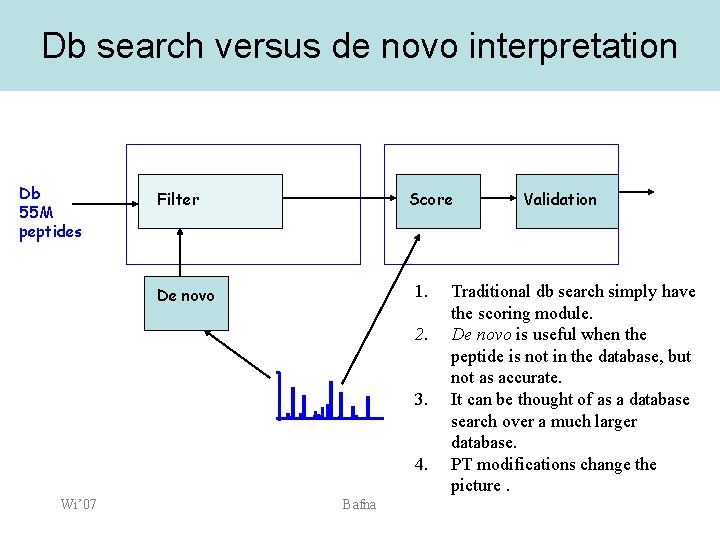
Db search versus de novo interpretation Db 55 M peptides Filter Score 1. De novo 2. 3. 4. Wi’ 07 Bafna Validation Traditional db search simply have the scoring module. De novo is useful when the peptide is not in the database, but not as accurate. It can be thought of as a database search over a much larger database. PT modifications change the picture.
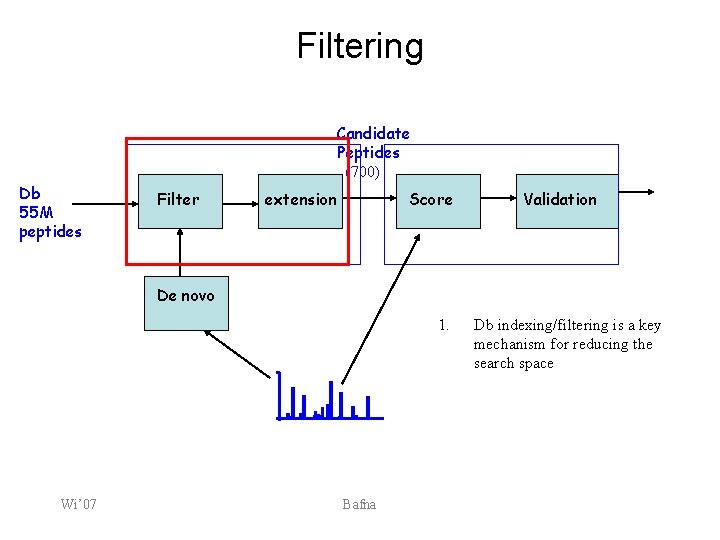
Filtering Candidate Peptides (700) Db 55 M peptides Filter extension Score Validation De novo 1. Wi’ 07 Bafna Db indexing/filtering is a key mechanism for reducing the search space
Filtering Define a filter as a computational tool that rapidly screens a database, removing much of it but retaining the true peptide. Can you suggest commonly used filters? • • 1. 2. Wi’ 07 Parent mass Trypsin digested peptides Bafna

Parent Mass filter • • • Sort all peptides in the database by their parent mass. Search only the peptides that are within some mass tolerance. The filter does not work when you have modifications. Wi’ 07 Bafna
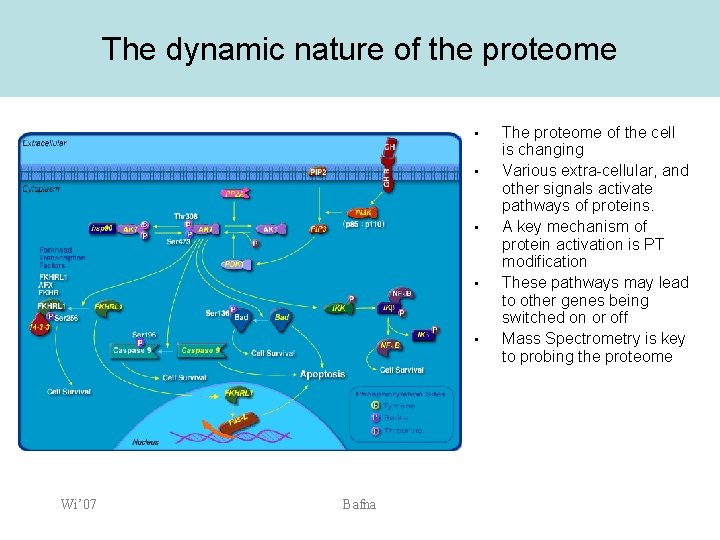
The dynamic nature of the proteome • • • Wi’ 07 Bafna The proteome of the cell is changing Various extra-cellular, and other signals activate pathways of proteins. A key mechanism of protein activation is PT modification These pathways may lead to other genes being switched on or off Mass Spectrometry is key to probing the proteome
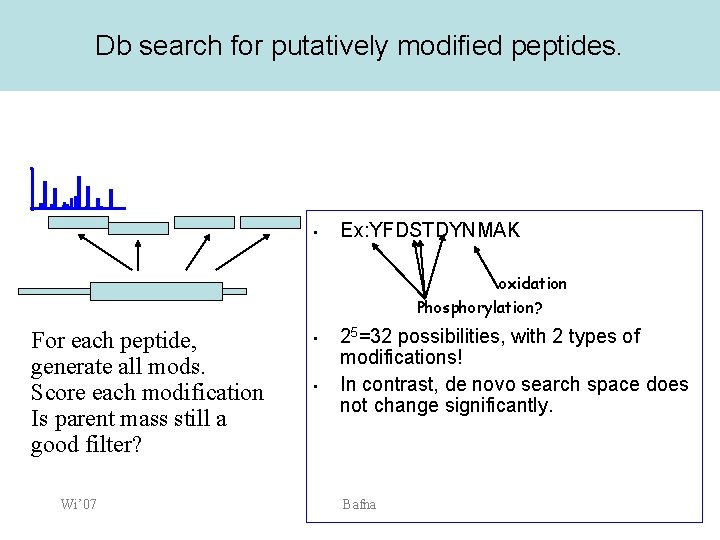
Db search for putatively modified peptides. • Ex: YFDSTDYNMAK oxidation Phosphorylation? For each peptide, generate all mods. Score each modification Is parent mass still a good filter? Wi’ 07 • • 25=32 possibilities, with 2 types of modifications! In contrast, de novo search space does not change significantly. Bafna

Enzymatic digestion rules as a filter • Consider only tryptic peptides – • Trypsin cleaves after R, K (not if RP, or KP) Tryptic peptide filters may not be very effective – – – Wi’ 07 Missed cleavage End-point degradation Endogenous peptide activity Bafna

Example of non-tryptic peptide analysis • • • Experiment: a massive oversampling of a proteome (14 M spectra of a prokaryotic genome) Plot the absolute postion of the most N-terminal peptide (not-necessarily tryptic) (Gupta et al. , 2007) Two peaks are seen, at position 2, and position ~22 Wi’ 07 Bafna

Signal Peptide discovery Wi’ 07 Bafna

PT modifications/processing • • • The problem was not intractable, but impractical. Identifications of modified peptides was not routine, and is left to specialized cases. Better filtering technology makes it practical to explore modifications on a large scale. Increase in time is modest with increasing number of modifications. The technology can only improve. Should be possible to improve speed by another order of magnitude. Wi’ 07 Bafna

We will filter databases via a trick from sequence searching Wi’ 07 Bafna

Sequence Search Basics 1: POTATO 2: POTASSIUM 3: TASTE POTASTPOTATO database dictionary • • Q: Given k words (si has length li), and a database of size n, find all matches to these words in the database string. How fast can this be done? Wi’ 07 Bafna

Dict. Matching & string matching • How fast can you do it, if you only had one word of length m? – – • Dictionary matching – – – • Trivial algorithm O(nm) time Pre-processing O(m), Search O(n) time. Trivial algorithm (l 1+l 2+l 3…)n Using a keyword tree, lpn (lp is the length of the longest pattern) Aho-Corasick: O(n) after preprocessing O(l 1+l 2. . ) We will consider the most general case Wi’ 07 Bafna

Sequence tag filters Basics 1. 2. 3. De novo sequencing can be used to get partial sequence information from the spectra. Exact matching for sequence is fast. We can search a database (size n) with k substrings (of any length) in time that is proportional to n, but independent of the size or number of substrings. • Aho-Corasick trie data-structure. Wi’ 07 Bafna
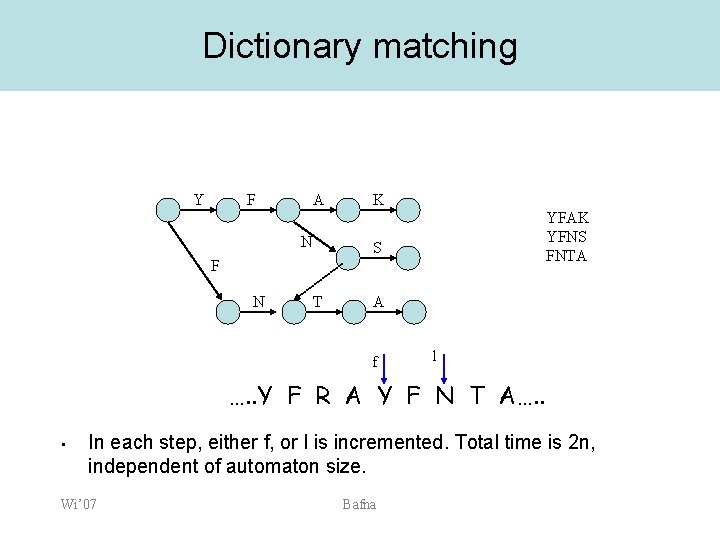
Dictionary matching Y F A N K YFAK YFNS FNTA S F N T A f l …. . Y F R A Y F N T A…. . • In each step, either f, or l is incremented. Total time is 2 n, independent of automaton size. Wi’ 07 Bafna

Tag-based filtering • • • A tag is a short peptide with a prefix and suffix mass Efficient: An average tripeptide tag matches Swiss-Prot ~700 times Tagging is related to de novo sequencing yet different. Objective: Compute a subset of short strings, at least one of which must be correct. Longer tags=> better filter. Analogy: Using tags to search the proteome is similar to moving from full Smith-Waterman alignment to BLAST Wi’ 07 Bafna
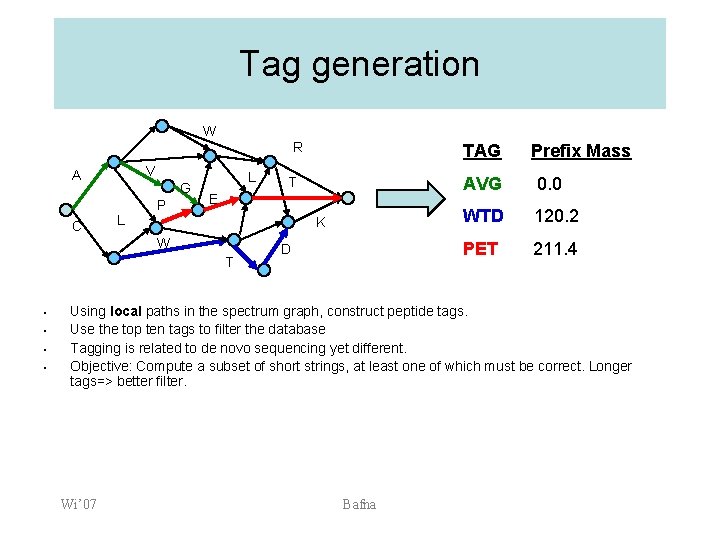
Tag generation W V A C R G L P L K T • • • T E W • TAG D Prefix Mass AVG 0. 0 WTD 120. 2 PET 211. 4 Using local paths in the spectrum graph, construct peptide tags. Use the top ten tags to filter the database Tagging is related to de novo sequencing yet different. Objective: Compute a subset of short strings, at least one of which must be correct. Longer tags=> better filter. Wi’ 07 Bafna

Tag-based search • • • Tags (from multiple spectra) are used to construct a trie For each string match, attempt to extend, matching the prefix and suffix mass using flanking sequence (and PTMs) Retain best matches for detailed scoring Wi’ 07 Bafna
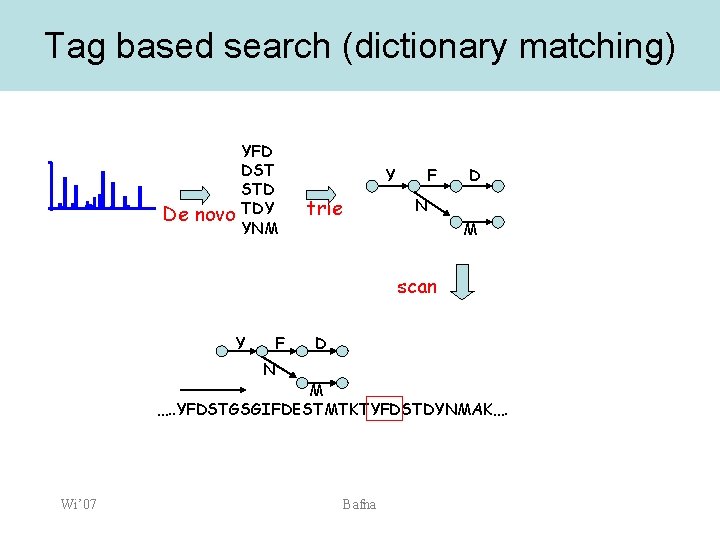
Tag based search (dictionary matching) De novo YFD DST STD TDY YNM Y trie F D N M scan Y F D N M …. . YFDSTGSGIFDESTMTKTYFDSTDYNMAK…. Wi’ 07 Bafna

Speed & Sensitivity • • • Filtered. Peptides: # peptides that pass the filter Time: Scan time + filteredpeptides * Scoring-time For sequence tag filters, the scan time can be amortized out, by combining scan for many spectra all at once. – Build one automaton from multiple spectra Thus, filter efficiency is key to speed. Filter sensitivity is also important Wi’ 07 Bafna

Fast Extension • Given: – – tag with prefix and suffix masses <m. P> xyz <m. S> match in the database <m. P>xyz<m. S> xyz • • Compute if a suffix and prefix match with allowable modifications. How fast can this be done? Compute a candidate peptide with most likely attachment point. Wi’ 07 Bafna

Filtering • • • Database filtering is a critical (and relatively unexplored) strategy for MS searches. De novo sequencing to get tags is an effective strategy Are other forms of filtering possible? – Wi’ 07 Alg. Question: given a spectrum, find all peptides that match a subset of theoretical fragments. How quickly can you do that? Bafna
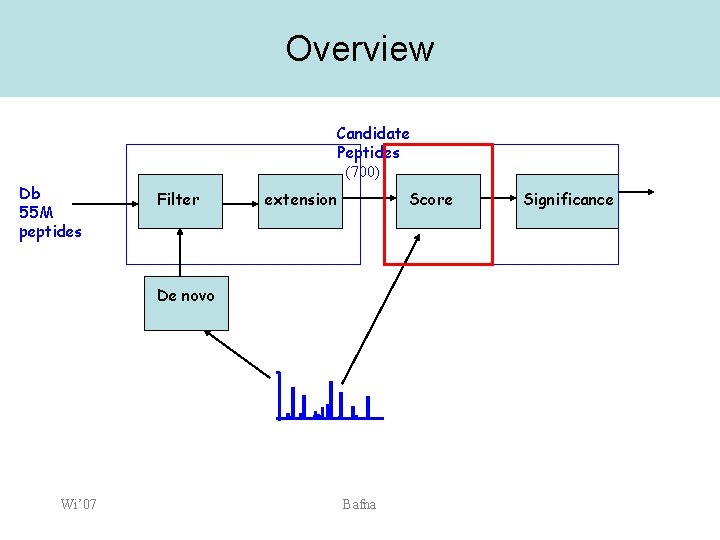
Overview Candidate Peptides (700) Db 55 M peptides Filter extension Score De novo Wi’ 07 Bafna Significance

Scoring • Input: – – • Candidate peptide with attached modifications Spectrum Output: – – Wi’ 07 Score function: Key: the score must normalize for length, as variable modifications can change peptide length. Bafna

Score function • • • Score is a log-odds function. Numerator: Prob. That the spectrum was generated by n theoretical fragments from the peptide. Denominator: Probability that the spectrum was generated by n randomly generated peaks Wi’ 07 Bafna
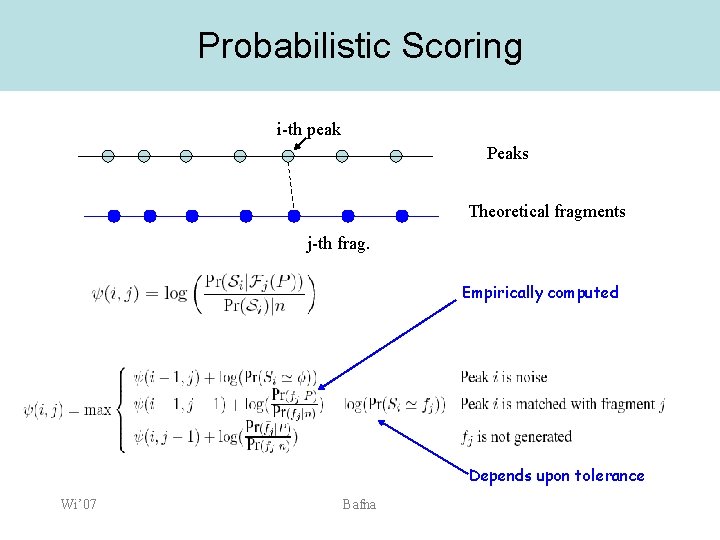
Probabilistic Scoring i-th peak Peaks Theoretical fragments j-th frag. Empirically computed Depends upon tolerance Wi’ 07 Bafna
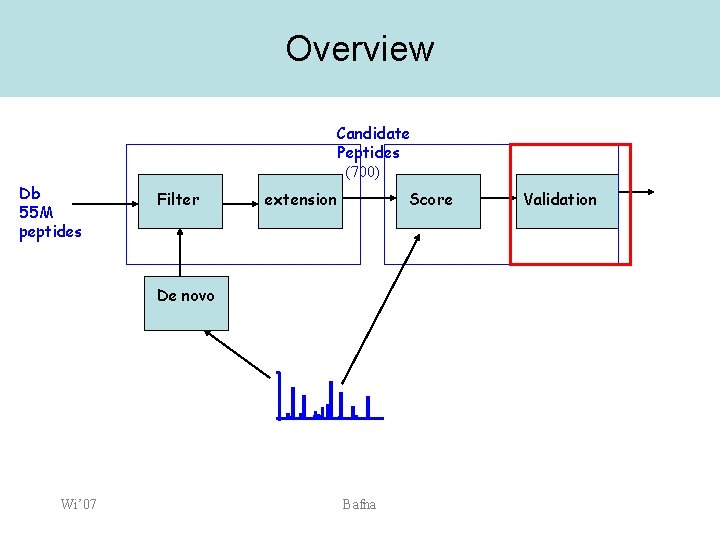
Overview Candidate Peptides (700) Db 55 M peptides Filter extension Score De novo Wi’ 07 Bafna Validation

P-value computation • • The score function allows us to rank peptides. The top scoring one may not be the correct one. We consider a collection of +ve (top scoring correct one), and -ve spectra. Consider a bunch of other scores that help separate the +ve from the -ve. Wi’ 07 Bafna

Quality scores and p-values • Features: • • • Score S: as computed Explained Intensity I: fraction of total intensity explained by annotated peaks. Explained peaks: fraction of top 25 peaks annotated. b-y score B: fraction of b+y ions annotated -score : difference between the best and second best Choose a final score as a (linear) combination of the features. The weights can be trained using a discriminative strategy. Wi’ 07 Bafna
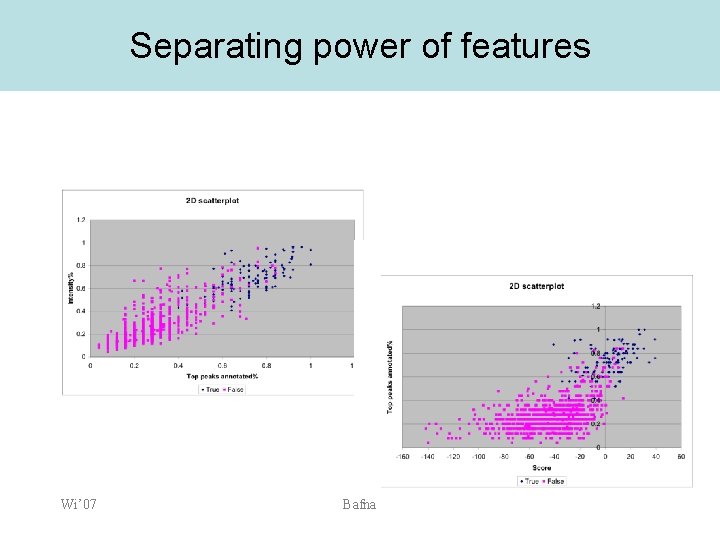
Separating power of features Wi’ 07 Bafna
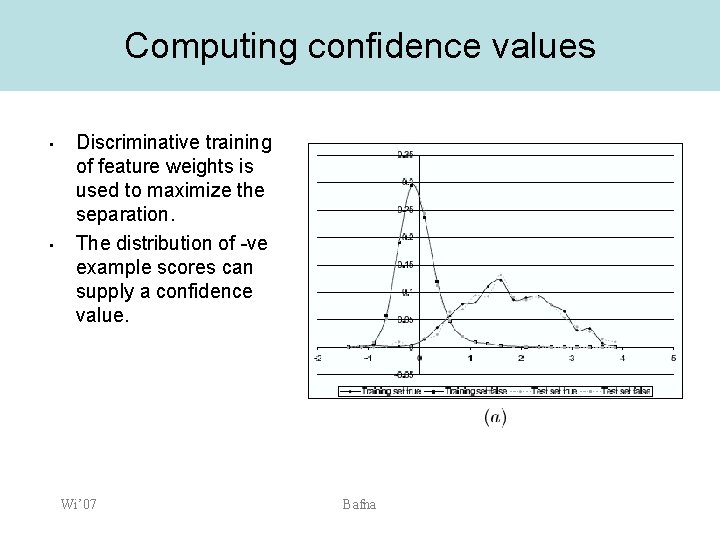
Computing confidence values • • Discriminative training of feature weights is used to maximize the separation. The distribution of -ve example scores can supply a confidence value. Wi’ 07 Bafna

Common ID Tools • Sequest – – – • • De Novo + Filtering: Parent Mass, Enzyme specificity Scoring: Simple cross-correlation of theoretical and experimental peaks Validation: Xcorr (based on difference between best and second best) Mascot: Similar MS-BLAST – – Wi’ 07 Uses 3 rd party de novo prediction Filtering using Blast code Limited scoring Validation at the protein level Bafna

Identification summary • • • While MS technologies are important for proteomics in general, they are the key technology for identification. They can probe the proteome dynamically, and help identify mutations and modifications. The algorithms are continually improved by improved versions of one or more modules. Wi’ 07 Bafna

Modifications • While we discussed modification in general, identification and validation of modifications remains an important theme. • Larger modifications are interesting in themselves (such that glycan chains in glycosylation), and might be identified using MS ABRF delta mass resource Wi’ 07 Bafna

Isotopic Profiles Wi’ 07 Bafna
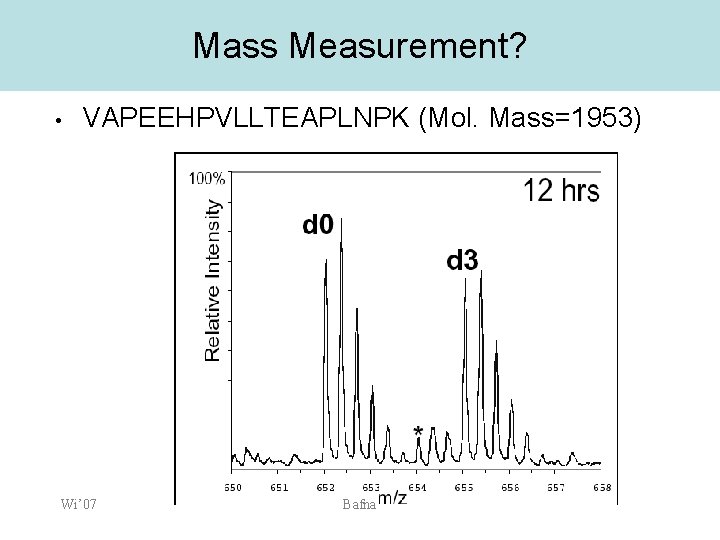
Mass Measurement? • VAPEEHPVLLTEAPLNPK (Mol. Mass=1953) Wi’ 07 Bafna

Mass-Charge ratio • • The charge is due to a proton (H+) which has a mass of ~1 Da The X-axis is (M+Z)/Z – – Z=1 implies that peak is at M+1 Z=2 implies that peak is at (M+2)/2 • – Wi’ 07 M=1000, Z=2, peak position is at 501 Suppose you see a peak at 501. Is the mass 500, or is it 1000? Bafna
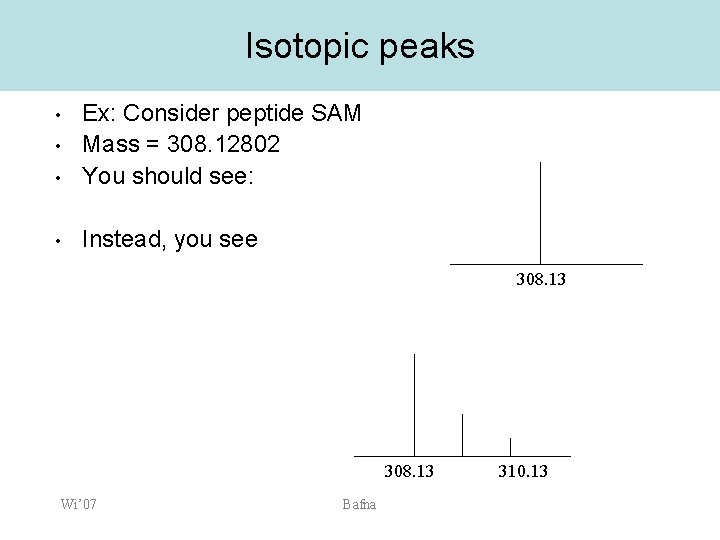
Isotopic peaks • Ex: Consider peptide SAM Mass = 308. 12802 You should see: • Instead, you see • • 308. 13 Wi’ 07 Bafna 310. 13
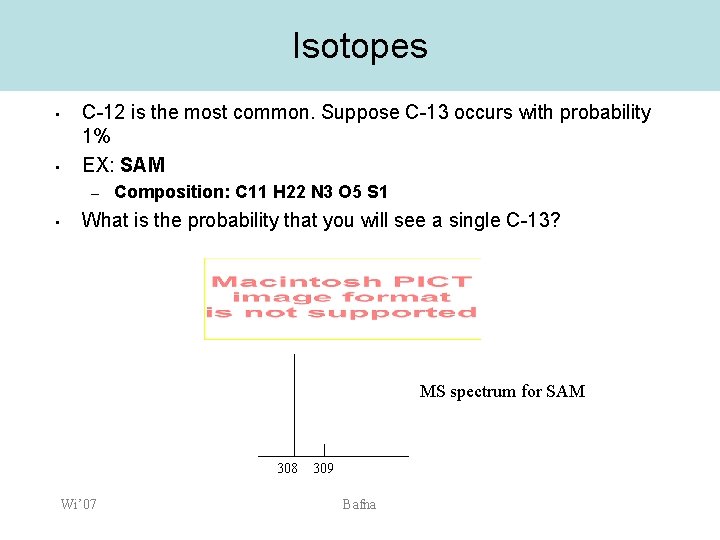
Isotopes • • C-12 is the most common. Suppose C-13 occurs with probability 1% EX: SAM – • Composition: C 11 H 22 N 3 O 5 S 1 What is the probability that you will see a single C-13? MS spectrum for SAM 308 Wi’ 07 309 Bafna

All atoms have isotopes • Isotopes of atoms – – • • • O 16, 18, C-12, 13, S 32, 34…. Each isotope has a frequency of occurrence If a molecule (peptide) has a single copy of C-13, that will shift its peak by 1 Da With multiple copies of a peptide, we have a distribution of intensities over a range of masses (Isotopic profile). How can you compute the isotopic profile of a peak? Wi’ 07 Bafna

Isotope Calculation • Denote: – – – Nc : number of carbon atoms in the peptide Pc : probability of occurrence of C-13 (~1%) Then Nc=50 +1 Nc=200 +1 Wi’ 07 Bafna

Isotope Calculation Example • • • Suppose we consider Nitrogen, and Carbon NN: number of Nitrogen atoms PN: probability of occurrence of N-15 Pr(peak at M) Pr(peak at M+1)? Pr(peak at M+2)? How do we generalize? How can we handle Oxygen (O-16, 18)? Wi’ 07 Bafna

General isotope computation • • • Definition: – Let pi, a be the abundance of the isotope with mass i Da above the least mass – Ex: P 0, C : abundance of C-12, P 2, O: O-18 etc. Characteristic polynomial Prob{M+i}: coefficient of xi in (x) (a binomial convolution) Wi’ 07 Bafna

Quiz • How can you determine the charge on a peptide? § Difference between the first and second isotope peak is 1/Z § Proposal: §Given a mass, predict a composition, and the isotopic profile § Do a ‘goodness of fit’ test to isolate the peaks corresponding to the isotope § Compute the difference Wi’ 07 Bafna

Isotopic Profile Application • • • In Dx. MS, hydrogen atoms are exchanged with deuterium The rate of exchange indicates how buried the peptide is (in folded state) Consider the observed characteristic polynomial of the isotope profile t 1, t 2, at various time points. Then The estimates of p 1, H can be obtained by a deconvolution Such estimates at various time points should give the rate of incorporation of Deuterium, and therefore, the accessibility. Wi’ 07 Bafna

Quantitation via MS Wi’ 07 Bafna

Quantitation • The intensity of the peak depends upon – • • Abundance, ionization potential, substrate etc. Two peptides with the same abundance can have very different intensities. Assumption: relative abundance can be measured by comparing the ratio of a peptide in 2 samples. Wi’ 07 Bafna

Quantitation issues • • • The two samples might be from a complex mixture. How do we identify identical peptides in two samples? In micro-array this is possible because the c. DNA is spotted in a precise location? Can we have a ‘location’ for proteins/peptides MS based quantitation must be coupled with quantitation Wi’ 07 Bafna

2 D Gel based separation • • • Intact proteins are used Iso-electric focusing and Mol Weight used to separate. Intensity of spot is used to measure abundance. Problems: All 3 measurements are not that precise (low reproducibility) Labor intensive Many proteins do not get separated. Wi’ 07 Bafna
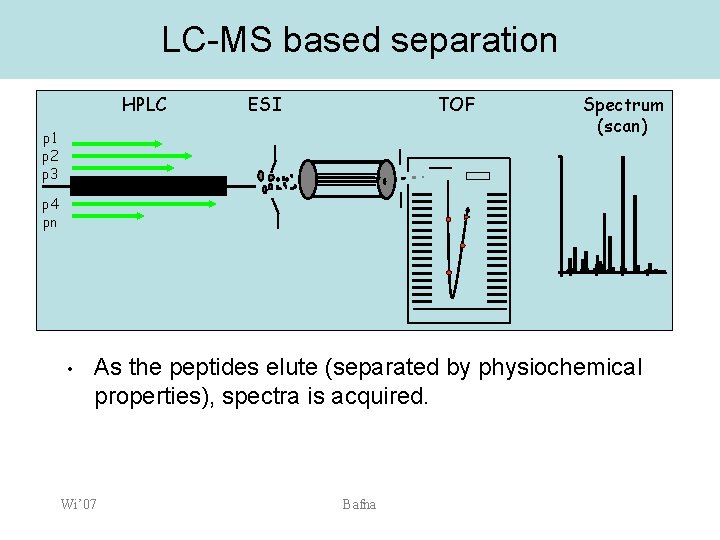
LC-MS based separation HPLC ESI TOF p 1 p 2 p 3 Spectrum (scan) p 4 pn • As the peptides elute (separated by physiochemical properties), spectra is acquired. Wi’ 07 Bafna
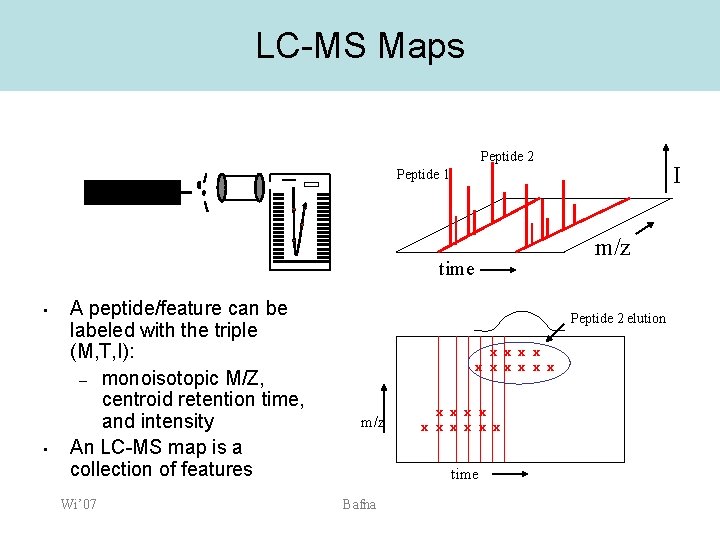
LC-MS Maps Peptide 2 I Peptide 1 m/z time • • A peptide/feature can be labeled with the triple (M, T, I): – monoisotopic M/Z, centroid retention time, and intensity An LC-MS map is a collection of features Wi’ 07 Peptide 2 elution x x x x x m/z x x x x x time Bafna
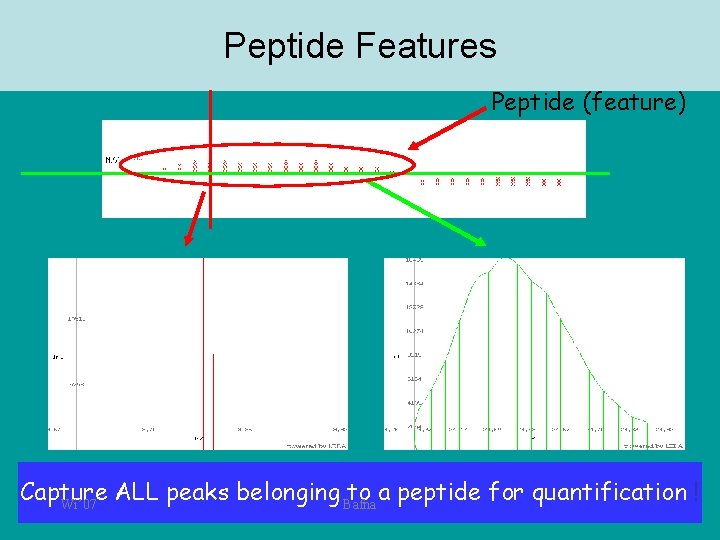
Peptide Features Peptide (feature) Isotope pattern Elution profile Capture ALL peaks belonging Bafna to a peptide for quantification ! Wi’ 07

Relative abundance using MS • • • Differential Isotope labeling (ICAT/SILAC) External standards (AQUA) Direct Map comparison Wi’ 07 Bafna
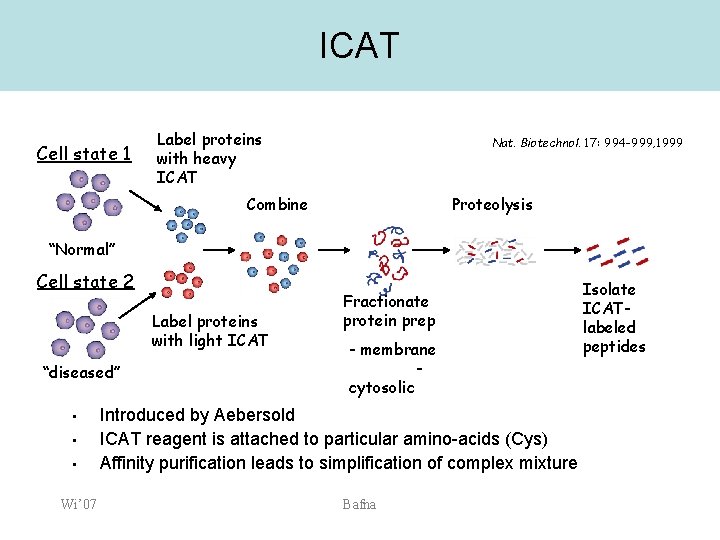
ICAT Cell state 1 Label proteins with heavy ICAT Nat. Biotechnol. 17: 994 -999, 1999 Combine Proteolysis “Normal” Cell state 2 Label proteins with light ICAT “diseased” • • • Wi’ 07 Fractionate protein prep - membrane cytosolic Introduced by Aebersold ICAT reagent is attached to particular amino-acids (Cys) Affinity purification leads to simplification of complex mixture Bafna Isolate ICATlabeled peptides
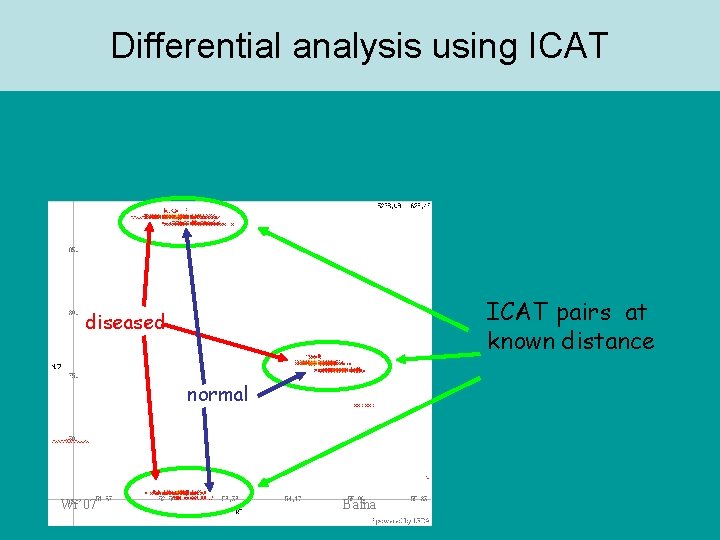
Differential analysis using ICAT pairs at known distance diseased normal Wi’ 07 Bafna

ICAT issues • • • The tag is heavy, and decreases the dynamic range of the measurements. The tag might break off Only Cysteine containing peptides are retrieved Non-specific binding to strepdavidin Wi’ 07 Bafna
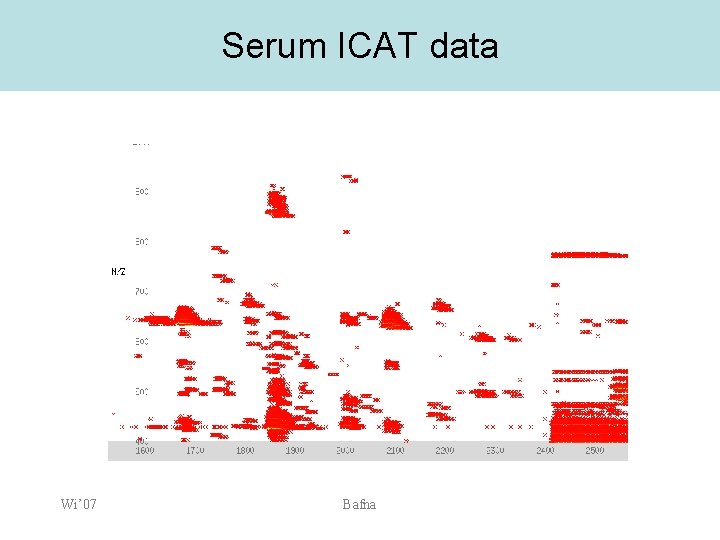
Serum ICAT data Wi’ 07 Bafna
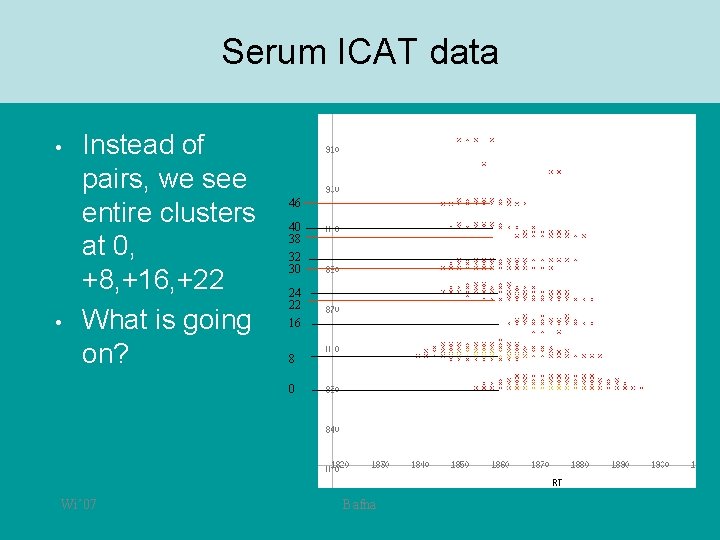
Serum ICAT data • • Instead of pairs, we see entire clusters at 0, +8, +16, +22 What is going on? 46 40 38 32 30 24 22 16 8 0 Wi’ 07 Bafna

SILAC • • • A different isotope labeling strategy Mammalian cells do not ‘manufacture’ all amino-acids. Where do they come from? Labeled amino-acids are added to amino-acid deficient culture, and are incorporated into all proteins as they are synthesized No chemical labeling or affinity purification is performed. Leucine can be used (10% abundance vs 2% for Cys) Wi’ 07 Bafna
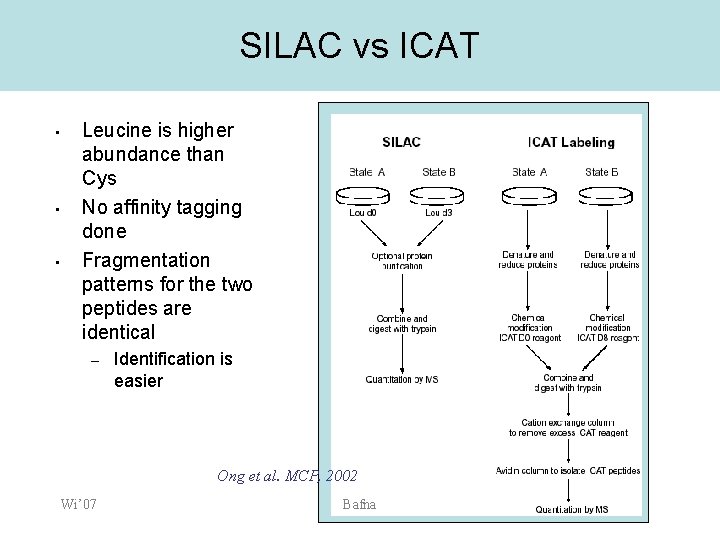
SILAC vs ICAT • • • Leucine is higher abundance than Cys No affinity tagging done Fragmentation patterns for the two peptides are identical – Identification is easier Ong et al. MCP, 2002 Wi’ 07 Bafna
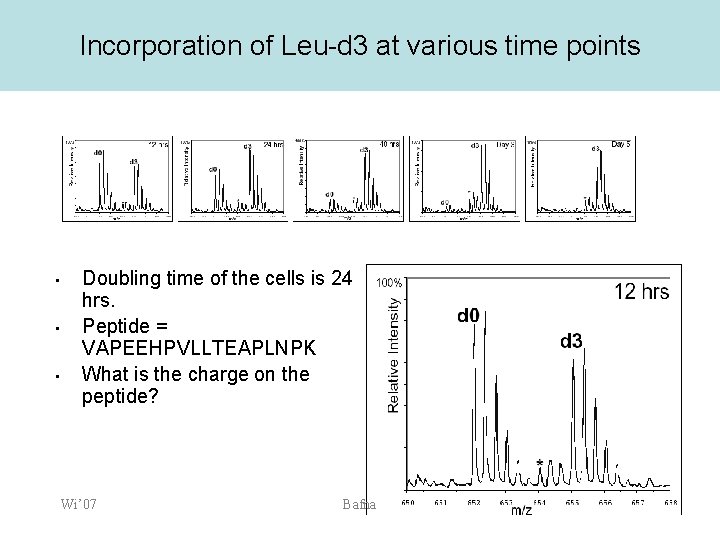
Incorporation of Leu-d 3 at various time points • • • Doubling time of the cells is 24 hrs. Peptide = VAPEEHPVLLTEAPLNPK What is the charge on the peptide? Wi’ 07 Bafna
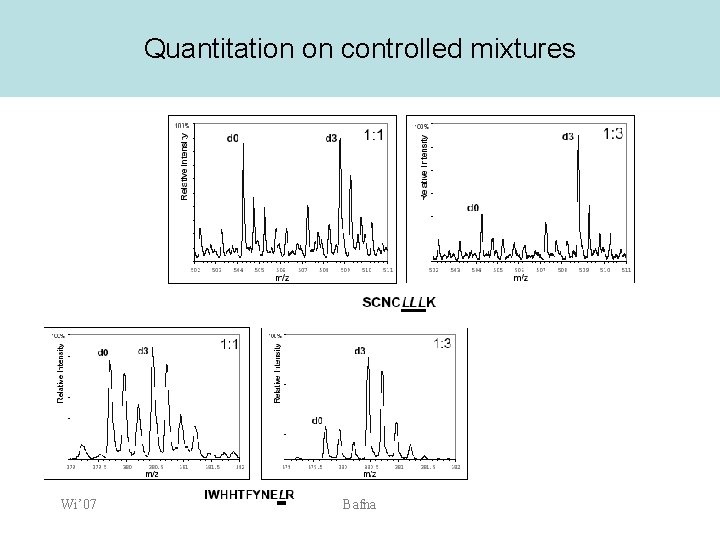
Quantitation on controlled mixtures Wi’ 07 Bafna
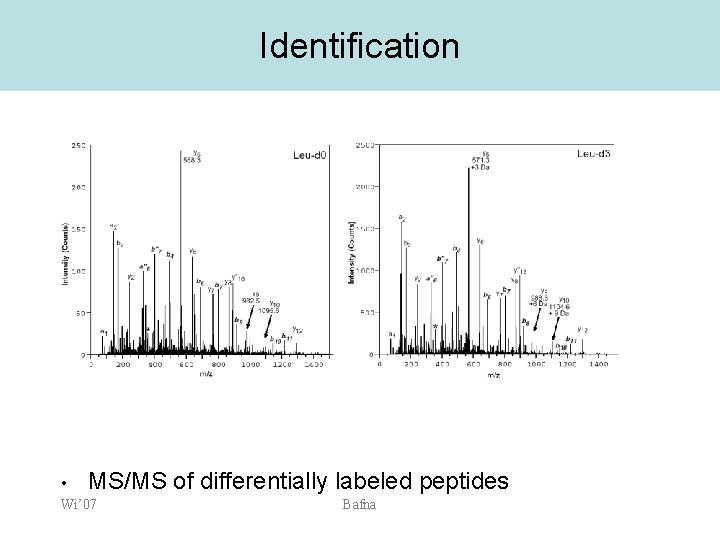
Identification • MS/MS of differentially labeled peptides Wi’ 07 Bafna

Questions • • • The quantitation of peptides also depends upon the amount mixed right in the beginning (what if you mixed more of one sample? ) How can you control for such errors? What happens when you see singletons (unpaired features)? – – – Wi’ 07 In ICAT, it can be a non-Cys peptide (chemical noise), OR Under-expressed Cys-peptide How can you differentiate between the two cases? Bafna
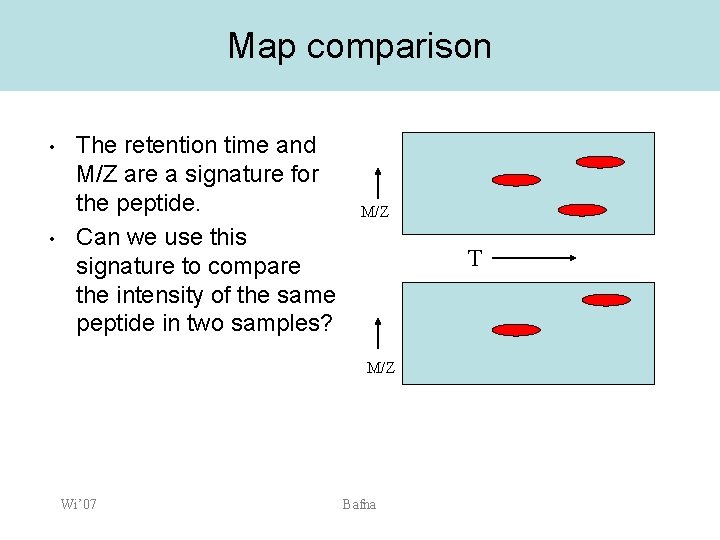
Map comparison • • The retention time and M/Z are a signature for the peptide. Can we use this signature to compare the intensity of the same peptide in two samples? M/Z T M/Z Wi’ 07 Bafna
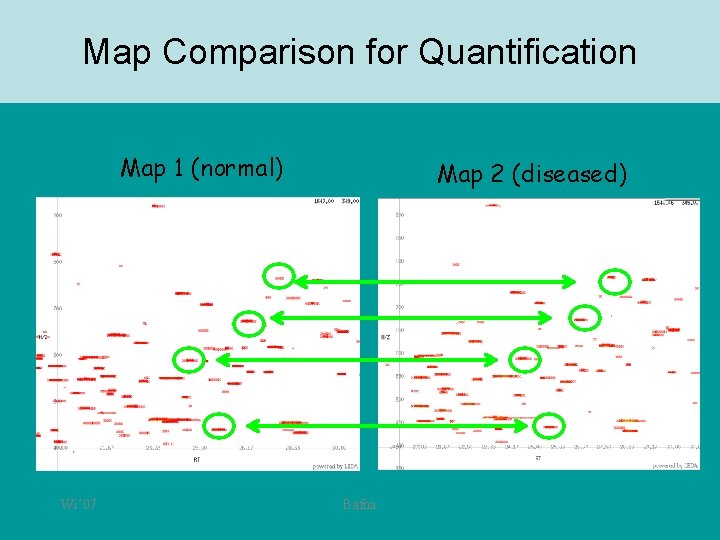
Map Comparison for Quantification Map 1 (normal) Wi’ 07 Map 2 (diseased) Bafna
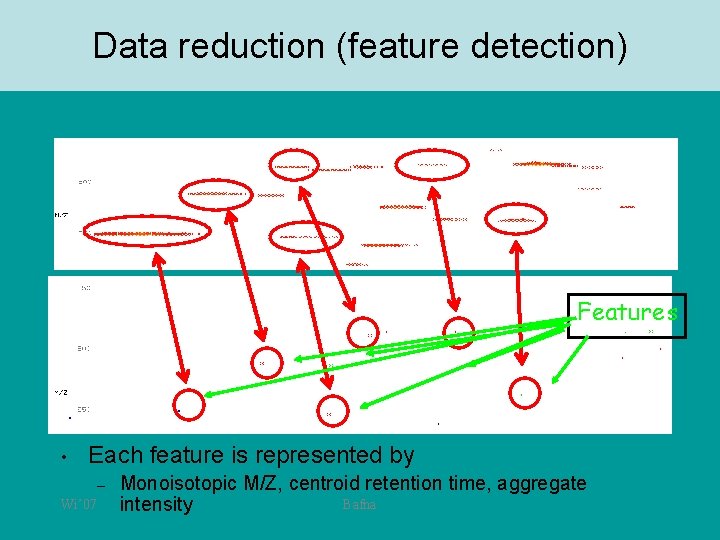
Data reduction (feature detection) Features • Each feature is represented by – Wi’ 07 Monoisotopic M/Z, centroid retention time, aggregate Bafna intensity
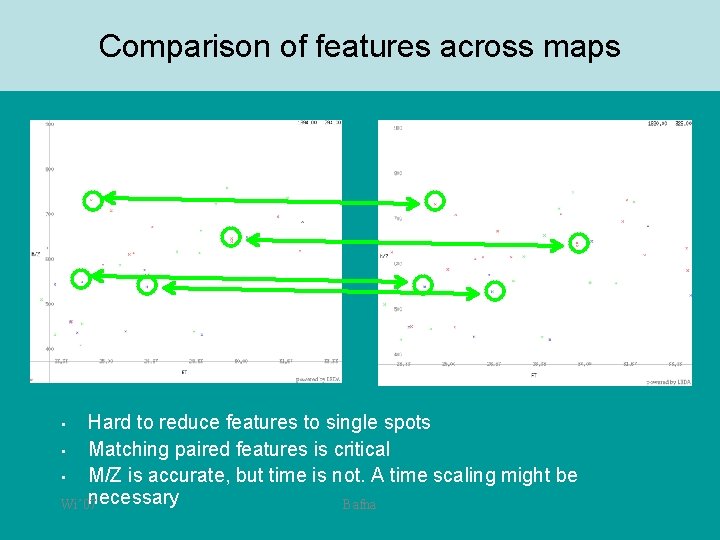
Comparison of features across maps Hard to reduce features to single spots • Matching paired features is critical • M/Z is accurate, but time is not. A time scaling might be necessary Wi’ 07 Bafna •
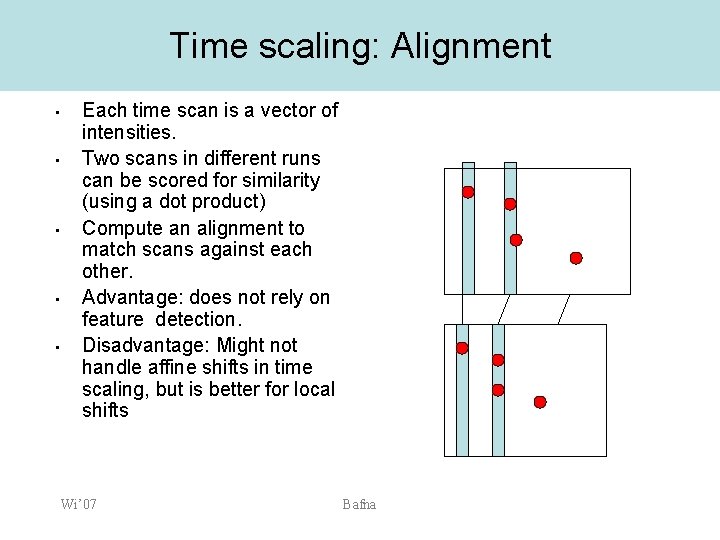
Time scaling: Alignment • • • Each time scan is a vector of intensities. Two scans in different runs can be scored for similarity (using a dot product) Compute an alignment to match scans against each other. Advantage: does not rely on feature detection. Disadvantage: Might not handle affine shifts in time scaling, but is better for local shifts Wi’ 07 Bafna
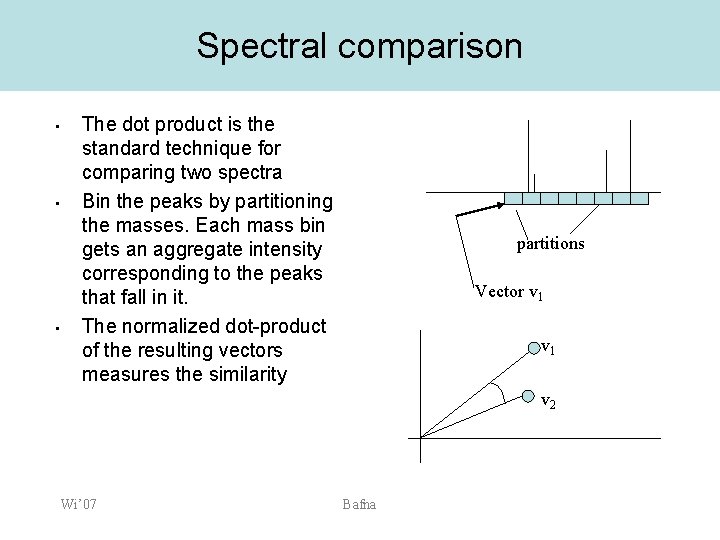
Spectral comparison • • • The dot product is the standard technique for comparing two spectra Bin the peaks by partitioning the masses. Each mass bin gets an aggregate intensity corresponding to the peaks that fall in it. The normalized dot-product of the resulting vectors measures the similarity partitions Vector v 1 v 2 Wi’ 07 Bafna
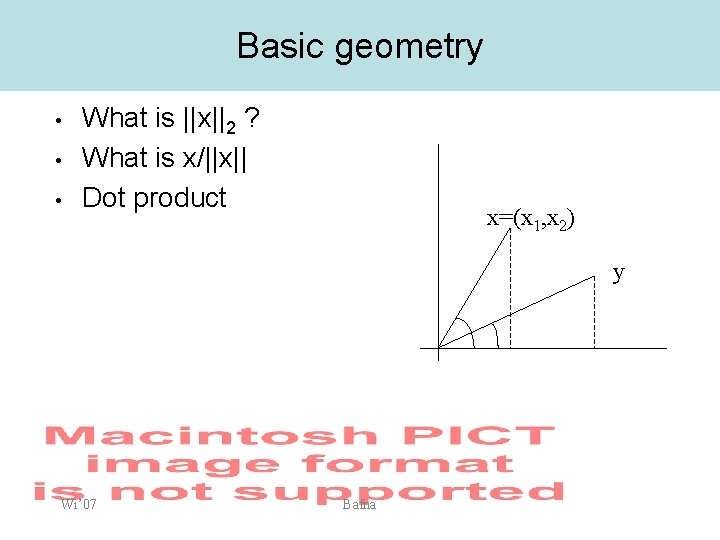
Basic geometry • • • What is ||x||2 ? What is x/||x|| Dot product x=(x 1, x 2) y Wi’ 07 Bafna
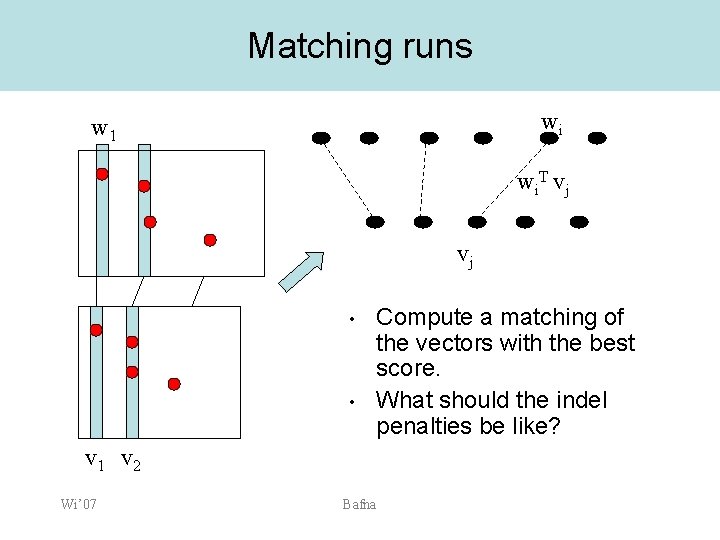
Matching runs wi w 1 wi. T vj vj • • Compute a matching of the vectors with the best score. What should the indel penalties be like? v 1 v 2 Wi’ 07 Bafna
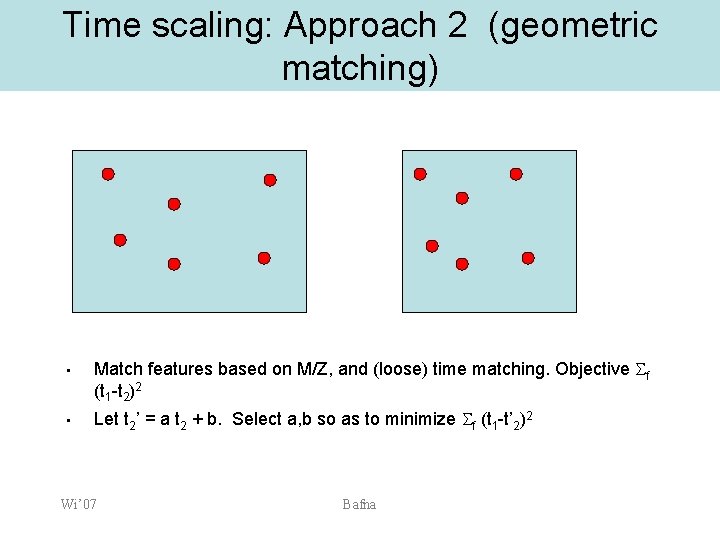
Time scaling: Approach 2 (geometric matching) • • Match features based on M/Z, and (loose) time matching. Objective f (t 1 -t 2)2 Let t 2’ = a t 2 + b. Select a, b so as to minimize f (t 1 -t’ 2)2 Wi’ 07 Bafna
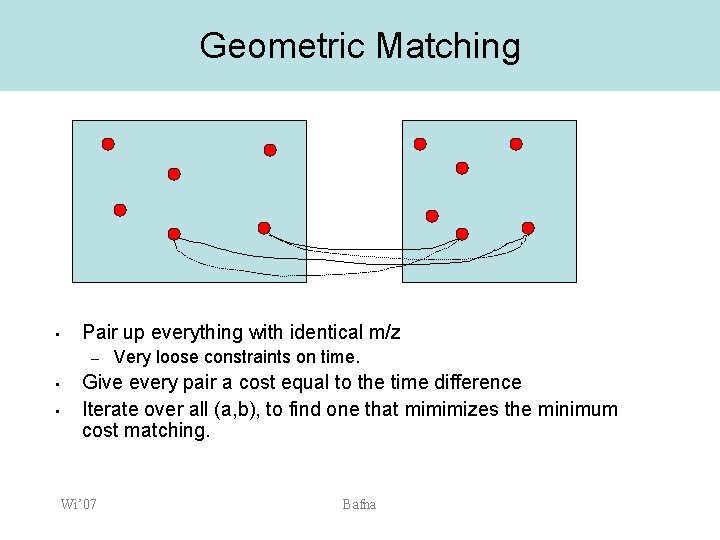
Geometric Matching • Pair up everything with identical m/z – • • Very loose constraints on time. Give every pair a cost equal to the time difference Iterate over all (a, b), to find one that mimimizes the minimum cost matching. Wi’ 07 Bafna
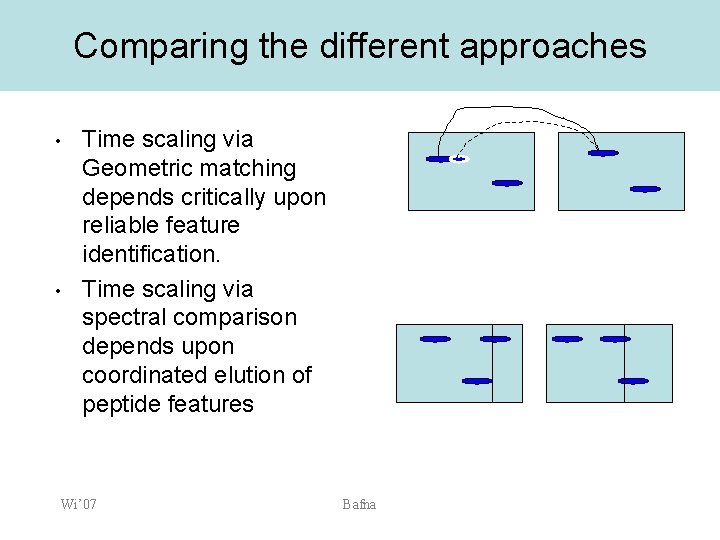
Comparing the different approaches • • Time scaling via Geometric matching depends critically upon reliable feature identification. Time scaling via spectral comparison depends upon coordinated elution of peptide features Wi’ 07 Bafna
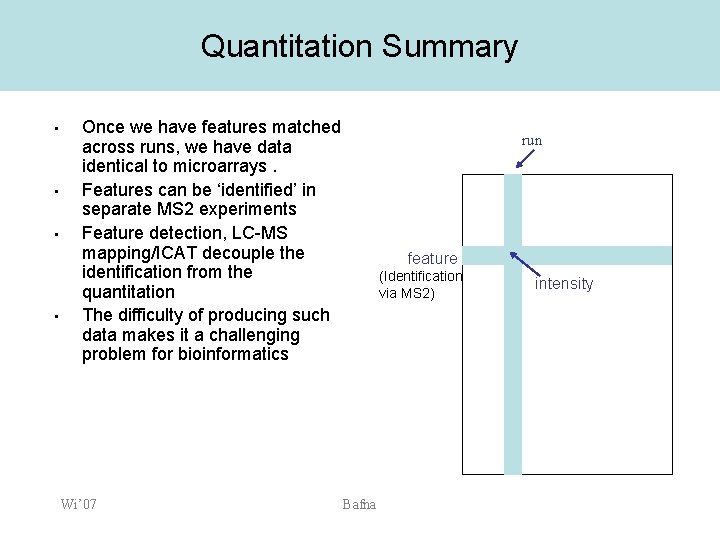
Quantitation Summary • • Once we have features matched across runs, we have data identical to microarrays. Features can be ‘identified’ in separate MS 2 experiments Feature detection, LC-MS mapping/ICAT decouple the identification from the quantitation The difficulty of producing such data makes it a challenging problem for bioinformatics Wi’ 07 run feature (Identification via MS 2) Bafna intensity

Other applications of MS Wi’ 07 Bafna

MS application: Protein-protein interaction • • Proteins combine to form functional complexes. An antibody is a special kind of protein that can recognize a specific protein Use an antibody to recognize a protein in a complex. Isolate & Purify the complex that binds to the antibody. Identify all the proteins in the complex via mass spectrometry. Wi’ 07 Bafna

MS application: Protein Structure • • Use chemical cross-linkers to link spatially proximal residues. Denature and digest the protein. Identify the cross-linked peptides. This provides extra structural constraints which help predict structure. Wi’ 07 Bafna
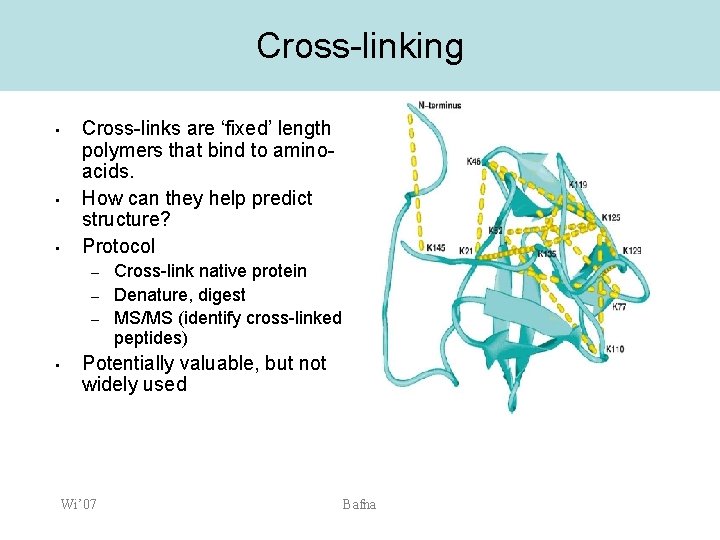
Cross-linking • • • Cross-links are ‘fixed’ length polymers that bind to aminoacids. How can they help predict structure? Protocol – – – • Cross-link native protein Denature, digest MS/MS (identify cross-linked peptides) Potentially valuable, but not widely used Wi’ 07 Bafna

Identifying Cross-linked peptides • • • Identify all peptide pairs, whose mass explains the parent mass. Given a list of peptide pairs, find the pair, and the linked position that best explains the MS 2 data. What is the number of possible candidate pairs. Fragmentation in the presence of linkers is poorly understood How do you separate crosslinked peptides from singly linked, and non-cross-linked peptides? Wi’ 07 Bafna

Overlap peptides and shotgun based identification Wi’ 07 Bafna

Motivation Database search of MS/MS spectra works well whenever the protein sequence is known and the protein is not modified/mutated BUT: – Not all protein sequences are available • • • – Wi’ 07 Examples include proteins from snake and scorpion venom Integrilin, a successful blood clot prevention drug distributed by Millenium, was derived from rattlesnake venom Sequences are still determined by Edman sequencing Database search is likely to fail if the analyzed protein contains unexpected post-translational modifications or mutations. Bafna
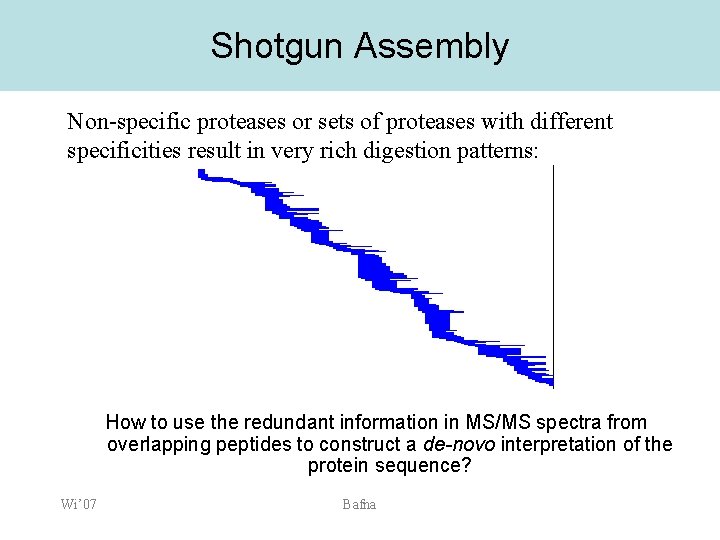
Shotgun Assembly Non-specific proteases or sets of proteases with different specificities result in very rich digestion patterns: How to use the redundant information in MS/MS spectra from overlapping peptides to construct a de-novo interpretation of the protein sequence? Wi’ 07 Bafna

Unanticipated modifications Wi’ 07 Bafna

MS-Alignment Tsur et al. ’ Nat. Bio. 2005 • • Wi’ 07 Dynamic Programming can be used to capture mass-offsets (putative PTMs). In large data-sets, true PTMs should be over-represented. Bafna

PTM Frequency Matrix 50, 000 spectra from a sample of IKKb were searched in blind mode, and identifications with pvalue <0. 05 were retained Cell shading indicates the number of annotations with modification ( , a) Wi’ 07 Bafna

PTM Frequency Matrix Methylation Oxidation Sodium Wi’ 07 Dimethylation Bafna Double oxidation

PTM selection: Output Wi’ 07 Bafna

Filtering -correct annotations M+17 (from oxidized methionine, incorrect mass) A+14 (from methylated lysine, incorrect placement) Wi’ 07 Bafna

Overlapping peptides Wi’ 07 Bafna

Features for robust PTM identification • A number of features are used to validate mass-shifts obtained from MS-alignment, including – – – • • evidence from overlapping peptides, number of spectra. Delta scores from other possibilities, Evidence from multiple sites, de-novo identifications etc. These features are used to train an SVM. Validation is done at spectrum, peptide, and site levels Final searches performed using a fixed false discovery rates on a random database. Wi’ 07 Bafna
Blind Search of a lens data-set Human lenses are a rich source of modified proteins: – Lens proteins do not turnover, but accumulate modifications over time – Hundreds of papers in the last 20 years reporting crystallin modifications – Yates‘ lab pioneered use of various proteases for high-throughput PTM validation/discovery in lenses (Mac. Coss et al, 2003) – Sample from a 93 -year old patient – Wilmarth et al. (Jnl. Prot. Res. , 06) Wi’ 07 Bafna
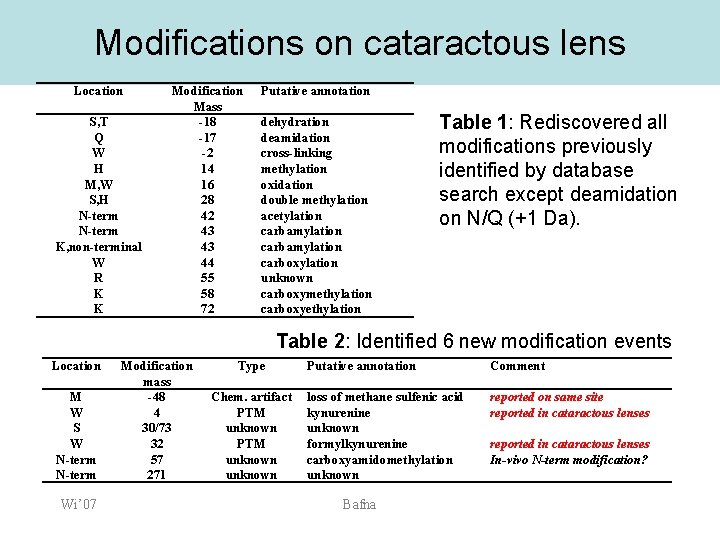
Modifications on cataractous lens Location S, T Q W H M, W S, H N-term K, non-terminal W R K K Modification Mass -18 -17 -2 14 16 28 42 43 43 44 55 58 72 Putative annotation dehydration deamidation cross-linking methylation oxidation double methylation acetylation carbamylation carboxylation unknown carboxymethylation carboxyethylation Table 1: Rediscovered all modifications previously identified by database search except deamidation on N/Q (+1 Da). Table 2: Identified 6 new modification events Location M W S W N-term Wi’ 07 Modification mass -48 4 30/73 32 57 271 Type Chem. artifact PTM unknown Putative annotation Comment loss of methane sulfenic acid kynurenine unknown formylkynurenine carboxyamidomethylation unknown reported on same site reported in cataractous lenses Bafna reported in cataractous lenses In-vivo N-term modification?

Lens: Unknown modifications • • Unknown modification R+55 was found on both data-sets, and assigned to overlapping sets of sites. Some evidence for Q+161 (glycosylation? ) in a few spectra Wi’ 07 Bafna
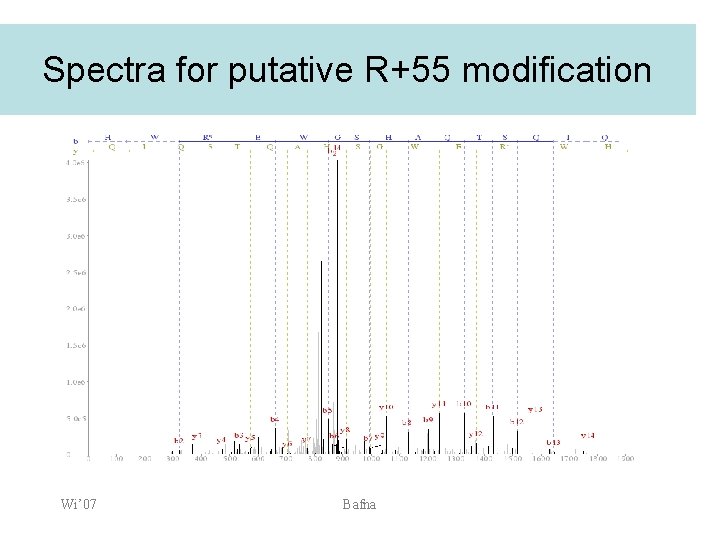
Spectra for putative R+55 modification Wi’ 07 Bafna

2. Selecting modifications (summary) • A ‘strength in numbers’ approach: The more spectra, the better. – • • A recent unpublished search with ~18 M spectra reveals ~500 modified sites, with a 2% FDR Overlapping peptides are strong evidence (incorrect matches unlikely to overlap) These and other features are being used in an automated tool for identifying modified sites. Wi’ 07 Bafna
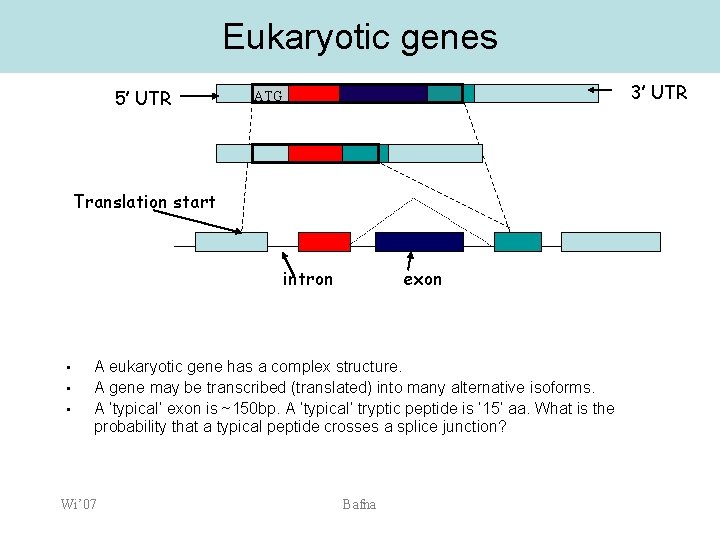
Eukaryotic genes 5’ UTR 3’ UTR ATG Translation start intron • • • exon A eukaryotic gene has a complex structure. A gene may be transcribed (translated) into many alternative isoforms. A ‘typical’ exon is ~150 bp. A ‘typical’ tryptic peptide is ‘ 15’ aa. What is the probability that a typical peptide crosses a splice junction? Wi’ 07 Bafna
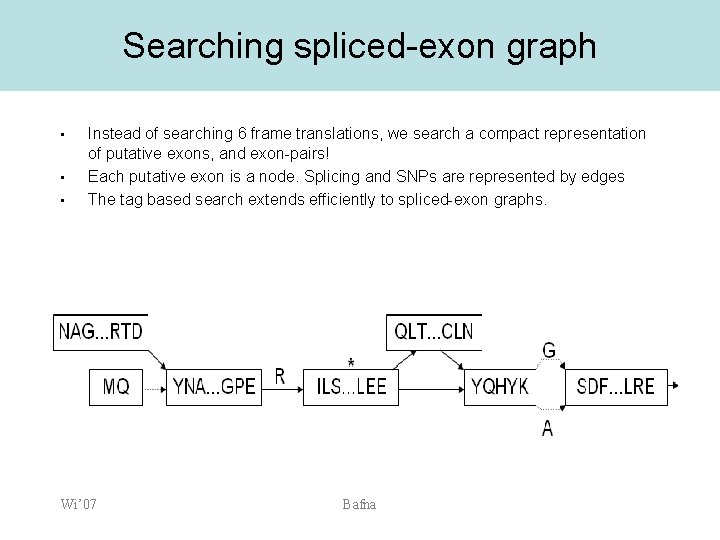
Searching spliced-exon graph • • • Instead of searching 6 frame translations, we search a compact representation of putative exons, and exon-pairs! Each putative exon is a node. Splicing and SNPs are represented by edges The tag based search extends efficiently to spliced-exon graphs. Wi’ 07 Bafna
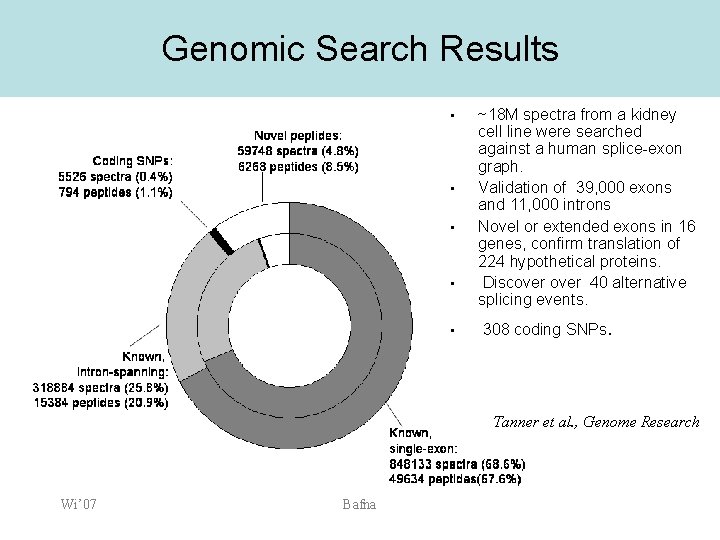
Genomic Search Results • • • ~18 M spectra from a kidney cell line were searched against a human splice-exon graph. Validation of 39, 000 exons and 11, 000 introns Novel or extended exons in 16 genes, confirm translation of 224 hypothetical proteins. Discover 40 alternative splicing events. 308 coding SNPs. Tanner et al. , Genome Research Wi’ 07 Bafna
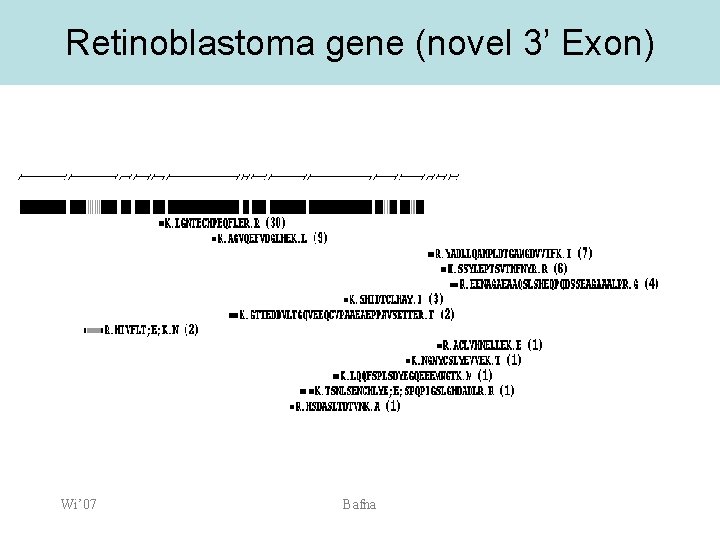
Retinoblastoma gene (novel 3’ Exon) Wi’ 07 Bafna

EX: Novel Exons intron Wi’ 07 Bafna

Validating hypothetical protein Wi’ 07 Bafna

Conclusion • • • Key technology for proteomics Leading technology for protein identification, PT modifications, and protein level quantitation Applications to protein structure, interactions, pathways and other proteomic problems Wi’ 07 Bafna
- Slides: 148