Proteomics Mass Spectrometry Nathan Edwards Center for Bioinformatics

Proteomics & Mass Spectrometry Nathan Edwards Center for Bioinformatics and Computational Biology

Outline • Proteomics • Mass Spectrometry • Protein Identification • Peptide Mass Fingerprint • Tandem Mass Spectrometry 2

Proteomics • Proteins are the machines that drive much of biology • Genes are merely the recipe • The direct characterization of a sample’s proteins en masse. • What proteins are present? • How much of each protein is present? 3

Systems Biology • Establish relationships by • Choosing related samples, • Global characterization, and • Comparison. Gene / Transcript / Protein Measurement Discrete (DNA) Continuous Predetermined Unknown Genotyping Sequencing Gene Expression Proteomics 4

Samples • Healthy / Diseased • Cancerous / Benign • Drug resistant / Drug susceptible • Bound / Unbound • Tissue specific • Cellular location specific • Mitochondria, Membrane 5

2 D Gel-Electrophoresis • Protein separation • Molecular weight (MW) • Isoelectric point (p. I) • Staining • Birds-eye view of protein abundance 6
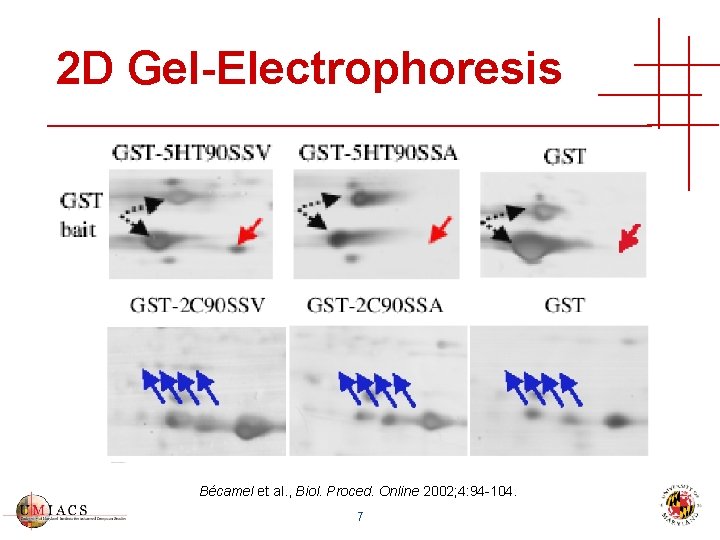
2 D Gel-Electrophoresis Bécamel et al. , Biol. Proced. Online 2002; 4: 94 -104. 7

Paradigm Shift • Traditional protein chemistry assay methods struggle to establish identity. • Identity requires: • Specificity of measurement (Precision) • Mass spectrometry • A reference for comparison (Measurement → Identity) • Protein sequence databases 8
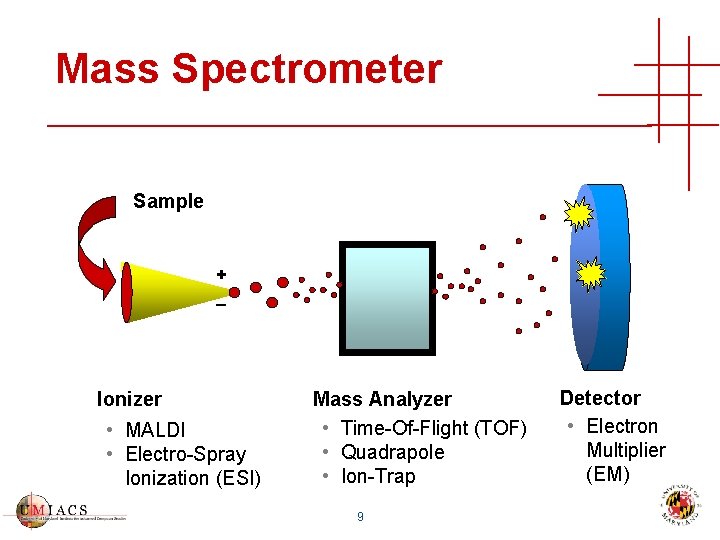
Mass Spectrometer Sample + _ Ionizer • MALDI • Electro-Spray Ionization (ESI) Mass Analyzer • Time-Of-Flight (TOF) • Quadrapole • Ion-Trap 9 Detector • Electron Multiplier (EM)
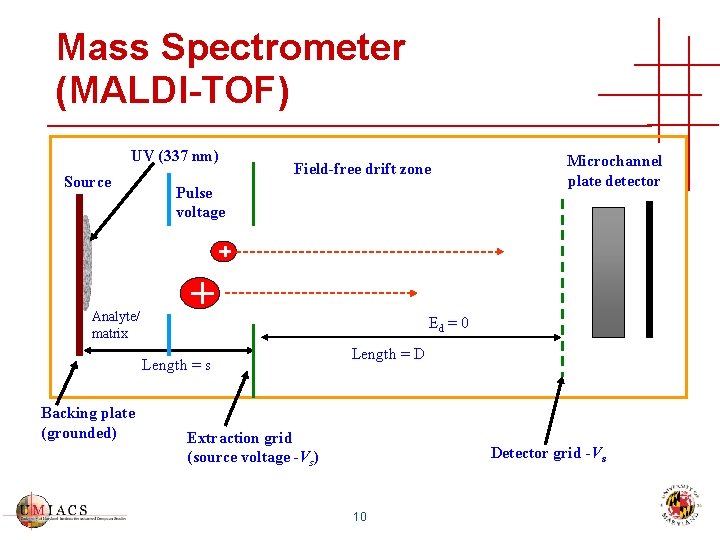
Mass Spectrometer (MALDI-TOF) UV (337 nm) Source Field-free drift zone Pulse voltage Analyte/ matrix Ed = 0 Length = s Backing plate (grounded) Microchannel plate detector Length = D Extraction grid (source voltage -Vs) Detector grid -Vs 10
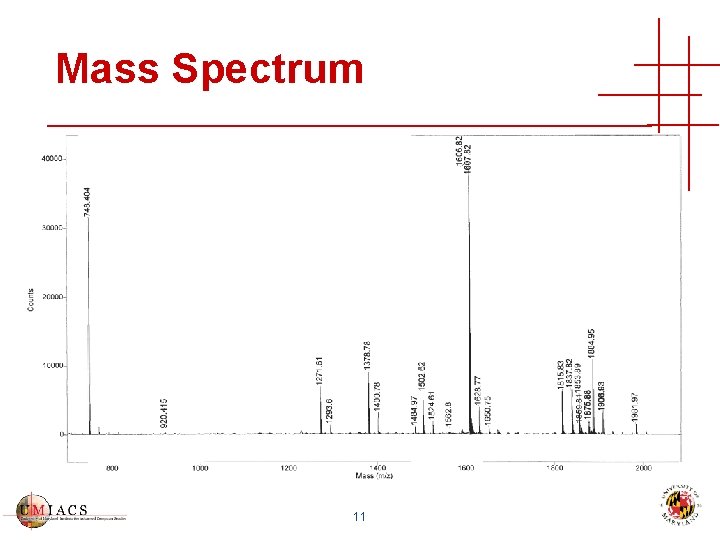
Mass Spectrum 11

Mass is fundamental 12
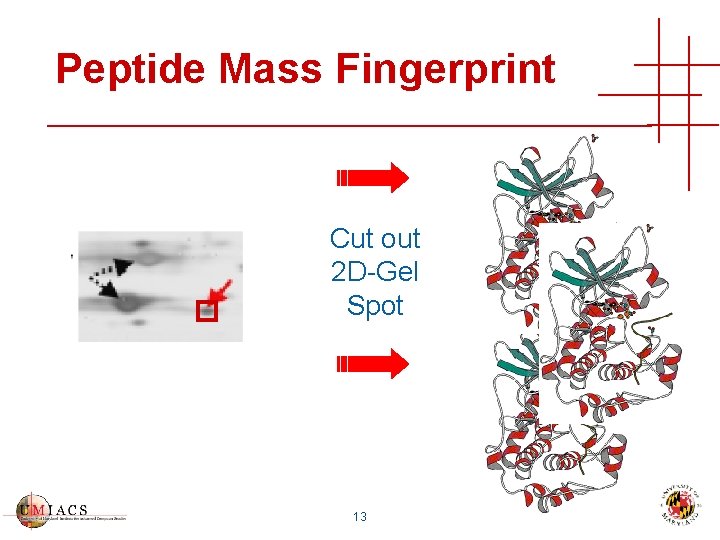
Peptide Mass Fingerprint Cut out 2 D-Gel Spot 13
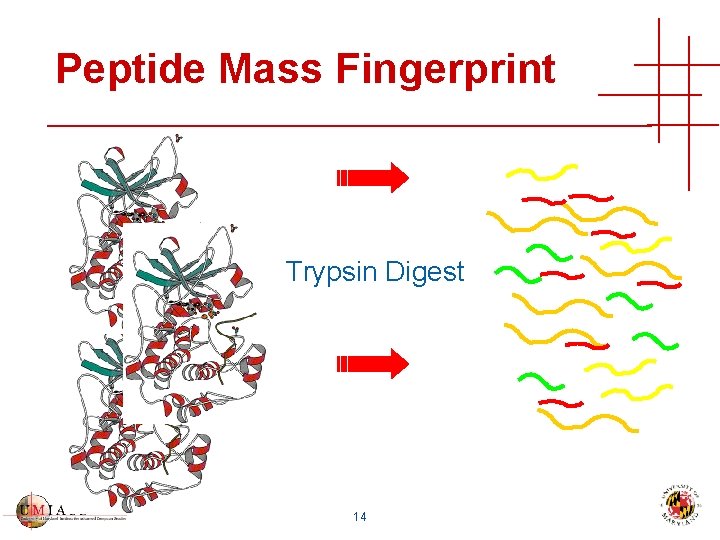
Peptide Mass Fingerprint Trypsin Digest 14
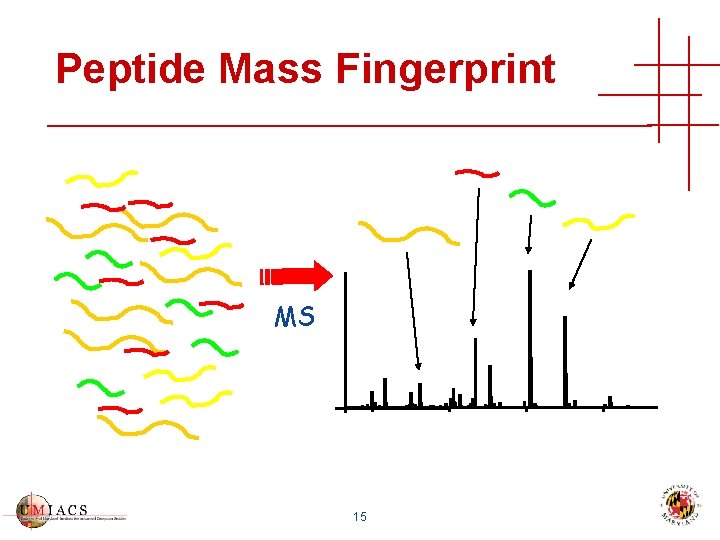
Peptide Mass Fingerprint MS 15
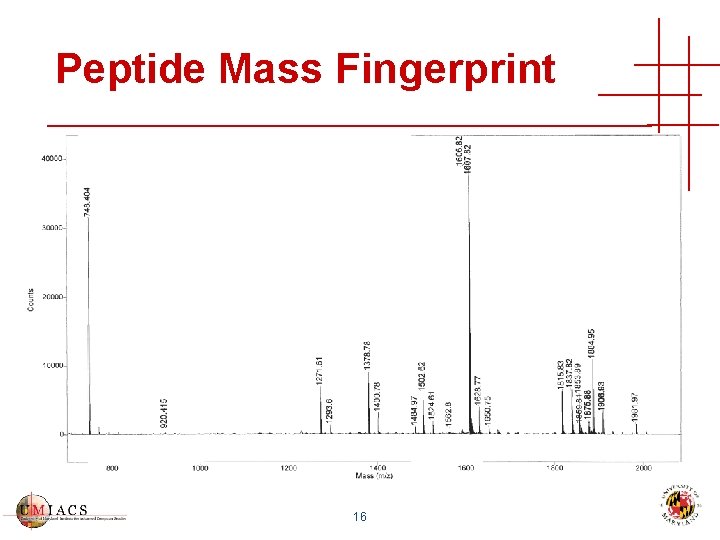
Peptide Mass Fingerprint 16

Peptide Mass Fingerprint • Trypsin: digestion enzyme • Highly specific • Cuts after K & R except if followed by P • Protein sequence from sequence database • In silico digest • Mass computation • For each protein sequence in turn: • Compare computer generated masses with observed spectrum 17

Protein Sequence • Myoglobin - Plains zebra GLSDGEWQQV RLFTGHPETL LKKHGTVVLT QSHATKHKIP GDFGADAQGA FQG LNVWGKVEAD EKFDKFKHLK ALGGILKKKG IKYLEFISDA MTKALELFRN 18 IAGHGQEVLI TEAEMKASED HHEAELKPLA IIHVLHSKHP DIAAKYKELG

Protein Sequence • Myoglobin - Plains zebra GLSDGEWQQV RLFTGHPETL LKKHGTVVLT QSHATKHKIP GDFGADAQGA FQG LNVWGKVEAD EKFDKFKHLK ALGGILKKKG IKYLEFISDA MTKALELFRN 19 IAGHGQEVLI TEAEMKASED HHEAELKPLA IIHVLHSKHP DIAAKYKELG

Peptide Masses 1811. 90 1606. 85 1271. 66 1378. 83 1982. 05 1853. 95 1884. 01 1502. 66 748. 43 GLSDGEWQQVLNVWGK VEADIAGHGQEVLIR LFTGHPETLEK HGTVVLTALGGILK KGHHEAELKPLAQSHATK YLEFISDAIIHVLHSK HPGDFGADAQGAMTK ALELFR 20
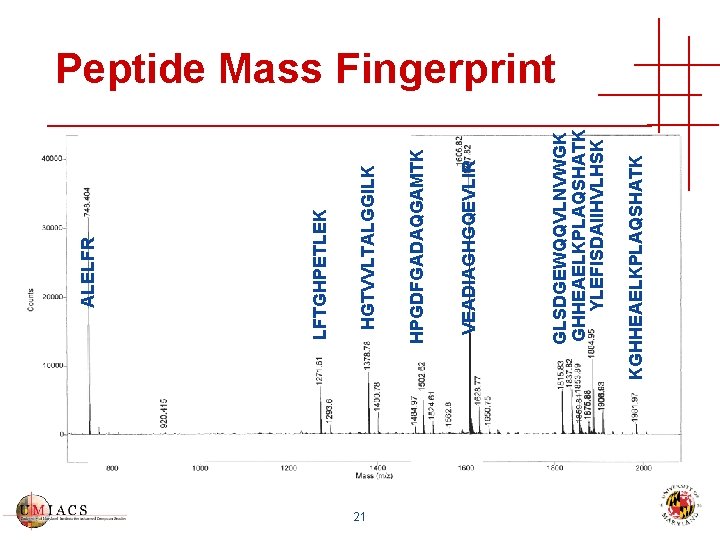
21 KGHHEAELKPLAQSHATK GLSDGEWQQVLNVWGK GHHEAELKPLAQSHATK YLEFISDAIIHVLHSK VEADIAGHGQEVLIR HPGDFGADAQGAMTK HGTVVLTALGGILK LFTGHPETLEK ALELFR Peptide Mass Fingerprint

Mass Spectrometry • Strengths • Precise molecular weight • Fragmentation • Automated • Weaknesses • Best for a few molecules at a time • Best for small molecules • Mass-to-charge ratio, not mass • Intensity ≠ Abundance 22
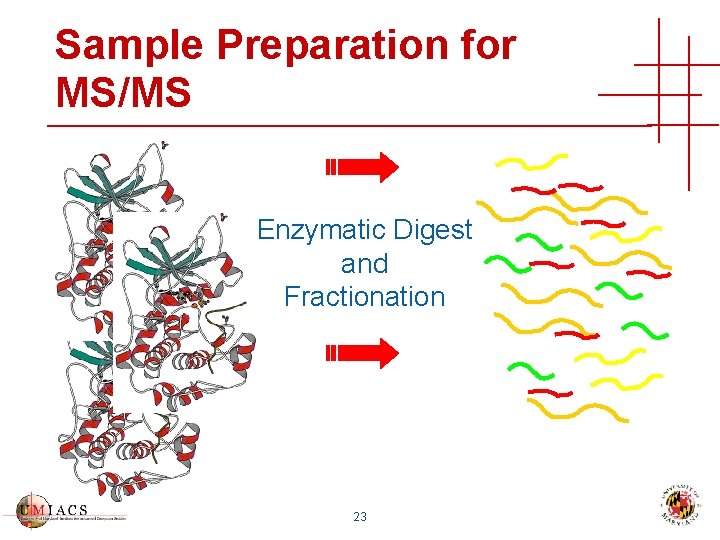
Sample Preparation for MS/MS Enzymatic Digest and Fractionation 23
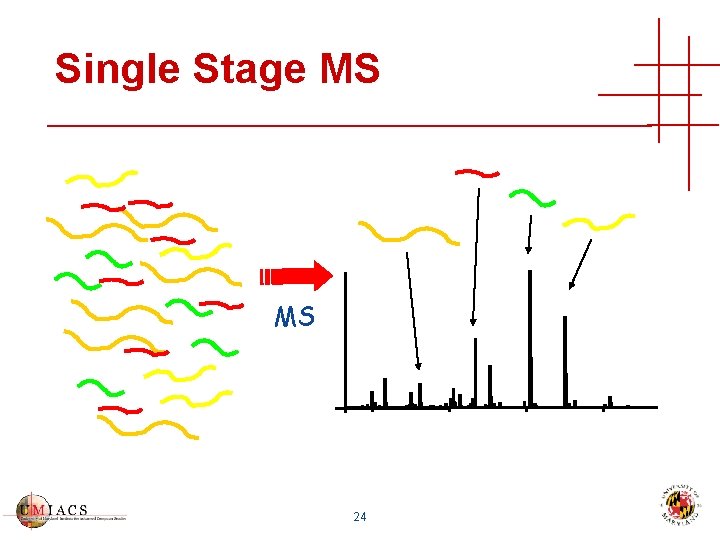
Single Stage MS MS 24
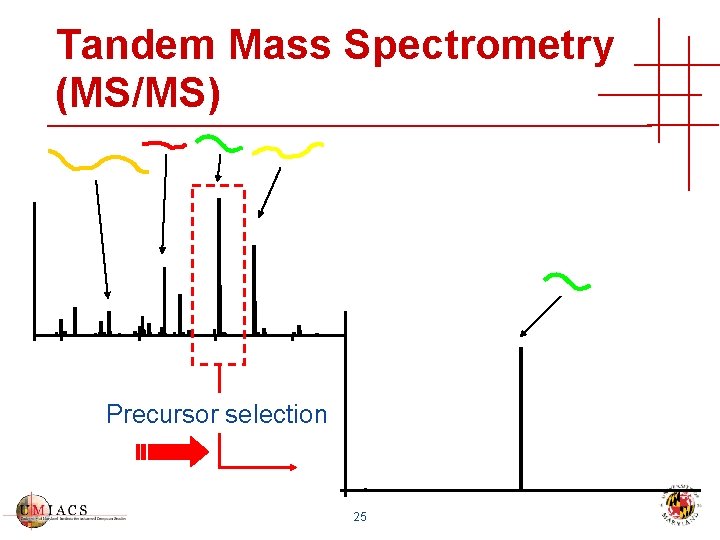
Tandem Mass Spectrometry (MS/MS) Precursor selection 25
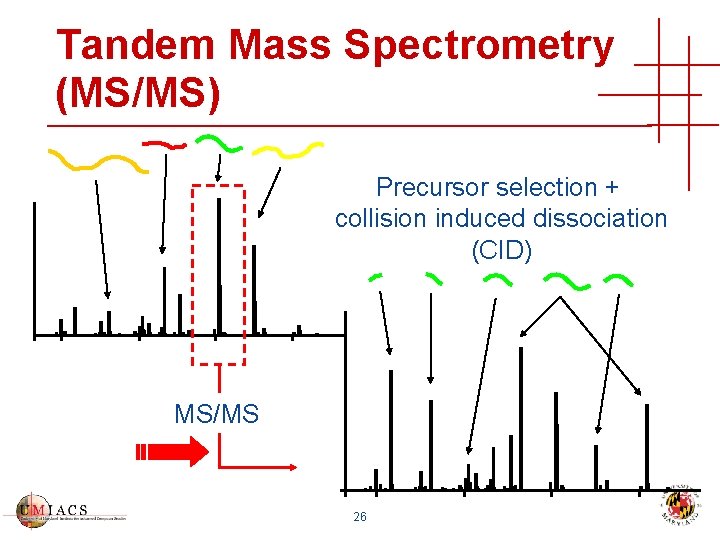
Tandem Mass Spectrometry (MS/MS) Precursor selection + collision induced dissociation (CID) MS/MS 26
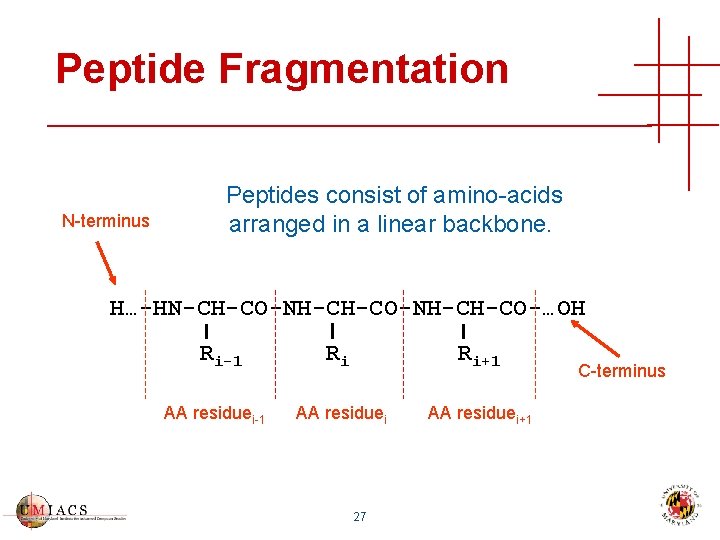
Peptide Fragmentation N-terminus Peptides consist of amino-acids arranged in a linear backbone. H…-HN-CH-CO-NH-CH-CO-…OH Ri-1 AA residuei-1 Ri Ri+1 AA residuei+1 27 C-terminus
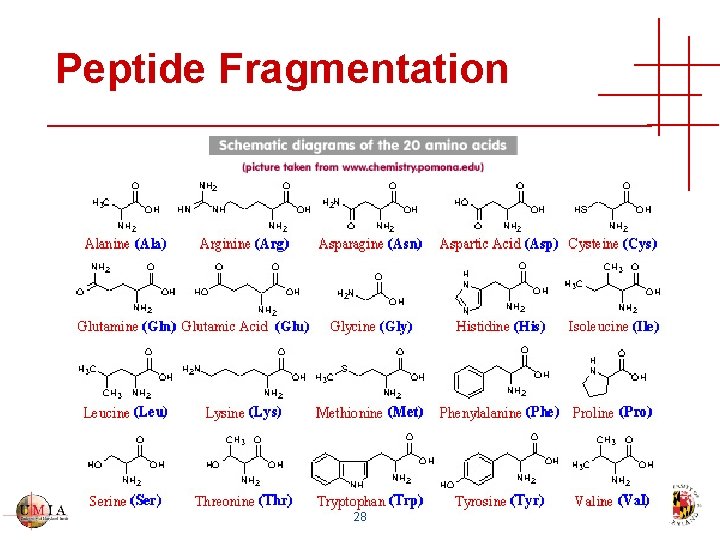
Peptide Fragmentation 28
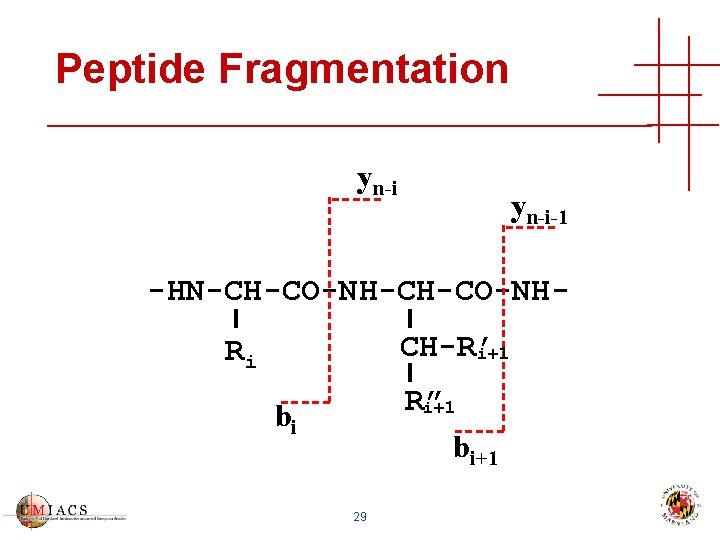
Peptide Fragmentation yn-i-1 -HN-CH-CO-NHCH-R’ i+1 Ri R” i+1 bi 29
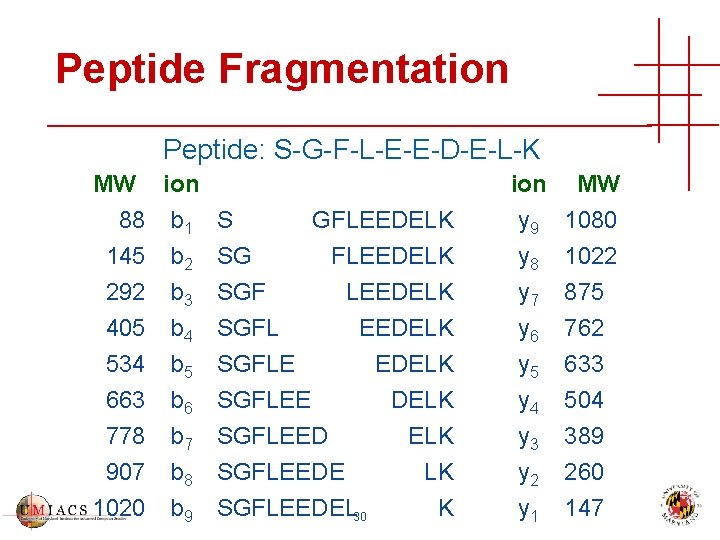
Peptide Fragmentation Peptide: S-G-F-L-E-E-D-E-L-K MW ion 88 b 1 S 145 b 2 SG 292 b 3 SGF 405 534 663 778 907 1020 b 4 b 5 b 6 b 7 b 8 b 9 GFLEEDELK SGFL EEDELK SGFLEE DELK SGFLEEDE LK SGFLEEDEL 30 K ion MW y 9 1080 y 8 1022 y 7 875 y 6 y 5 y 4 y 3 y 2 y 1 762 633 504 389 260 147
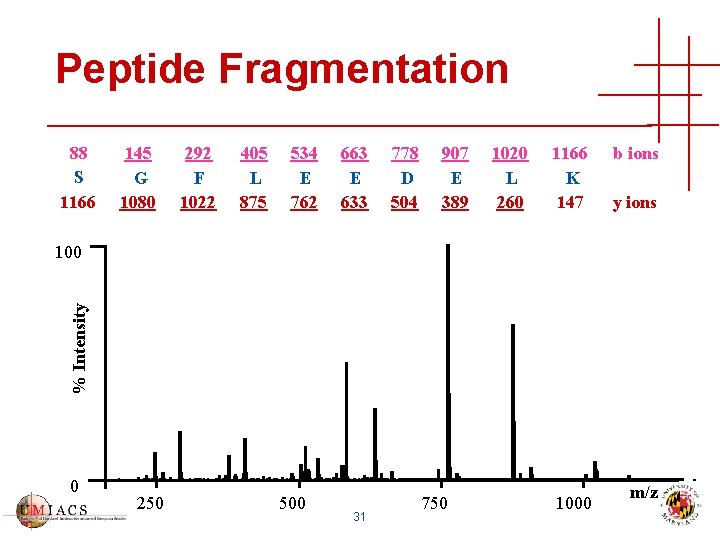
Peptide Fragmentation 88 S 1166 145 G 1080 292 F 1022 405 L 875 534 E 762 663 E 633 778 D 504 907 E 389 1020 L 260 1166 K 147 b ions y ions % Intensity 100 0 250 500 31 750 1000 m/z
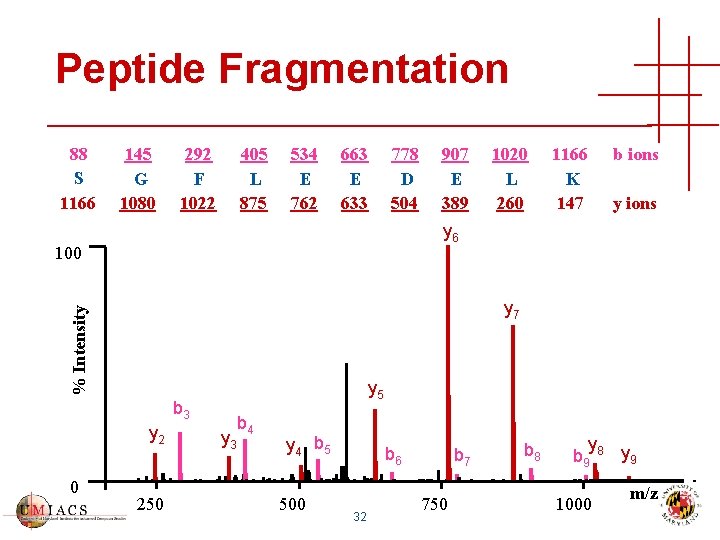
Peptide Fragmentation 88 S 1166 145 G 1080 292 F 1022 405 L 875 534 E 762 663 E 633 778 D 504 1020 L 260 1166 K 147 b ions y 6 100 % Intensity y 7 y 5 b 3 y 2 0 907 E 389 250 y 3 b 4 y 4 b 5 500 b 6 32 b 7 750 b 8 y b 9 8 y 9 1000 m/z

Peptide Identification Given: • The mass of the precursor ion, and • The MS/MS spectrum Output: • The amino-acid sequence of the peptide 33

Peptide Identification Two paradigms: • De novo interpretation • Sequence database search 34
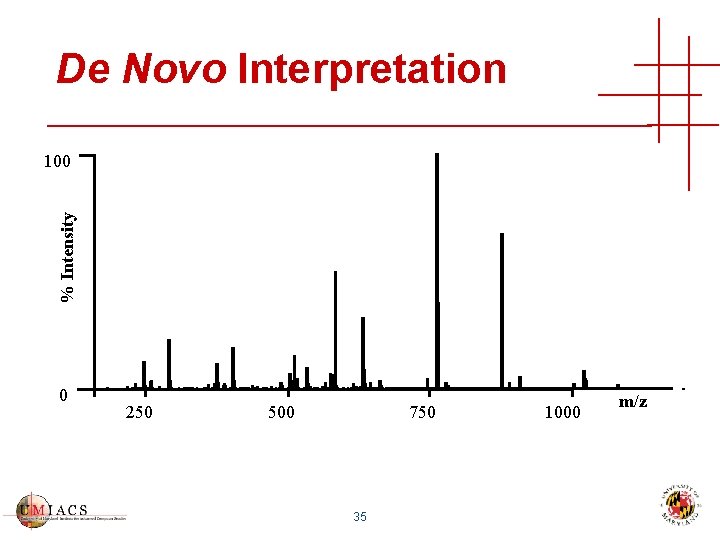
De Novo Interpretation % Intensity 100 0 250 500 750 35 1000 m/z
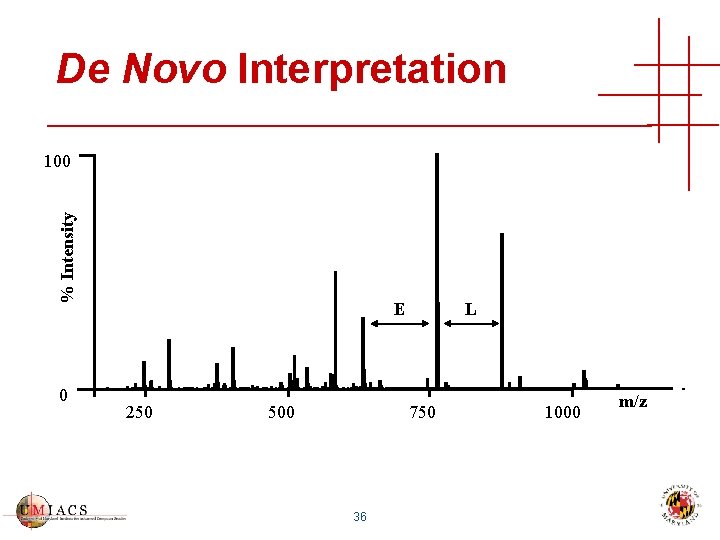
De Novo Interpretation % Intensity 100 0 E 250 500 L 750 36 1000 m/z
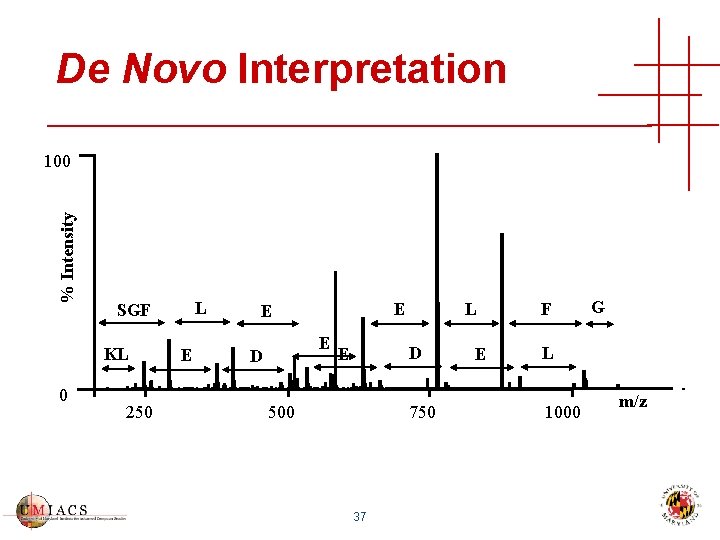
De Novo Interpretation % Intensity 100 KL 0 L SGF 250 E E D L D E 500 750 37 E F G L 1000 m/z
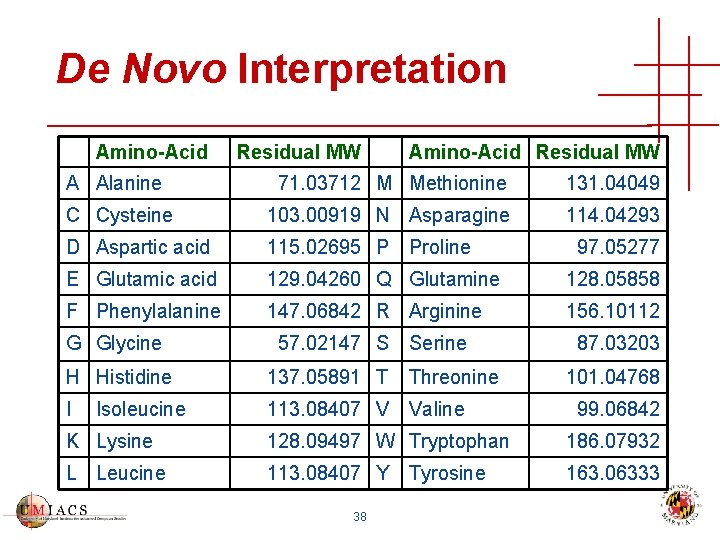
De Novo Interpretation Amino-Acid Residual MW A Alanine 71. 03712 M Methionine 131. 04049 C Cysteine 103. 00919 N Asparagine 114. 04293 D Aspartic acid 115. 02695 P E Glutamic acid 129. 04260 Q Glutamine 128. 05858 F Phenylalanine 147. 06842 R Arginine 156. 10112 G Glycine 57. 02147 S Proline Serine H Histidine 137. 05891 T Threonine I 113. 08407 V Valine Isoleucine 97. 05277 87. 03203 101. 04768 99. 06842 K Lysine 128. 09497 W Tryptophan 186. 07932 L Leucine 113. 08407 Y 163. 06333 38 Tyrosine
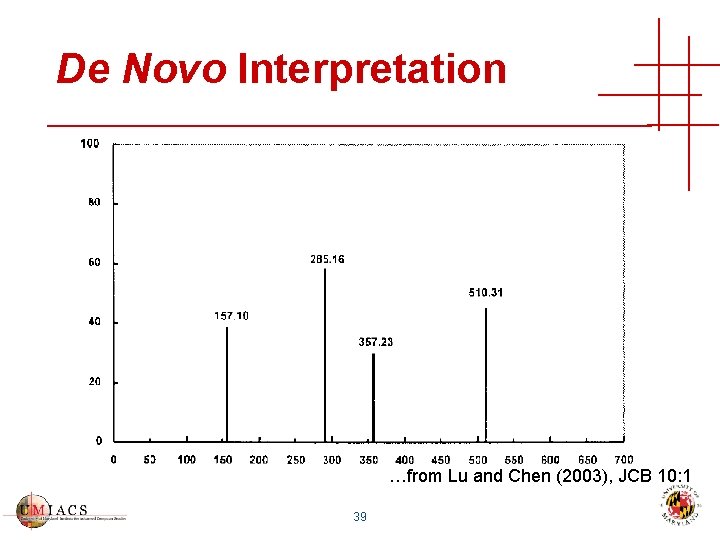
De Novo Interpretation …from Lu and Chen (2003), JCB 10: 1 39
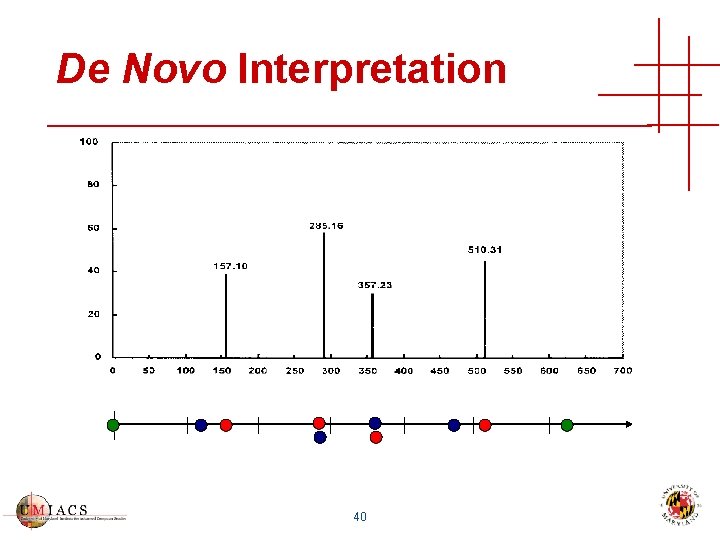
De Novo Interpretation 40
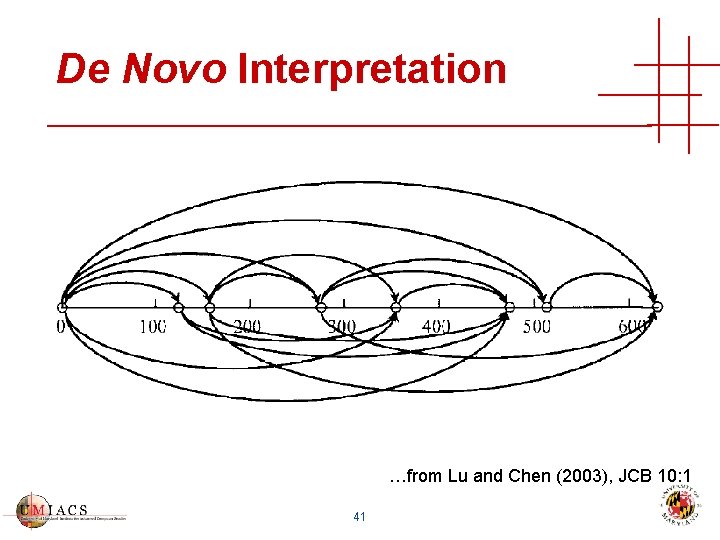
De Novo Interpretation …from Lu and Chen (2003), JCB 10: 1 41

De Novo Interpretation • Find good paths in spectrum graph • Can’t use same peak twice • Simple peptide fragmentation model • Usually many apparently good solutions • Amino-acids have duplicate masses! • “Best” de novo interpretation may have no biological relevance • Identifies relatively few peptides in highthroughput workflows 42

Sequence Database Search • Compares peptides from a protein sequence database with spectra • Filter peptide candidates by • Precursor mass • Digest motif • Score each peptide against spectrum • Generate all possible peptide fragments • Match putative fragments with peaks • Score and rank 43
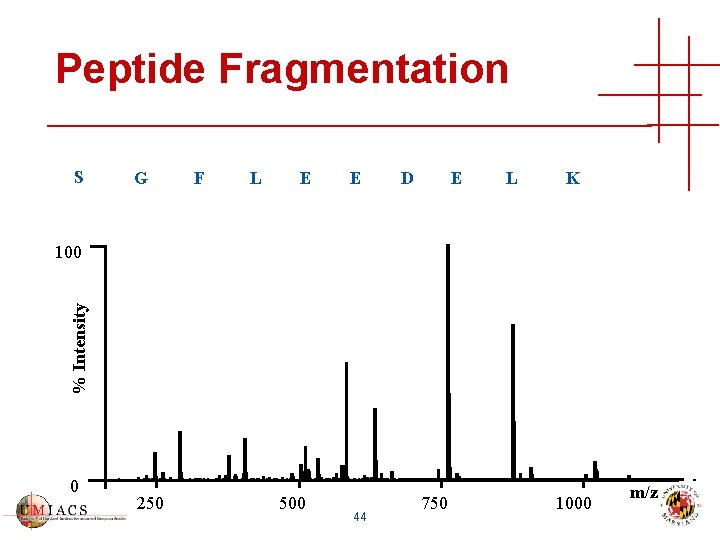
Peptide Fragmentation S G F L E E D E L K % Intensity 100 0 250 500 44 750 1000 m/z
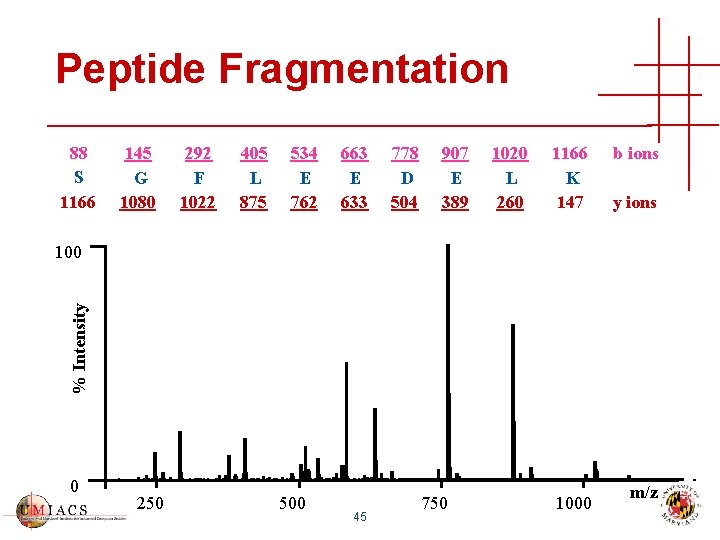
Peptide Fragmentation 88 S 1166 145 G 1080 292 F 1022 405 L 875 534 E 762 663 E 633 778 D 504 907 E 389 1020 L 260 1166 K 147 b ions y ions % Intensity 100 0 250 500 45 750 1000 m/z
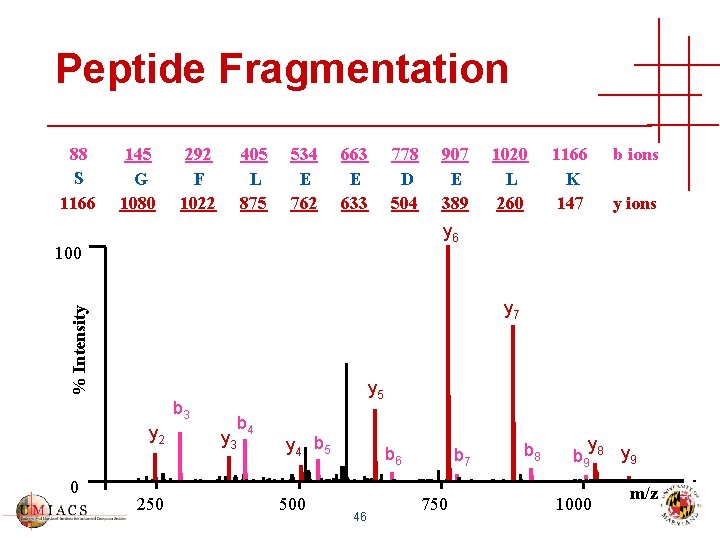
Peptide Fragmentation 88 S 1166 145 G 1080 292 F 1022 405 L 875 534 E 762 663 E 633 778 D 504 1020 L 260 1166 K 147 b ions y 6 100 % Intensity y 7 y 5 b 3 y 2 0 907 E 389 250 y 3 b 4 y 4 b 5 500 b 6 46 b 7 750 b 8 y b 9 8 y 9 1000 m/z

Sequence Database Search • Sequence fills in gaps in the spectrum • All candidates have biological relevance • Practical for high-throughput peptide identification • Correct peptide might be missing from database! 47

Peptide Candidate Filtering Digestion Enzyme: Trypsin • Cuts just after K or R unless followed by a P. • Must allow for “missed” cleavage sites • “Average” peptide length about 10 -15 amino-acids 48

Peptide Candidate Filtering >ALBU_HUMAN MKWVTFISLLFLFSSAYSRGVFRRDAHKSEVAHRFKDLGEENFKAL VLIAFAQYLQQCPFEDHVKLVNEVTEFAK… No missed cleavage sites MK WVTFISLLFLFSSAYSR GVFR R DAHK SEVAHR FK DLGEENFK ALVLIAFAQYLQQCPFEDHVK LVNEVTEFAK 49 …

Peptide Candidate Filtering >ALBU_HUMAN MKWVTFISLLFLFSSAYSRGVFRRDAHKSEVAHRFKDLGEENFKAL VLIAFAQYLQQCPFEDHVKLVNEVTEFAK… One missed cleavage site MKWVTFISLLFLFSSAYSRGVFRR RDAHKSEVAHRFK FKDLGEENFKALVLIAFAQYLQQCPFEDHVKLVNEVTEFAK 50 …

Peptide Scoring • Peptide fragments vary based on • The instrument • The peptide’s amino-acid sequence • The peptide’s charge state • Etc… • Search engines model peptide fragmentation to various degrees. • Speed vs. sensitivity tradeoff • y-ions & b-ions occur most frequently 51
Mascot Search Engine 52

Mascot MS/MS Ions Search 53
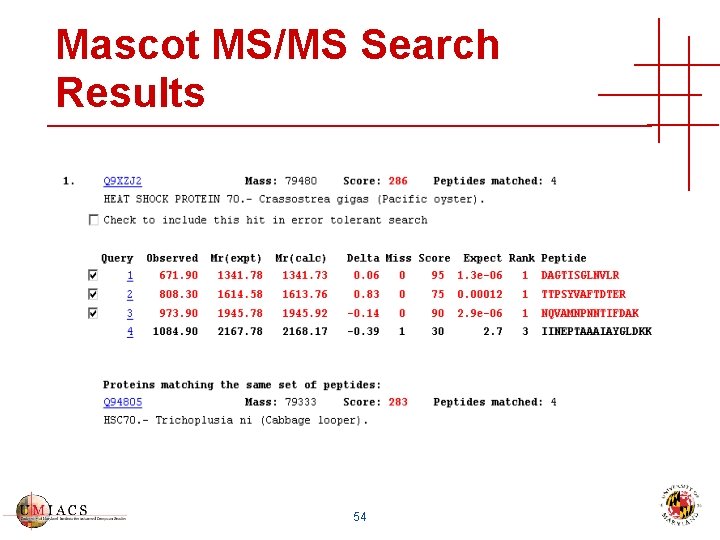
Mascot MS/MS Search Results 54

Mascot MS/MS Search Results 55
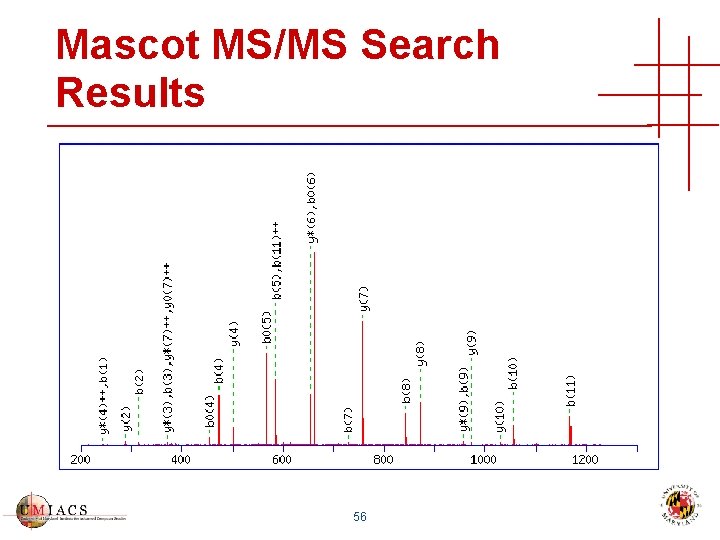
Mascot MS/MS Search Results 56
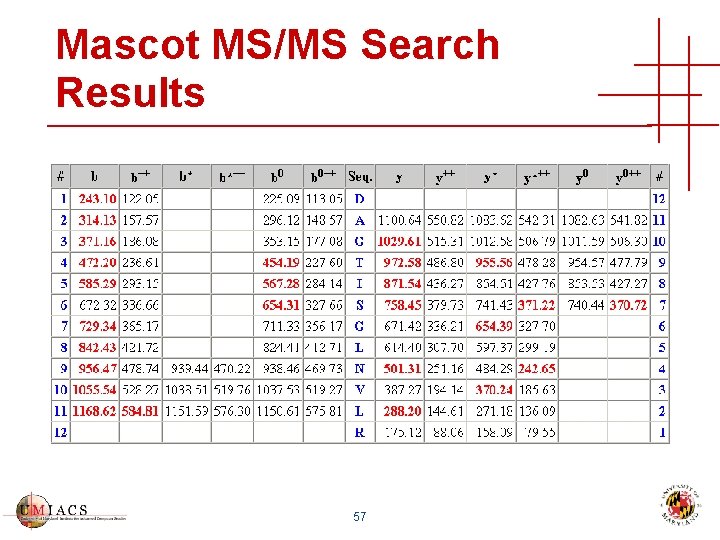
Mascot MS/MS Search Results 57
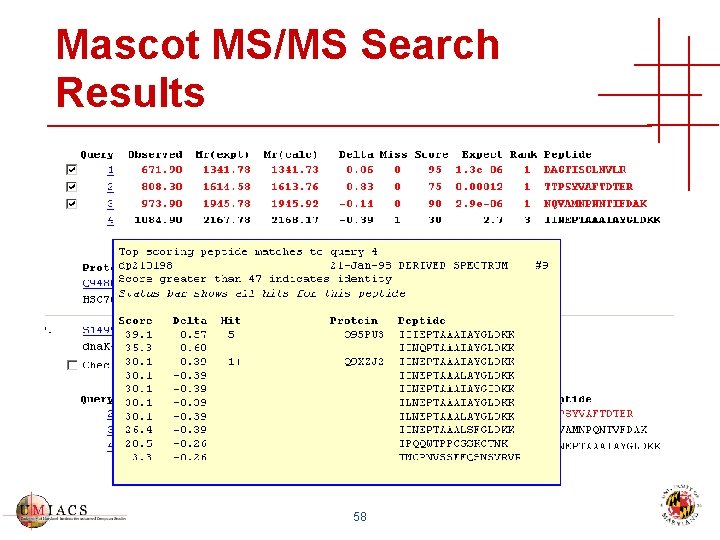
Mascot MS/MS Search Results 58
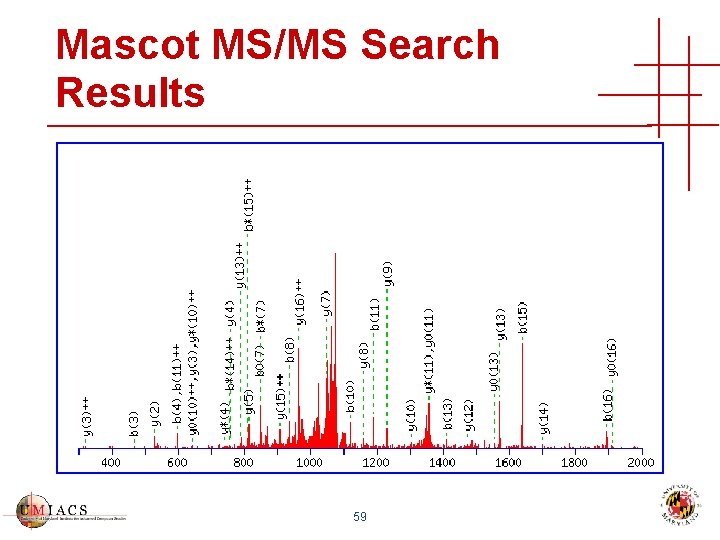
Mascot MS/MS Search Results 59
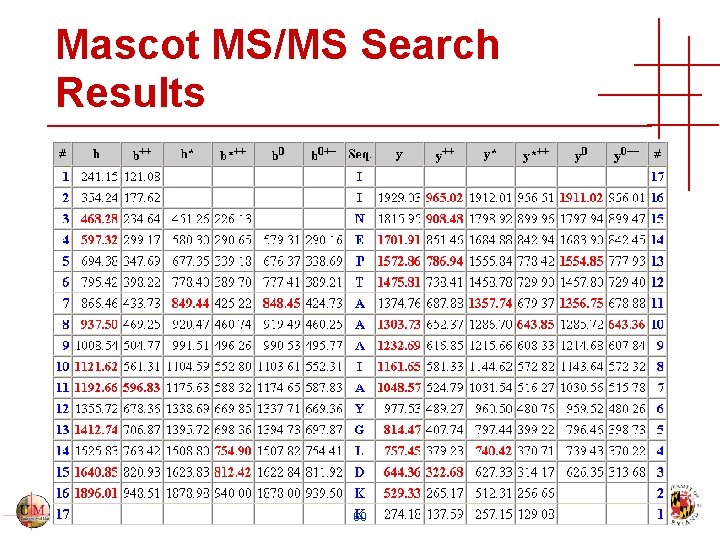
Mascot MS/MS Search Results 60

Mascot MS/MS Search Results 61
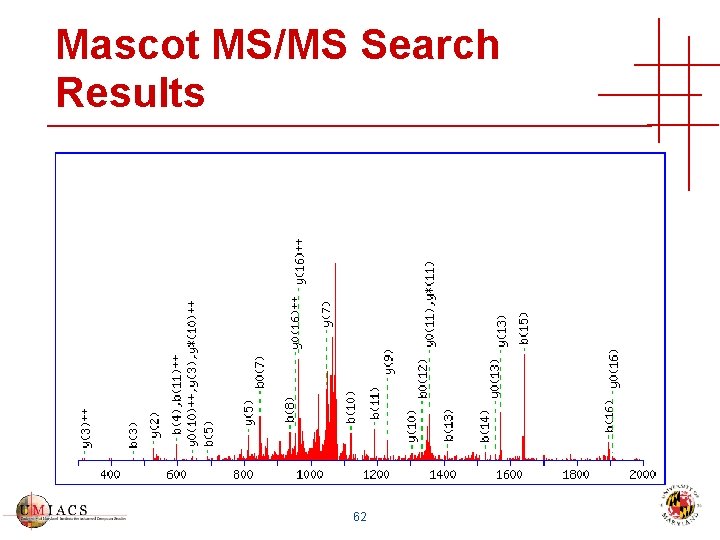
Mascot MS/MS Search Results 62
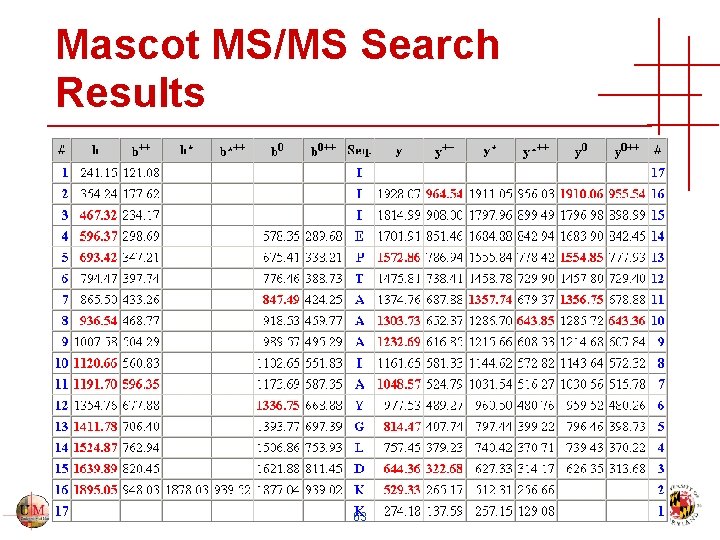
Mascot MS/MS Search Results 63

Summary • Protein identification by mass spectrometry is a key element of proteomics and systems biology. • Mass spectrometry + sequence databases represent a huge leap for protein (bio-)chemistry. • Sample prep, instruments and algorithms still maturing, much work to be done. 64

Further Reading • Matrix Science (Mascot) Web Site • www. matrixscience. com • Seattle Proteome Center (ISB) • www. proteomecenter. org • Proteomic Mass Spectrometry Lab at The Scripps Research Institute • fields. scripps. edu • UCSF Protein. Prospector • prospector. ucsf. edu 65
- Slides: 65